est également classé zone humide d'importance internationale depuis 1987
Contexte historique pour situer la particularité de l’emplacement étudié (reconnaissance UNESCO, une certaine protection).
est également classé zone humide d'importance internationale depuis 1987
Contexte historique pour situer la particularité de l’emplacement étudié (reconnaissance UNESCO, une certaine protection).
Des biopsies de peau et de lard ont été réalisées chez des dauphins du parc national des Everglades et à Key West (dans les Keys - petites îles au sud de la Floride) afin de comprendre l’influence de leur habitat sur leurs niveaux et leurs profils de contamination
Description sommaire de la méthode et exposition des buts de la recherche.
huella computacional,
Es interesante como se puede llegar a rastrear todo, hasta los archivos eliminados o fantasmas todo bajo el rastreo de la huella computacional
Agregar 2 rondas más de combates a la tabla pokemenTournament, para un total de 3 rondas, incluyendo la primera ronda agregada en el código previo.
Al realizar la ejecutar el combate en el torneo pokemon entre Pikachu y Pidgey, siguiendo la narrativa de datos se logro adicionar las filas correspondientes a los tres round en la tabla
Vista errorea del Data Frame para guardar el torneo Pokemon. Si que quiere ver la vista correcta hay que cliquera la soalapa “Tree”, en lugar de la solapa “Data”, en la parte superior.
Me presenta un error y no estoy seguro si se debe a que coloqué varios Fighter en el código. Posiblemente estoy repitiendo nombres o variables sin control, lo que podría estar generando conflicto entre lo que quiero ejecutar.
corte1 := HedgeDoc new url: 'https://docutopia.sustrato.red/unisema:25A13'; retrieveContents
HedgeDoc es una herramienta colaborativa en línea que permite crear y editar documentos de manera simultánea entre varios usuarios, facilitando la sincronización de contenidos entre diferentes clases. Además, puede integrarse con Glamorous Toolkit, lo que permite sincronizar y gestionar esos documentos dentro de un entorno de desarrollo avanzado basado en Pharo Smalltalk.


Data Frame
Un DataFrame es una tabla de datos estructurada en filas y columnas que permite organizar, manipular y analizar información de forma sencilla y ordenada en programación y ciencia de datos.

No me está funcionando la tabla, creo que me falta definir alguna variable o estoy olvidando algún paso.
fighter1 := Pokemon new name: 'pikachu'. fighter2 := Pokemon new name: 'ditto'.
Estoy ejecutando Fighter 1 hasta Fighter 6 para organizar la lista de los Pokémon que tengo en sus Pokébolas, de esta manera puedo mantener un orden claro y definido sobre su respectiva posición o turno.
Pero no se si estoy ejecuntado mal al tener varios Fighter.

Pokemon,

La primera generación de Pokémon, que inició con los juegos de Game Boy (Rojo, Verde y Azul en Japón), presenta 151 especies diferentes. Estos 151 Pokémon son todos originarios de la región de Kanto. El último Pokémon de esta generación es Mew, el número 151.
Pokemones e información a partir de datos en JSON
Se puede utilizar JSON para almacenar la información de un Pokémon, incluyendo su nombre, tipo, habilidades y estadísticas, organizados en una estructura de datos clara y accesible.
Es importante tener conocimientos básicos de programación, ya que al momento de ejecutar código pueden presentarse errores en algunos casos. Contar con esas bases permite identificar, corregir y entender mejor los problemas que surgen durante el desarrollo.
Grafoscopio
Grafoscopio es una plataforma de código abierto basada en Pharo Smalltalk, una versión actualizada que facilita la creación, lectura y distribución de cuadernos computacionales interactivos.
Ideal para dejar documentado no solo los resultados sino también el proceso que se lleva, permitiendo a otros replicarlo o adaptarlo.
>>> 'https://chiselapp.com/user/Valentina.P/repository/Valentina-P/timeline'
El enlace que compartiste lleva a la línea de tiempo del repositorio “Valentina-P” en ChiselApp, gestionado por Valentina Penagos Ariza.
Este repositorio documenta sus actividades académicas y de programación, especialmente en temas de computación y lecturas hipertextuales.
