Author response:
The following is the authors’ response to the original reviews.
Reviewer #1 (Public Review):
Summary:
Sun et al. generated germline-specific cKO mice for the Znhit1 gene and examined its effect on male meiosis. The authors found that the loss of Znhit1 affects the transcriptional activation of pachytene. Znhit1 is a subunit of the SRCAP chromatin remodeling complex and a depositor of H2AZ, and in cKO spermatocytes, H2AZ is not deposited into the gene region. The authors claim that this is why the PGA was not activated. These findings provide important insights into the mechanisms of transcriptional regulation during the meiotic prophase.
Strengths:
The authors used samples from their original mouse model, analyzing both the epigenome and the transcriptome in detail using diverse NGS analyses to gain new insights into PGA. The quality of the results appeared excellent.
Weaknesses:
Overall, the data is inconsistent with the authors' claims and does not support their final conclusions. In addition, the sample used may not be the most suitable for the analysis, but a more suitable sample would dramatically improve the overall quality of the paper.
Thank you for your comprehensive summary of our study and your thoughtful insights into its strengths and weaknesses. We greatly appreciate this valuable feedback, which helps us further improve our work. Below, we provide a detailed response addressing each of the points you raised.
Reviewer #1 (Recommendations For The Authors):
Major revisions:
Surprisingly, many genes were upregulated in the scRNA-seq results. How many XY genes are included? Discuss why many genes are up-regulated in Fig. 5E whereas bulk RNA-seq showed only 70 genes were down-regulated. Since apoptosis-related factors are up-regulated in Fig5E, could these up-regulated genes be due to the high content of the transcriptome of dead cells? As you know, cell death starts, but randomly and violently disrupts the transcriptome, so we think it is not desirable to analyze the transcriptome with dead cells in the mix. Describe this point appropriately in the text or generate new data without dead cells.
We sincerely appreciate the reviewer’s critical points. Below, we address each point sequentially:
(1) To address the question about XY-linked genes, we utilized scRNA-seq data to identify differentially expressed sex chromosome genes in spermatocytes at different stages. Our analysis revealed an aberrant activation of XY-linked genes relative to controls. Specifically, 120 XY-linked genes were aberrantly activated in zygotenestage spermatocytes, and 119 XY-linked genes showed aberrant activation in pachytene-stage spermatocytes (revised Fig. 4F). This observation directly indicates that Znhit1 knockout impairs Meiotic Sex Chromosome Inactivation (MSCI), a finding that aligns with our prior characterization of XY chromosome synapsis defects in Znhit1-deficient spermatocytes.
(2) Two key reasons explain the discrepancy between scRNA-seq and bulk RNA-seq results:
First, scRNA-seq employs a more permissive threshold for identifying DEGs (log2 fold change [log2FC] = 0.25), thereby enhancing sensitivity to subtle expression changes and enabling the detection of more upregulated genes. In contrast, bulk RNAseq uses a stricter threshold (log2FC = 1), which filters out these subtly upregulated transcripts, resulting in fewer DEGs overall.
Second, scRNA-seq can capture cell subset-specific differential expression. In contrast, bulk RNA-seq averages signals across mixed cells, masking such subsetspecific expression changes.
These clarifications have been included in the Data Analysis section of the revised manuscript.
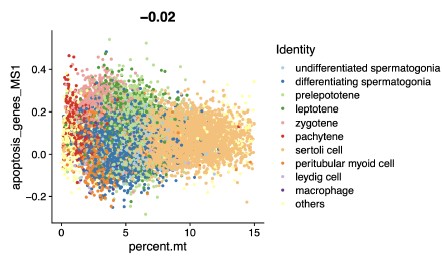
(3) We fully agree with the reviewer’s concern that dead cells could confound transcriptomic analyses. Before downstream analysis, we excluded non-viable cells via stringent QC: cells with mitochondrial RNA (mtRNA) content exceeding 15% were removed, as high mtRNA content is a well-established marker of cell death or compromised viability. To further validate that upregulated genes were not driven by dead cell contamination, we analyzed the correlation between the expression of apoptosis-related genes and mtRNA fractions in our data. This analysis revealed no significant correlation (Pearson correlation coefficient, r = -0.02; please see Author response image 1). These results collectively rule out dead cell transcriptome contamination as the primary cause of the observed gene upregulation.
Author response image 1.
Scatter Chart showing the Pearson correlation between apoptosisrelated genes and mitochondrial RNA fractions in scRNA-seq data.