En la línea de tiempo se pueden ver los distintos cambios y publicaciones que ha hecho en sus proyectos.
3 de junio de 2025: Eliminó un archivo con un nombre incorrecto.
2 de junio de 2025: Publicó el proyecto Batalla Pokémon.
1 de junio de 2025: Subió su proyecto final.
21 de mayo de 2025: Publicó una actividad autónoma usando JSON.
7 de mayo de 2025: Actualizó y publicó el ejercicio Challenge Yourself.
26 y 28 de abril de 2025: Publicó lecturas hipertextuales y ejercicios de clase.
pokemonTournament
En este caso la tabla sale en blanco
Y agreguémosla a nuestro torneo:
siguiendo las instrucciones no agrega la fila porque devuelve un valor nulo
e libro de Data Frame,
Si bien pude guiarme con un ejemplo del libro, me fue imposible generar la tabla en su totalidad, no pude nombrar las columnas, pero si pude ingresar los contenidos.
Vista de impresión de la pelea entre dos Pokemones.
Cuando ejecuto el código y dejo en vista de impresión un resultado pero no como un arreglo como se ve en el ejemplo
Importe esta narrativa a su propio GToolkit/Grafoscopio y exportarla en su propio portafolio
Respecto a este apartado tengo problemas para poder importar esta misma narrativa. Puesto que cuando coloco el enlace como aprendimos a hacerlo en clase del 31 de marzo se me congela por completo el programa de Glamorous Toolkit. No sé si haya otra estrategia para poder importar la narrativa o si deba presentar una nueva narrativa completamente hecha por mí.

Si se alcanza, intentar traducir esas frases anteriores en español a su equivalente en código.
En este apartado, realice el ejercicio con las partes de código del torneo Pokémon, pero no pude organizar los datos en la tabla, porque se me genera solo con los daos vacíos. Entonces seguí los ejemplos del libro de los DataFrame, pero me sigue generando el mismo error, se crea la tabla, pero no se puede organizar dentro de la estructura y en algunos lados genera errores que desconozco si son de la ejecución del código, o es que falta adicionar algo al Software Grafoscopio
pokemonTournament addRow: (roundBetweenFighters value: fighter1 value: fighter2) named: 'round 1'
No me deja realizar la acción me sale este error

pokemonTournament := DataFrame withColumnNames: #(figther1 fighter2 moveFigther1 moveFigther2 winner)
Aqui solo le estamos diciendo que le ponga columnas a la tabla pero no las filas ¿ hay que ponerlas?
randomMove := [ :fighter | (fighter data at: 'moves') atRandom at: 'move' at: 'name' ]
Lo que decimos aquí es: es tráigame aleatoriamente de los datos de cualquier luchador de la lineal movimientos un movimiento al azar.
fighter1 data
Con este comando podemos traer de la API todos los atributos del objeto

fighter1 := Pokemon new name: 'pikachu'. fighter2 := Pokemon new name: 'ditto'.
Para el proyecto final he creado la CLASE: pokemon con sus objetos " fighter" y a la vez cada uno de estos con sus atributos para mi proyecto he creado 6 luchadores:



round := { fighter1 name. fighter2 name. (randomMove value: fighter1). (randomMove value: fighter2). { fighter1 . fighter2 } atRandom name }
Sobre los códigos de las rondas tengo un problema. Precisamente cuando ejecuto el código tal cual como lo vimos en clase me aparece "value was sent to nil"
Estuve revisando las grabaciones de las clases pero no encuentro ningún error anterior, de hecho, me había funcionado hasta la parte donde debemos utilizar los dataframes. Pero a partir de ahí cuando quiero poner una ronda no me lo permite.
Este es el código que he estado utilizando.
round1 := { fighter1 name. fighter2 name.
(randomMove value: fighter1) at: 'move' at: 'name'.
(randomMove value: fighter2) at: 'move' at: 'name'.
{ fighter1 . fighter2 } atRandom }
Y esto es lo que aparece cuando lo ejecuto.
 De saber alguna solución posible lo agradecería
De saber alguna solución posible lo agradecería
STONJSON fromString: ('https://pokeapi.co/api/v2/pokemon?limit=-1' asUrl retrieveContents
Extrañamente el código se me ejecuta correctamente, pues me da la misma respuesta, pero no entiendo cómo podemos extraer los nombres, pues al no poder acceder a este Pokemon atRandom no me permite elegir en aleatorio entre todos los nombres de todos los Pokemones.
winner := { fighter1 . fighter2 } atRandom
No declara ganador ya que no esta definida la variable
finalRound := { randomMove value: fighter1 . randomMove value: fighter2 }
Al ejecutarla para seguir el ejercicio se produjo error de respuesta nula como en la ante rior
rior
fighter1 := Pokemon new name: 'pikachu'. fighter2 := Pokemon new name: 'ditto'.
Cuando estoy realizando el mismo ejemplo no lo ejecuta porque hay un error esobre que declarada la varaiable.
Pokemon atRandom
Si bien entiendo la utilidad del código, pues nos asegura elegir que las peleas sean elegidas por dos pekemon aleatorios, por el mismo error de 'data' no puedo ejecutar este código, pues no se reconoce 'atRandom', supongo que por la misma razón. De igual manera, como mi código está modificado, haré a mi propia manera una forma de elegir los pokemones aleatorios.
pokemonTournament
Con este código podemos ver nuestra tabla de le grabamos a nuestra DataFrame (si no estoy mal, por lo menos lo que yo entiendo que estamos haciendo)
pokemonTournament addRow: (roundBetweenFighters value: fighter1 value: fighter2) named: 'round 1'
Algo importante que aclarar con este código es que, como agrega un row, no se puede ejecutar dos veces, porque el row que volvemos a ejecutar ya existe, lo que nos produce un error.
fighter1 := Pokemon new name: 'pikachu'. fighter2 := Pokemon new name: 'ditto'.
Desde aquí tuve una incidencia debido a la instalación incorrecta de minidocs. Cosa que pude resolver con ayuda de otra persona que tuvo el mismo error en su debido momento
pokemonTournament := DataFrame withColumnNames: #(figther1 fighter2 moveFigther1 moveFigther2 winner)
En pocas palabras, esto lo que hace es dejar el nombre de nuestras columnas hechas para luego llenarlas con la información de las batallas. Prácticamente estamos dejando el nombre de las columnas que luego llenaremos.
fighter1 data
Durante la lectura anotada e interactiva de este documento en mi GToolkit, parece que la clase Pokemon no me entiende el 'data'. En lo que he experimentado autónomamente con la herramienta, puedo suponer que es porque el 'data' debe agregarse como newMethods dentro de la clase de Pokemon. Si bien esto no impide el trabajo del proyecto, me parece importante mencionarlo.

pokemonDataDictionaries := #('pikachu' 'ditto' 'charizard') collect: [:name | pokemonDataDictionary value: name ]
Funciona para obtener los datos de diferentes pokemones a la vez, y almacenarlos en una variable.
pokemonDataDictionary := [ :name | | dataLink pokemonRawData | dataLink := 'https://pokeapi.co/api/v2/pokemon/', name. pokemonRawData := dataLink asUrl retrieveContents. STONJSON fromString: pokemonRawData ]
Este código particularmente no me salió, pero con la lógica del mismo realice el cambio para que se hiciera con Snorlax
pokemonExampleLink value: 'pikachu'
Te devuele la URL del pokemon que pongas entre comilas
pokemonExampleLink := 'https://pokeapi.co/api/v2/pokemon/pikachu'
realicé el cambio de Pikachu por ''Snorlax'' y el resultado fue el mismo, únicamente con la edición del Pokémon que indiqué
Pokemones e información a partir de datos en JSON
Ha sido una forma diferente de entender la programación y la cantidad de posibilidades que existen una vez que aprendes a entender la lógica de los lenguajes. Pokémon en este caso es el conjunto de datos con los que estamos trabajando (Criaturas con diferentes poderes).
studentRepositories keys atRandom
Atrandom es el comando que he estado implementando para elegir un Pokémon al azar, y hará parte da la construcción de los round de mis pokemones.
pokemonDataDictionaries collect: [:item | {item at: 'name' . item at: 'height' } ]
Si lo interpreto bien, se supone que lo que le estamos pidiendo es: en bloque tomando la variable 'item' que será cada valor de nuestro arreglo anterior, entrá a cada uno y tre el valor de los metadatos 'name' y 'height'. Así el resultado será un listado del número de valores de nuestro arreglo, con su respectivo nombre y altura.