Line 280-286: The data in Figures 7I and J are confusing: as shown by KAS-seq, it is natural that ssDNA is not formed in the promoter region in Znhit1-cKO sample because transcription does not proceed, but why is ssDNA formed in the enhancer region in the first place in control and then lost in Znhit1-cKO sample? Generally, it is said that in the enhancer region, including the super-enhancer region, doublestranded DNA is not dissociated, thus not forming ssDNA. Discuss why the loss of ssDNA in the enhancer region affects transcription with appropriate citations. Also, show whether genes downstream of the missing ssDNA in the promoter region have abnormal transcriptional activity, along with the RNA-seq data. Furthermore, in the region shown in Figure 7I, why the chromatin is even more open, as shown by ATACseq in Znhit1-cKO. Discuss whether this is related to transcriptional progression or aberrant substitution with H2A. If the function of ZNHIT1 is to replace H2A with H2AZ for PGA, it is not necessary to show the H2A level in Znhit1-cKO.
We appreciate the reviewer’s constructive comments.
(1) ssDNA dynamics in enhancer regions: Emerging evidence demonstrates that active enhancers undergo transient DNA unwinding to form ssDNA, a process critical for transcriptional regulation by transcribing enhancer RNAs (eRNA). KAS‑seq is sufficiently sensitive to detect ssDNA in enhancer regions (Kim et al., 2010; Wu et al., 2020). It has been shown that H2A.Z (deposited by the ZNHIT1-SRCAP complex) is required for maintaining enhancer accessibility and dynamic unwinding (Sporrij et al., 2023). In this study, we found that Znhit1 deletion and defective H2A.Z incorporation impaired enhancer ssDNA formation, indicating that ZNHIT-H2A.Z plays an important role in the activity of both promoter and enhancer.
(2) Impact of ssDNA loss on transcription: To address how missing ssDNA affects transcriptional activity, we further analyzed changes in KAS‑seq signals following Znhit1 knockout. Overall, KAS‑seq signals were significantly reduced upon Znhit1 depletion, confirming that Znhit1 is essential for ssDNA formation. Further examination of KAS‑seq signals at promoters of downregulated genes also revealed reduced signals (revised manuscript, Fig. S8). In contrast, KAS-seq signals of upregulated genes remained relatively low and showed no changes in both the control and knockout groups, and their upregulation probably results from indirect regulation. These results underscore the importance of ZNHIT1-mediated chromatin states in regulating ssDNA formation and gene expression.
(3) Aberrant chromatin openness in Znhit1-cKO (ATAC-seq): The increased chromatin accessibility detected by ATAC-seq likely represents a disorganized, nonfunctional state rather than productive transcriptional openness. H2A.Z normally constrains chromatin dynamics to facilitate ordered transcriptional regulation (Cole et al., 2021); its absence in Znhit1-cKO leads to higher ATAC-seq signals, suggesting that this aberrant openness fails to support proper assembly of the transcriptional machinery.
Minor revisions:
Line 106. The text says that they looked for chromatin factors, but the legend says that they looked for epigenetic factors. The text must be consistent.
We have corrected it in the revised manuscript (line 801).
Line 107. Although it is stated that the transcriptional data published here were used, it appears from the cited references that they are scRNA-seq data. A clear explanation is required in the text or legend.
We have revised this data as scRNA-seq data (line 107).
Line 141-143: Using TUNEL analysis in Figure 4F, the authors show that Znhit1cKO testis cells contain many dead cells. Describe the type or stage of the apoptotic cells.
We appreciate the reviewer’s suggestion. Specifically, we performed TUNEL staining on testes isolated from P14 mice, a critical time point for pachytene development (revised Fig. 2D). We tested this by showing that apoptosis-related genes were significantly upregulated in pachytene-stage spermatocytes in scRNA-seq data (revised Fig. 4D). To further validate this observation, we performed scRNA-seq from P35 testis samples. The results revealed a significant reduction in late pachytene-stage spermatocytes in Znhit1-cKO samples (revised Fig. 2F), consistent with apoptotic loss of pachytene cells. Collectively, these data confirm that Znhit1 knockout impairs pachytene-stage spermatocyte development.
The authors claimed that the loss of Znhit1 lowers the transcription of a group of genes involved in homologous recombination, including Rnf212, causing a delay in homologous recombination; however, if the process of homologous recombination is delayed, homologous chromosome pairing and synapsis are affected unless DSB repair is completed. Provide a satisfactory explanation for the fact that DNA damage remains on autosomes despite complete synapsis, as shown in Figure 3C, which is likely not solely due to delayed homologous recombination.
Thank you for this insightful comment. We fully agree that persistent autosomal DNA damage cannot be explained solely by delayed homologous recombination. To resolve this question, we further analyzed autosomal synapsis through SYCP1 and SYCP3 staining. While autosomal synapsis appeared morphologically complete, we identified subtle but significant synapsis defects in autosomal terminal regions (revised Fig. 3A). This suggests that Znhit1 knockout also results in autosomal synapsis defects. We speculate that these synapsis defects are associated with the unresolved autosomal DNA damage we observed.
Lines 150-163. With regard to XY unpairing in Znhit1-cKO pachytene spermatocytes, there is insufficient discussion as to whether this is due to transcriptional aberrations.
Thank you for highlighting the need to link transcriptional aberrations to XY unpairing in Znhit1-cKO pachytene spermatocytes. To address this, we analyzed sex chromosome transcription using scRNA-seq data. Relative to controls, 120 XYlinked genes were aberrantly activated at zygotene, and 119 were upregulated at pachytene in Znhit1-cKO spermatocytes (revised Fig. 4F), directly demonstrating Znhit1 knockout disrupts Meiotic Sex Chromosome Inactivation (MSCI). Given that intact MSCI is required to stabilize XY synapsis in pachytene spermatocytes, we conclude that the observed XY unpairing is likely a direct consequence of these sex chromosome transcriptional abnormalities. We add this information to the revised manuscript (lines 221-226).
Line 187-194. Analysis of the scRNA-seq data is shown in Figure 4, but it lists several genes as stage-specific markers, some of which do not have well-understood meiotic functions. Please cite a reference paper that provides sufficient evidence to qualify this stage.
In response to this comment, we have refined the presentation of marker genes used for cell annotation (revised Fig. S4B). We have incorporated relevant references supporting their utility as stage-specific markers for the meiotic stages (line 187).
Line 225-233: If Znhit1 is important for H2AZ deposition and regulates PGA through it, how does it regulate HR-related genes that are expressed earlier through H2AZ deposition during the pachytene stage? For example, Rnf212 is not specifically expressed during the pachytene stage but is one of the targets of MEIOSIN, so it is expressed at an earlier stage.
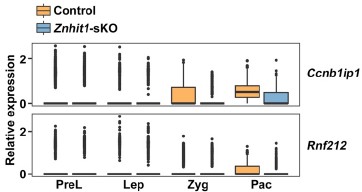
Thank you for this insightful comment. We fully acknowledge the reviewer’s key observation that HR-related genes such as Rnf212 are MEIOSIN targets that initiate transcription at earlier meiotic stages, before the pachytene stage. Our stage-resolved scRNA-seq data further showed that the expression of Ccnb1ip1 and Rnf212 was significantly upregulated from zygotene to pachytene, following their initial transcriptional onset. We next showed that the loss of H2A.Z deposition induced by Znhit1 deletion specifically impaired this pachytene-specific secondary transcriptional activation, rather than the early MEIOSIN-driven expression onset (please see Author response image 2).
Author response image 2.
Plots showing the expression level of indicated genes in scRNAseq data.
Line 245-251: As shown in Figure 6E, more than 14,000 genes have H2AZ peaks. In contrast, only approximately 60% of the genes downregulated by Znhit1-cKO appeared to be directly affected by H2AZ. Are the remaining 40% of genes regulated in a different way that is not mediated by H2AZ? Also, only a few percent of the genes with H2AZ peaks are affected, but why are only genes with A-MYB involvement affected, as shown in Figure 7?
Thank you for these insightful and constructive comments. For the ~40% of downregulated genes not directly linked to H2A.Z, they were likely regulated through indirect mechanisms. H2A.Z deposition mediated by ZNHIT1 may influence upstream transcriptional regulators (e.g., transcription factors or coactivators), whose dysregulation in turn affects these genes.
The selective effect of H2A.Z loss on A-MYB target genes is explained by the strict context-dependent function of H2A.Z, which requires stage-specific partner transcription factors to exert its regulatory activity. During the zygotene-to-pachytene transition, A-MYB acts as the master regulator of pachytene gene activation and forms a functional collaborative complex with H2A.Z to drive target gene transcription. Disrupted H2A.Z deposition upon Znhit1 deletion specifically impairs the activity of this A-MYB-H2A.Z complex, leading to selective downregulation of A-MYB targets. Other H2A.Z peak-associated genes may rely on alternative cofactors and compensatory mechanisms.
Line 245-256: Figures 6 and F show that the localization of H2AZ is reduced in Znhit1-cKO mice, which means that no substitution with H2A occurs. If so, show it in the data because the localization of H2A should be increased compared to that in the control.