Lo único que me molesta es que el orden que presenta es justamente como lo pedimos, es decir, los resultados están en el mismo orden que los valores de nuestro arreglo. No sé si haya una forma de volver la información de 'height' en un dato al que luego podamos ponerle una condición de mayor a menor número.
pokemonDataDictionaries := #('pikachu' 'ditto' 'charizard') collect: [:name | pokemonDataDictionary value: name ]
Al final esto es justamente lo que intenté hacer en la última clase, tener un listado con los nombres de los Pokemones y un bloque que me permita visualizar la información estructurada de cada uno. No pensé que este sería el resultado de ello, pero me voy satisfecha.
Este tiene dos partes, que se muestran en la figura anterior: las llaves, ubicadas a la izquierda, y los valores, ubicados a la derecha.
Esta representación es muy útil si lo queremos adaptar a otros ejercicios. Si bien el reto es, precisamente, como adaptamos este código para una nueva necesidad que surja, me parece interesante como los nombres son las llaves y el link personal de cada repositorio. Lo interpretó como una especie de BUSCARV de excel interesante para analizar a profundidad.
capturar información de los repositorios de código en Fossil para las distintas asignaturas que usaran dicha herramienta
Sería bueno seguir desarrollando como capturar información de los repositorios de código podría servirnos para otras asignaturas. Obviando el hecho de su relación en búsqueda de información, es importante resaltar que importancia puede tener esta captura de información en espacios de otras materias que pueden usar esta herramienta. Si bien el ejercicio con PokeAPI nos permite hacer la parte práctica, se identifica el vacío teórico de esta relevancia del ejercicio práctico.
El microentorno en el que comienza a surgir la respuesta inmune también puede influir en el resultado; el mismo patógeno puede tratarse de manera diferente dependiendo del contexto en el que se encuentra.
Diferente tratamiento para el mismo patógeno
Pigs are crucial sources of meat and protein, valuable animal models, and potential donors for xenotransplantation. However, the existing reference genome for pigs is incomplete, with thousands of segments and missing centromeres and telomeres, which limits our understanding of the important traits in these genomic regions. To address this issue, we present a near complete genome assembly for the Jinhua pig (JH-T2T), constructed using PacBio HiFi and ONT long reads. This assembly includes all 18 autosomes and the X and Y sex chromosomes, with only six gaps. It features annotations of 46.90% repetitive sequences, 35 telomeres, 17 centromeres, and 23,924 high-confident genes. Compared to the Sscrofa11.1, JH-T2T closes nearly all gaps, extends sequences by 177 Mb, predicts more intact telomeres and centromeres, and gains 799 more genes and loses 114 genes. Moreover, it enhances the mapping rate for both Western and Chinese local pigs, outperforming Sscrofa11.1 as a reference genome. Additionally, this comprehensive genome assembly will facilitate large-scale variant detection and enable the exploration of genes associated with pig domestication, such as GPAM, CYP2C18, LY9, ITLN2, and CHIA. Our findings represent a significant advancement in pig genomics, providing a robust resource that enhances genetic research, breeding programs, and biomedical applications.
A version of this preprint has been published in the Open Access journal GigaScience (see paper https://doi.org/10.1093/gigascience/giaf048), where the paper and peer reviews are published openly under a CC-BY 4.0 license.