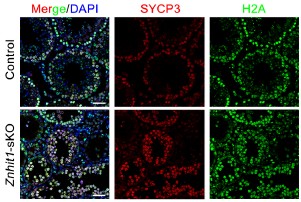
To clarify the status of H2A, we have now detected immunofluorescent staining against H2A. While H2A.Z deposition was clearly impaired following Znhit1 deletion, the global level of H2A did not change significantly (Author response image 3). We speculate that this observed absence of a compensatory increase in H2A is likely due to the intrinsically low abundance of the histone variant H2A.Z relative to canonical histone H2A under physiological conditions.
Author response image 3.
Immunostaining of SYCP3 and H2A in spermatocyte testis sections of control and Znhit1-sKO mice, Scale bar, 40 μm.
Reviewer #2 (Public Review):
Summary:
The study demonstrates that Znhit1 regulates male meiosis, with deletion causing pachytene failure associated with defective expression of pachytene genes and subtle effects on X-Y pairing and DSB repair. The authors attribute this phenotype to the defective incorporation of the Znhit1 target H2A.Z into chromatin.
Strengths:
The paper and the figures are well presented and the narrative is clear. Evidence that the conditional deletion strategy removes Znhit1 is strong, with multiple orthogonal approaches used. Most of the meiotic phenotyping is well performed, and the omics analysis clearly identifies a dramatic effect on the meiotic gene expression program. The link to H2A.Z and A-MYB adds a mechanistic angle to the study.
Weaknesses:
(1) Current literature demonstrates that meiotic mutants arrest at one of two stages: midpachytene (stage IV of the seminiferous cycle) or metaphase I (stage XII of the seminiferous cycle). This study documents that in the Znhit1 KO the midpachytene marker H1t appears normally, but that cells arrest before diplotene. If this is true, then arrest must occur during late pachytene, which based on my knowledge has never been documented for a meiotic KO. To resolve this, the authors should present stronger histological substaging evidence to support their claim.
Thank you for this insightful and constructive comment. To achieve highresolution tracking of cell lineage progression, we performed scRNA-seq analysis using P35 testes in this revised manuscript. scRNA-seq data showed that germ cells normally progressed through all meiotic stages and successfully gave rise to spermatids in control groups. By contrast, in the Znhit1 knockout group, late pachytene spermatocytes decreased significantly, and only very few subsequent germ cell types were observable (revised Fig. 2F, G). In scRNA-seq data, although very few diplotene spermatocytes and meiotic metaphase I cells were detectable, these cells still appeared abnormal, as evidenced by their extremely low Pou5f2 expression. We have revised our description of the meiotic arrest stage in the manuscript.
(2) The authors overlooked the possible effects of Znhit1 deletion on MSCI. Defective MSCI is a well-established cause of pachytene arrest. Actually, the fact that they see X-Y pairing failure should alert them even more strongly to this possibility because MSCI failure is often associated with defective X-Y pairing. This could be easily addressed by examination of their RNAseq data.
To address the concern that Znhit1 deletion may impact Meiotic Sex Chromosome Inactivation (MSCI), we analyzed XY-linked gene expression using scRNA-seq data from spermatocytes at distinct stages. Our analysis revealed aberrant activation of XY-linked genes in Znhit1-CKO spermatocytes relative to controls. Specifically, 120 XY-linked genes were activated at zygotene, and 119 XY-linked genes were upregulated at pachytene (revised Fig. 4F). This observation directly demonstrates that Znhit1-CKO impairs MSCI, which aligns with our prior characterization of defective X-Y chromosome synapsis in Znhit1-deficient spermatocytes. To explicitly resolve this concern, we have integrated these MSCIfocused RNA-seq analyses into the revised Results section (lines 221-226).
(3) The recombination assays need attention.
In the text the authors state that they studied RPA2 and DMC1, but the figures show RPA2 and RAD51.
The RPA counts are not quantitated.
The conclusion that crossover formation fails (based on MLH1 staining) is not justified. This marker does not appear in wt males until late pachytene, so if cells in this mutant are dying before that stage, MLH1 cannot be assessed.
The authors state that gH2AZ persists in the KO, but I'm not convinced that they are comparing equivalent stages in the wt and KO. In Figure 3C, the pachytene cell is late, whereas in the mutant the pachytene cell is early or mid (when residual gH2AX is expected, even in wt males).
Previous work (PMID: 23824539) has shown that antibodies reportedly detecting pATM in the sex body are non-specific. I therefore advise caution with the data shown in Figure 3D.
We appreciate the reviewer’s detailed feedback on our recombination assays and have addressed each concern as follows:
(1) Discrepancy between text and figures (RPA2/DMC1 vs. RPA2/RAD51): We have corrected this in the revised manuscript.
(2) Quantitation of RPA2 foci: We have supplemented quantitative analysis of RPA2 foci (revised Fig. S3).