Revision 1 version
Reviewer 1: Martien Groenen
In their revised version of the manuscript, the authors have addressed all my major concerns raised in my earlier review and have made the many editorial edits as suggested. I only have a few (mostly editorial) comments for the revised version. The most important one is the title of the manuscript. I realize I did not mention this in my earlier review, but I think the title is not very appropriate and could be more informative. I suggest something like "A telomere-to-telomere genome assembly of the Jinhua pig"
Minor editorial comments: Line 40: Replace "provides" by "provide"; "genome" to "genomes" and "JH" to "Jinhua" Lines 50-51: "This study produced a gapless and near-gapless assembly of the pig genome, and provides a set of diploid JH reference genome." Should be changes to something like "This study produced a near-gapless assembly of the pig genome and provides a set of haploid Jinhua reference genomes." Line 177: Change "with with" to "with" Line 194: Replace "population" by "populations" Lines 232-233: Referring to human as a "closely related species" is rather awkward and not correct. I suggest replacing this with "eleven other mammals" Lines 299, 301 and 303: Insert "of" after "consisting" Line 317: Insert "and" before "2.33 Gb" Line 319: Insert "and" before "2.17 Gb" Line 320-321: Change to "The more continuous contigs of the two assemblies were selected to construct the final haploid assemblies". Line 323: Replace "assembly" by "assembler "Line 354: Delete "ranging" Lines 358359: Change "The average properly mapped rate" to "The average rate of properly mapped reads" Line 379: Insert "respectively" after "60.07"Line 380: "suggested" (remove space)Line 385: Change "indicate a gapless and near-gapless" to "indicate a near-gapless" Line 455: Change "were overlapped with" to "were overlapping with" Lines 557-559" The sentence "The insertion found in the SLA-DOB gene, which serves to enhance the immune system's response and is relevant to transplant rejection" seems incomplete and sound awkward. Perhaps you mean something like "The insertion found in SLA-DOB, a gene involved in enhancing the immune system's response to infection, might be relevant in relation to transplant rejection"
Reviewer 2: Benjamin D Rosen
The first near-complete genome assembly of pig: enabling more accurate genetic research
General comments: I thank the authors for addressing most of my points and providing more details on the parameters they have used. Unfortunately, I still have some unanswered questions regarding the methodology. My current understanding from the authors responses to my previous comments leads me to believe that the assembly has been scaffolded incorrectly. If the authors did indeed use HiC data to place 8 contigs into gaps and then joined those contigs without placing gaps at the joins or doing any further gap filling, that calls into question the validity of the assembly. Finally, the language needs further improvement for readability.
Specific comments: Line 85 - *will contribute to. Lines 187-191 - HiC interaction maps do not provide information for gap filling. Either this has been explained insufficiently, or it has been done incorrectly. Placing assembled sequences in the correct order does not mean that it is okay to join them without a gap. It is necessary to return to the gap filling procedure now that the contigs are in the correct order and attempt to fill them as done previously. Line 191 - Figure S3 - These HiC contact maps are not very informative they need to be labeled and have a scale bar. Additionally, contact maps can have a lack of signal due to a gap in the sequence or due to multimapping reads in repetitive regions being filtered so it's not clear what they are trying to show in A-C. The authors reply to my previous concern regarding the labeling of this figure does not help, furthermore, the figure legend in the supplemental materials is still insufficient. I think I understand that panels D and E are chr3 before and after misassembly correction, it would be helpful if the two panels were at the same scale. I still don't know why panel F is shown, how is this related to panel C and I don't see any red ellipses indicated by the legend. Line 275 - "ensemble from Duroc pigs" is incorrect. It is an "assembly of a Duroc pig". Lines 299, 301, 303 - "containing" not "consisting" Lines 306-308 - Again, HiC data orders and orients contigs, but it does not fill gaps. Please clarify how the assembly was reduced from 14 gaps to 6 gaps with HiC data. Was an additional round of gap filling performed? Lines 313-314 - How is the contig N50 larger than the scaffold N50 above? Lines 335-336 - Does this refer to the Merqury analysis? I don't think "using mapped K-mers" is correct here, please reword. Lines 367-368 - what does it mean that "8 out of 63 gaps were corrected" is this from the HiC ordering of contigs? Line 369 - what does the mapping between Sscrofa11.1 and JH-T2T shown in figure S6 have to do with the JH-T2T gap filling being described here? Line 369 - I previously asked about this supplemental table only containing 55 entries. The authors response "The other