(3) Conclusion on crossover failure: Single-cell RNA sequencing data from P35 testes definitively confirmed that Znhit1 knockout spermatocytes successfully progressed to the late pachytene stage, ruling out the possibility that our MLH1 staining results are confounded by cell death or arrest before this critical stage. In addition, analysis of transcriptome datasets revealed significant downregulation of important genes required for homologous recombination and crossover formation, including Ccnb1ip1 and Rnf212. Reduced expression of these essential factors may impair the assembly of MLH1 crossover foci. These data demonstrate that ZNHIT1 is essential for proper homologous recombination and crossover formation during male meiosis. We have revised the text to emphasize this context.
(4) γH2AX persistence and stage matching: We have replaced the images with more representative, stage‑matched pachytene spermatocytes from wild‑type and Znhit1‑KO mice (revised Fig. 2C). Furthermore, prompted by the insightful comment from Reviewer 1, we carefully re‑examined autosomal synapsis and identified abnormal synapsis specifically at the terminal regions of autosomes in Znhit1‑deficient spermatocytes (revised Fig. 3A). These data together confirm that ZNHIT1 is essential for DSB repair during male meiotic prophase I.
(5) pATM staining issue: Following the reviewer’s advice, we carefully reviewed the relevant literature (PMID: 23824539) and confirmed that the anti‑pATM antibody may exhibit non‑specific staining on the XY chromosomes. Accordingly, we have removed the pATM staining data presented in Figure 3D from the revised manuscript to ensure the accuracy and rigor of our results.
(4) RNAseq data. The authors show convincingly that Znhit1 activates genes that are normally upregulated at the zyg-pachytene transition. They should repeat the analysis for genes normally upregulated at the prelep- lep and lep-zyg transition to show that this effect is really pachytene-gene specific.
We appreciate this suggestion. To clarify the stage specificity of ZNHIT1’s regulatory role, we analyzed genes upregulated at the prelep-lep and lepzyg transitions. Our results showed that Znhit1 knockout had little impact on the overall expression levels of these genes (as shown in revised Fig. 4B). In contrast, as we previously reported, genes upregulated at the zygotene-pachytene transition were remarkably downregulated in Znhit1-cKO. These findings further confirm the specificity of ZNHIT1 in regulating pachytene gene expression.
(5) I am puzzled that the title and overall gist of the study focuses on H2A.Z, when it is Znhit1 that has been deleted.
We appreciate the reviewer’s observation and have revised the study title as suggested. Specifically, the title is now updated to “ZNHIT1-dependent H2A.Z deposition at meiotic prophase I underlies pachytene gene expression and meiotic progression during male meiosis.”
Reviewer #3 (Public Review):
Summary:
Sun et al. present a manuscript detailing the phenotypic characterization of loss of Znhit1 in male germ cells. Znhit1 is a subunit of the chromatin regulating complex SRCAP that functions to deposit the histone variant H2A.Z. Given that meiosis, and specifically meiotic recombination, occurs in the context of the dynamic condensing of chromosomes, the role of chromatin regulators in general, and histone variants specifically, in mammalian meiosis is an active area of research. Previous work has shown that H2A.Z is found at the locations of recombination in plants, although H2A.Z was previously not found at recombination sites in mammalian meiosis. Here the authors use a conditional approach to ablate Znhit1 in spermatocytes and characterize a block in meiosis in prophase I in the transition from pachytene to diplotene stage.
Strengths:
The authors combine current methods in immunohistochemistry and functional genomics to provide strong evidence of meiotic block upon the loss of Znhit1. They find that loss of Znhit1 leads to reduced incorporation of the histone variant H2A.Z, specifically at promoters and enhancers. Further, RNA sequencing found more genes are down-regulated upon loss of Znhit1 compared to upregulated, suggesting that incorporation of H2A.Z is critical for the expression of genes necessary for successful meiotic progression.
A strength of the manuscript is tying the locations of changes in H2A.Z deposition with binding of the transcription factor A-MYB, providing a mechanism that can potentially combine the changes in chromatin regulation with variable binding of a transcription factor in gene expression in pachytene stage spermatocytes.
Weaknesses:
A weakness in the single-cell RNA experiment using cells from 16-day-old male mice. The authors suggest that the rationale for the experiment was to determine where the Znhit1-sKO mutant showed an arrest in meiosis, and claim that this is the pachytene stage. However, in the 'first wave' of meiosis 16-day-old mice are just beginning to enter pachytene, so cells from later meiotic stages will be largely absent in these tubules. This is clear from the UMAP showing a similar pattern of cell distributions between wild-type and mutant mice. Using older mice would have better demonstrated where the mutant and wild-type mice differ in cell-type composition.