filled 8 gaps were resolved through adjustments made to the Hi-C map to correct misassembles. As a result, these gaps cannot be precisely located within the existing order of the assembly." indicates that contigs must have been incorrectly joined solely based on the HiC signal between contigs. The authors must know what contigs were added or joined to form the final assembly. It would be trivial to align the two assembly versions and identify the positions of the old contigs in the new assembly. I believe that these incorrectly joined contigs should be broken and put through the same gap filling procedure as performed earlier. Lines 375-378 - Dramatic coverage changes in read mappings as found in these figures are usually indicative of assembly errors. I do not agree that "These findings confirmed the accuracy and reliability" of the assembly. I suggest replacing the last sentence with something more measured such as "Although supported by some read data, the inconsistency of coverage across these gap filled regions suggests that caution should be used when interpreting findings in these regions, cross-referencing results with the gap positions (Supplementary Table S9) is advised." Line 375 - "evidenced by fully coverage" remove "fully", it isn't proper usage of the word and I wouldn't interpret the low coverage in many of these regions as "full coverage". Line 385 - should read "Overall, our assembly quality metrics indicate a near-gapless assembly of the pig genome" Line 390 - should read "a gapless T2T sequence for 16 out of 20" Line 396 - Supplemental table 10 not 9.Lines 398399 - according to supplemental table S4 and figure 3A, chromosome 2 also has a single telomere. Line 402 - the centromeres are not marked in Figure 3A.Line 402 - Figure S8 - please rename chr19 and chr20, chrX and chrY. Line 406 - "at early research" unclear what is meant by this. please reword. Line 423 - as indicated on line 397, 33 telomeres were identified, not 35.Line 426 - "The JH-T2T assembly IDENTIFIED 17 centromeres" Line 450 - "are located in" Line 453 - "these SVs are located in" Line 455 - Moreover, 12,129 genes overlap these SVs" Line 502 - "which contained 544 gaps" Line 841 - Figure 2 legend description is still incorrect. Only A is mapping rates, B and C are PM rates and base error rates.
Pigs are crucial sources of meat and protein, valuable animal models, and potential donors for xenotransplantation. However, the existing reference genome for pigs is incomplete, with thousands of segments and missing centromeres and telomeres, which limits our understanding of the important traits in these genomic regions. To address this issue, we present a near complete genome assembly for the Jinhua pig (JH-T2T), constructed using PacBio HiFi and ONT long reads. This assembly includes all 18 autosomes and the X and Y sex chromosomes, with only six gaps. It features annotations of 46.90% repetitive sequences, 35 telomeres, 17 centromeres, and 23,924 high-confident genes. Compared to the Sscrofa11.1, JH-T2T closes nearly all gaps, extends sequences by 177 Mb, predicts more intact telomeres and centromeres, and gains 799 more genes and loses 114 genes. Moreover, it enhances the mapping rate for both Western and Chinese local pigs, outperforming Sscrofa11.1 as a reference genome. Additionally, this comprehensive genome assembly will facilitate large-scale variant detection and enable the exploration of genes associated with pig domestication, such as GPAM, CYP2C18, LY9, ITLN2, and CHIA. Our findings represent a significant advancement in pig genomics, providing a robust resource that enhances genetic research, breeding programs, and biomedical applications.
A version of this preprint has been published in the Open Access journal GigaScience (see paper https://doi.org/10.1093/gigascience/giaf048), where the paper and peer reviews are published openly under a CC-BY 4.0 license.
Original version
Reviewer 1: Martien Groenen
The manuscript describes the T2T genome assembly for the Chinese pig breed Jinhua, which presents a vast improvement compared to the current reference genome of the Duroc pig TJTabasco (build11.1). The results and methodology use for the assembly are described clearly and the authors show the improvement of this assembly by a detailed comparison with the current reference 11.1. While clearly of interest to be published, several aspects of the manuscript should be improved. Most of these changes are minor modifications or inaccuracies in the presentation of the results.
However, there are two major aspects that need further attention:
The T2T assembly presented, represents a combination of the two haplotypes of the pig sequenced. I am surprised why the authors did not also develop two haplotype resolved assemblies of this genome. Haplotype resolved assemblies will be the assemblies of choice for future developments of a reference pan-genome for pigs. The authors describe that they have sequenced the two parents of the sequenced F1 individual, so why did they not use the trio-binning approach to also develop haplotype resolved assemblies. I, think adding these to the manuscript would be a vast improvement for this important resource.