We appreciate the reviewer’s constructive comment. To resolve this issue, we have added new scRNA‑seq data from testes of P35 mice, which harbor a full spectrum of meiotic stages, including late pachytene, diplotene, metaphase I spermatocytes, and post-meiotic spermatids. Compared with wild-type controls, Znhit1-sKO testes exhibited a marked reduction in late pachytene spermatocytes and a near-complete loss of post-pachytene cell types, directly validating the pachytenestage meiotic arrest (revised Fig. 2F, G). All updated analyses have been integrated into the manuscript to strengthen our conclusions.
The authors use the term pachytene genome activation (PGS) in the manuscript to suggest a novel process by which genes are specifically increased in expression in the pachytene stage of meiotic prophase I, without reference to literature that establishes the term. If the authors are putting forward a new concept defined by this term, it would strengthen the manuscript to describe it further and delineate what the genes are that are activated and discuss potential mechanisms.
We appreciate the reviewer’s valuable feedback on our use of the term "pachytene genome activation (PGA)".
To address this, we have revised the text to explicitly frame PGA as a stage-specific transcriptional program observed in our data, defined by the coordinated upregulation of a distinct set of genes during the pachytene stage of meiotic prophase I.
(1) Definition and Gene Set: Using the scRNA-seq dataset, we formally defined PGA as the transcriptional wave characterized by genes with increased expression in pachytene vs. zygotene spermatocytes (n = 1,560 genes). Functional enrichment analysis shows these genes are primarily involved in DNA repair, cilium organization, and spermatid development (Table S3), consistent with the biological process of germ cell development.
(2) Relationship to existing literature: While PGA as a term is not widely established, our data align with prior observations of pachytene-specific transcriptional upregulation (Alexander et al., 2023; Ernst et al., 2019; Turner, 2015). Importantly, Alexander et al reveals that in late meiotic stages, starting from pachynema, chromatin has a ~3-fold increase in transcription. We have added these citations to clearly illustrate the relevant advances in the field (lines 68-71).
(3) Regulation of pachytene-stage gene expression: We further delineate that PGA is regulated by ZNHIT1-dependent H2A.Z deposition. Znhit1 deletion resulted in significant downregulation of 70.1% (1,094 out of 1,560) of these genes. This links PGA to chromatin-based regulation, where ZNHIT1-dependent H2A.Z deposition enables pachytene-specific transcription.
Generally speaking, the authors present solid evidence for a pachytene block in male germ cell development in mice lacking Znhit1 in spermatocytes. The evidence supporting a change in gene expression during pachytene, that more genes are downregulated in the mutant compared to increased expression, and changes in histone modification dynamics and placement of H2A.Z all support a role in alterations in meiotic gene regulation. However, the support that changes in H2A.Z impacting meiotic recombination (as suggested in the manuscript title) is less supported, rather than a general cell arrest in the pachytene stage leading to cell death. The conclusions around the role of Znhit1 influencing meiotic recombination directly could use further justification or mechanistic hypothesis.
We acknowledge the reviewer’s comments. Indeed, existing data support the presence of a pachytene block in spermatocytes of Znhit1-deficient mice, along with aberrant pachytene gene expression and impaired H2A.Z deposition.
In response, we made the following revisions: (1) we adjusted the manuscript title and conclusion to reduce emphasis on a direct H2A.Z-recombination link, and focus instead on ZNHIT1/H2A.Z in pachytene gene regulation and meiotic progression; (2) recombination defects may be indirect consequences of failed pachytene gene regulation, rather than a direct regulatory effect of ZNHIT1 on recombination machinery (lines 314-319).
Reviewer #3 (Recommendations For The Authors):
Quality of the images for meiotic spreads - images have low contrast and are tiny. It is difficult to see the SYCP3 results even when the images are magnified on the computer screen.
We have provided new images with high resolution to ensure a clear visualization of SYCP3 signals.
Line 165 - indicates the results for DMC1, although the figure suggests the results are for RAD51 foci.
We have corrected this mistake.
Line 306 - this manuscript 'confirms' that H2AZ is not found at mammalian recombination sites, a result already in the literature.
We have corrected this mistake (lines 309-312).
Reviewing Editor Comments:
Major points and revisions highlighted by the reviewers:
(1) Meiotic prophase in Znhit1KO: The main questions to clarify are the stage and status of progression, the analysis of apoptosis, and the consequences of gene expression on the X and Y. Additional analysis for DSB repair foci, gH2AX is also required. Those analysis are needed to answer to reviewer 2. Even if H2AZ was not detected at recombination hotspots, it may be possible that it plays a role in DSB repair but the level is too low for detection. This should be discussed as H2AZ was shown to be involved in DNA repair.