The results described for the identification of selective sweep regions is not very convincing. This analysis shows differences in the genomes of two breeds: Duroc and Jinhua. However, these breeds have a very different origin of domestication of wild boars that diverged 1 million years ago, followed by the development of a wide range of different breeds selected for different traits. Therefore, the comparison made by the authors cannot distinguish between differences in evolution of Chinese and European Wild Boar, more recent selection after breed formation and even drift. To be able to do so, these analyses would need the inclusion of additional breeds and wild boars from China and Europe. Alternatively, the authors can decide to tone down this part of the manuscript or even delete it altogether, as it does not add to the major message of the manuscript.Minor comments Line 34: Change the sentence to: "with thousands of segments and centromeres and telomeres missing" Line 37: Insert "and Hi-C" after "long reads "Line 46: Delete " such as GPAM, CYP2C18, LY9, ITLN2, and CHIA" Line 54: Insert "potential" before "xenotransplantation" Line 82: Delete "in response to the gap of a T2T-level pig genome" as this does not add anything and the use of "gap" in this context is confusing. Line 93: Change "The fresh blood" to "Fresh blood" Line 100: The authors need to provide a reference for the SDS method. Lines 152-153, line 444, and table S6: This is confusing. The authors mention Genotypes from 939 individuals, but in the table it is shown that they have used WGS data. You need to describe how the WGS data was used to call the genotypes for these individuals. Furthermore, in line 444 you mention 289 JH pigs and 616 DU pigs which together is 905. What about the other 34 individuals shown in table S6?Line 244: Replace "were" by "was" and delete "the" before "fastp" Lines 287292: Here you use several times "length of xx Gb and yy contigs". This is not correct as the value for the contigs refers to a number and not a length. Rephase e.g. like "length of xx Gb and consisting of yy contigs" Line 294: The use of "bone" sems strange. Either use "backbone" or "core"Line 306: Replace "chromosome" by "genome" Lines 308-309: For the comment "Second, 16 of the 20 chromosomes were each represented by a single contig" you refer to figure 1D however from this figure it cannot be seen if the different chromosomes consist of a single or multiple contigs. Line 346: Do you mean build 11.1 with "historical genome version". If so, please use that instead. Line 349: "post-gap filled" Line 353: The largest gap is 35 kb not 36 kb. Figures 2F-I should be better explained in the legends and the main text (lines 353-358). Lines 378: For the 23,924 genes you refer to supp table S13. However, that table shows a list of SV enriched QTL not these genes. Furthermore, I checked all tables but a table with all the protein coding genes is missing. Line 380: For the 799 newly anchored genes, refer to table S10. Now you refer to table S17 which shows genes enriched KEGG pathways. Lines 383-386: For the higher gene density in GC rich regions, you refer to figure 1D, but it is impossible to see this correlation from figure 1D. For the density of genes and telomeres, you refer to figure 1G. However, that figure does not show gene densities only repeat densities. Line 406-407. This should be table S11.Lines 409412: For this result you refer to table S11. However, that table only shows data for the gained genes, not the lost genes. Lines 419-420: You refer to table S12 and figure 3B, but the information is only shown in figure 3B and not in table S12.Line 420: Replace "were" by "is" Line 422: Better to use "repeats" instead of "they" Line 425: "Moreover, 12,129 genes located in these SVs". Unclear to what "these" refers to and I assume that you mean genes that (partially) overlap with SVs? Also, this is an incomplete sentence (verb missing). Likewise, this number is not very meaningful as many of these SVs are within introns. It is much more informative to mention for how many genes SVs affect the CDS. Line 433 and table S14: This validation is not clear at all. What exactly are these numbers that are shown? You also mention "greater than 1.00" but the table does not contain any number that is greater than 1.00. Line 435: "Table" not "Tables" Line 436: Change to " SVs with a length larger than 500 bp "The term "invalidate" in figure 3D is rather awkward. Better to use "not-validated" and "validated" in this figure. Line 449: This should be Table S16. Line 452: There is not Table S18Lines 484-486: Change to "Similarly, in human, the use of the T2T-CHM13 genome assembly yields a more comprehensive view of SVs genome-wide, with a greatly improved balance of insertions and deletions [61]." Lines 500-501: Change to "For example, in human, the T2T-CHM13 assembly was shown to improve the analysis of global" Lines 517-528: This paragraph should be deleted as these genes have already been annotated and described in previous genome builds including 11.1. Why discuss these genes here? Following that line of thinking, almost every gene of the 20,000 can be discussed. Line 532: "%" instead of "%%" and insert "which" after "SVs" Lines 537-542: These sentences should be deleted. It is common knowledge that second generation sequencing is not very sensitive to identify SVs. The authors also do not provide any results about dPCR. Line 544: "affect" rather than "harbor" Lines 544-547: This is repetitive and has been stated multiple times so better to delete. Line 561: "which is serve to immune system's response and relevant to transplant rejection" This is an incorrect sentence and should rephrased. Lines 562-568: I don't agree with is statement and suggest to remove it from the discussion.