We sincerely appreciate the reviewing editor’s constructive comments.
(1) Stage and progression of meiotic prophase: We supplement P35 testes for scRNAseq. Results confirmed Znhit1-KO spermatocytes arrest at late pachytene, and postpachytene stages (diplotene, metaphase I) were nearly absent (revised Fig. 2F, G).
(2) Apoptosis analysis: We studied this by demonstrating that apoptosis-related genes were upregulated in pachytene spermatocytes at the single-cell level (revised Fig. 4D). To further validate this finding, we performed scRNA-seq analysis on P35 testis samples. Our results revealed a marked reduction in late pachytene spermatocytes in Znhit1-cKO testes (revised Fig. 2F, G), consistent with apoptotic depletion of pachytene-stage cells. Together, these data confirm that Znhit1 ablation impairs pachytene-stage spermatocyte development.
(3) X/Y gene expression consequences: To address this key point, we performed stage-resolved analysis of XY-linked gene expression using scRNA-seq data from different-stage spermatocytes. Compared with controls, we detected aberrant ectopic activation of XY-linked genes in Znhit1-KO spermatocytes: 120 XY-linked genes were inappropriately activated at zygotene, and 119 remained abnormally upregulated at pachytene (revised Fig. 4F). These results provide direct evidence that Znhit1 deletion impairs Meiotic Sex Chromosome Inactivation (MSCI).
(4) DSB repair issue: We have replaced the images with more representative, stage‑matched pachytene spermatocytes (revised Fig. 3C). The revised images show consistently increased γH2AX signals in Znhit1-KO spermatocytes. Prompted by Reviewer 1’s comment, we identified abnormal synapsis at autosomal terminal regions in mutant cells. Together, these results confirm that ZNHIT1 is essential for DSB repair during male meiotic prophase I.
(5) Potential role of H2A.Z in DSB repair: Though H2A.Z was nearly undetectable at recombination hotspots, we discuss two possibilities: (1) ZNHIT1-H2A.Z depletion dysregulated DSB repair-related genes; (2) Current ChIP-seq sensitivity may miss low-abundance H2A.Z at hotspots, which could support repair via chromatin remodeling. Future high-resolution assays (super-resolution imaging, DSB-targeted ChIP-seq) are proposed to validate this. We agree that recombination defects may be indirect consequences of failed pachytene gene regulation, rather than a direct regulatory effect of ZNHIT1 on recombination machinery.
(2) Gene expression analysis. The first consequence of H2AZ depletion is gene expression downregulation. However, it may be not surprising that some genes are down and others upregulated. There are likely secondary and indirect effects including the upregulation of some genes. The authors should explain and discuss this point such as to answer to questions raised by reviewer 1 and 2.
The primary consequence of H2A.Z depletion in pachytene spermatocytes is indeed widespread downregulation of genes. For the coexistence of upregulated genes, we explain this via three key points.
(1) Technical differences between scRNA-seq and bulk RNA-seq (addressing Reviewer 1): scRNA-seq captures cell-type-specific differentially expressed genes that bulk RNA-seq masks (bulk averages signals across mixed cells, hiding changes in rare subsets). Additionally, scRNA-seq uses a lower log2(fold change) threshold (0.25 vs. 1 in bulk RNA-seq), detecting subtle upregulations missed by bulk analysis.
(2) No dead cell contamination (addressing Reviewer 1): Stringent quality control excluded cells with >15% mitochondrial RNA. Apoptosis-related genes showed no significant correlation with mitochondrial RNA fractions (Pearson correlation coefficient, r = -0.02; please see Author response image 1), ruling out dead cell transcriptome interference.
(3) Secondary/indirect effects (addressing Reviewers 1 & 2): Upregulated genes likely result from indirect regulatory cascades. H2AZ depletion may disrupt upstream transcription factors, leading to compensatory upregulation of their downstream genes or cell stress responses to meiotic arrest. Notably, Znhit1 knockout specifically impacts genes upregulated at the zygotene-pachytene transition, while genes upregulated at preleptotene-leptotene or leptotene-zygotene transitions remain largely unaffected (revised Fig. 4B), confirming the specificity of H2A.Z’s direct regulatory role and framing upregulation as non-targeted indirect effects.
(3) The authors should also test the effect of Znhit1KO on the 1196 genes (up PreL/L) and 1325 (up L/Z) as shown in Figure 5D for the PGA. Also in Figure 5B, there is no evaluation of the statistical significance of the variation, this should be revised. X and Y genes should be analysed. KAS-Seq should be correlated with gene expression analysis, and several points as mentioned in the reviews below should be better explained and discussed.
(1) Effect of Znhit1-KO on PreL/L- and L/Z-upregulated genes: we analyzed the 1196 genes upregulated at the PreL/L transition and 1325 genes upregulated at the L/Z transition. Znhit1 knockout had minimal effect on the expression of these early meiotic gene sets (revised Fig. 4B), whereas genes activated at the zygotene‑pachytene transition were strongly downregulated in Znhit1-KO spermatocytes. These results confirm the specific role of ZNHIT1 in regulating pachytene‑stage gene expression. We have also added a statistical evaluation for the variation shown in Fig. 4B.
(2) X/Y-linked gene analysis: Analysis of stage‑resolved scRNA‑seq revealed aberrant ectopic activation of 120 XY‑linked genes at zygotene and 119 at pachytene in Znhit1-KO spermatocytes (revised Fig. 4F), demonstrating impaired Meiotic Sex Chromosome Inactivation (MSCI).
(3) KAS-seq correlation with gene expression: We analyzed the link between KAS‑seq signals and gene expression, and we found that Znhit1 depletion caused a global reduction in KAS‑seq signals, especially at promoters of downregulated genes (revised Fig. S8). Genes with increased expression showed low KAS‑seq signals in both control and mutant groups, likely reflecting indirect regulation. These results highlight the essential role of ZNHIT1 in transcriptional regulation.
(4) The title should refer to Znhit1, and the effect on meiotic recombination activities may be an indirect consequence of prophase progression arrest, even if some recombination genes are downregulated. This point is important as noted by reviewer 3.
We fully acknowledge Reviewer 3’s key point and have revised the manuscript title to “ZNHIT1-dependent H2A.Z deposition at meiotic prophase I underlies pachytene gene expression and meiotic progression during male meiosis” to reduce emphasis on a direct H2A.Z-recombination link.
Regarding meiotic recombination activities: The downregulation of recombinationrelated genes (e.g., Ccnb1ip1, Rnf212) stems from impaired pachytene-stage transcriptional programs caused by ZNHIT1-dependent H2A.Z deposition defects, which in turn leads to prophase progression arrest. Thus, the observed recombination abnormalities may be a secondary consequence of the meiotic prophase arrest, rather than a direct regulatory effect of ZNHIT1 on recombination machinery. This clarification has been integrated into the Discussion section (lines 314-318).
(5) The recent structural analysis of SRCAP should be cited: Yu et al. Cell Discovery (2024) 10:15 https://doi.org/10.1038/s41421-023-00640-1.
We have cited this reference in this revised manuscript (lines 234-236).
(6) The authors should read and answer the specific revisions asked for by the reviewers.
We have thoroughly read and systematically addressed all specific revisions requested by Reviewers 1, 2, and 3, as detailed in the revised manuscript and supplementary data.
References
Alexander, A.K., Rice, E.J., Lujic, J., Simon, L.E., Tanis, S., Barshad, G., Zhu, L., Lama, J., Cohen, P.E., and Danko, C.G. (2023). A-MYB and BRDT-dependent RNA Polymerase II pause release orchestrates transcriptional regulation in mammalian meiosis. Nature communications 14.
Cole, L., Kurscheid, S., Nekrasov, M., Domaschenz, R., Vera, D.L., Dennis, J.H., and Tremethick, D.J. (2021). Multiple roles of H2A.Z in regulating promoter chromatin architecture in human cells. Nature communications 12, 2524.
Ernst, C., Eling, N., Martinez-Jimenez, C.P., Marioni, J.C., and Odom, D.T. (2019). Staged developmental mapping and X chromosome transcriptional dynamics during mouse spermatogenesis. Nature communications 10, 1251.
Kim, T.K., Hemberg, M., Gray, J.M., Costa, A.M., Bear, D.M., Wu, J., Harmin, D.A., Laptewicz, M., Barbara-Haley, K., Kuersten, S., et al. (2010). Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182-187.
Sporrij, A., Choudhuri, A., Prasad, M., Muhire, B., Fast, E.M., Manning, M.E., Weiss, J.D., Koh, M., Yang, S., Kingston, R.E., et al. (2023). PGE(2) alters chromatin through H2A.Z-variant enhancer nucleosome modification to promote hematopoietic stem cell fate. Proceedings of the National Academy of Sciences of the United States of America 120, e2220613120.
Turner, J.M. (2015). Meiotic Silencing in Mammals. Annu Rev Genet 49, 395-412. Wu, T., Lyu, R., You, Q., and He, C. (2020). Kethoxal-assisted single-stranded DNA sequencing captures global transcription dynamics and enhancer activity in situ.
Nature methods 17, 515-523.





 - dill green
- dill green