Reviewer 2: Benjamin D Rosen
The first near-complete genome assembly of pig: enabling more accurate genetic research. The authors describe the telomere-to-telomere assembly of a Jinhua breed pig. They sequenced genomic DNA from whole blood with PacBio HiFi and Oxford Nanopore (ONT) long-read technologies as well as Illumina for short reads. They generated HiC data for scaffolding from blood and extracted RNA from 19 tissues for short read RNAseq for gene annotation. A hifiasm assembly was generated with the HiFi data and scaffolded with HiC to chromosome level with 63 gaps. The scaffolded assembly was gap filled with contigs from a NextDenovo assembly of the ONT data bringing the gaps down to 14. Finally, the assembly was manually curated with juicebox somehow closing a further 8 gaps. This needs to be clarified. Standard assembly assessments were performed as well as genome annotation. The authors compared their assembly to the current reference, Sscrofa11.1, and called SVs between the assemblies. The SVs were validated with additional Jinhua and Duroc animals. They then identified signatures of selection present in some of the largest SVs.
General comments: The manuscript is mostly easy to read but would benefit from further editing for language throughout. The described assembly appears to be high quality and quite contiguous. Although the authors do mention obtaining parental samples and claim the assembly is fully phased, there is no mention of how this was done. There are many additional places where the methods could be described more fully including the addition of parameters used.
Specific comments: Line 39 - Figure 1 only displays 34 telomeres, not 35. Additionally, I was only able to detect 33 telomeres using seqtk telo. Seqtk only reports telomeres at the beginning and end of sequences, digging further, the telomere on chr2 is ~59kb from the end of the chromosome, perhaps indicating a misassembly. Lines 79-81 - there are not hundreds of species with gap free genome assemblies and reference 19 does not claim that there are. Line 82 - the assembly is not gap-free, replace with "nearly gap-free" Line 95 - were these parental tissue samples ever used? Lines 151-156 - this section would be better located below the assembly methods. Please number supplementary tables in order of their appearance in the text. Line 171 - please provide parameters used here and for all analyses. Lines 187-188 - how did rearranging contigs decrease the gaps? Was the same gap filling procedure used after HiC manual adjustments? Line 188 - Figure S3 - I don't understand the relationship between the panels nor what the authors are attempting to show. If panels A-C display chromosomes 2, 8, and 13, Why does D display chr3? Both panels C and E are labeled chr13 but they look nothing alike. Are D-E whole chromosomes or zoomed in views? Missing description of panel F. Lines 222-224 - why weren't pig proteins used? Ensembl rapid release has annotated protein datasets for 9 pig assemblies. Line 264 - although most will know this, make it clear that Sscrofa11.1 is an assembly of a Duroc pig. Line 292 - how was polishing performed? This is missing from the methods. Line 294 - should this read "selected it for the backbone of the genome assembly."? Lines 298-299 - methods? Line 314 - what is meant by "using mapped K-mers from trio Illumina PCR-free reads data"? Line 331 - accession numbers for assemblies would be useful. Line 333 - what is "properly mapped rate"? Do you mean properly paired mapping rate? Line 346 - what is the historical genome version? Line 349 - Supplemental Table S8 only has 55 entries including the 6 remaining gaps. Where are the other filled 8 gaps located? Lines 350-358 - read depth displays wouldn't show the presence of clipped reads which would indicate an improperly closed gap. It would be more convincing to display IGV windows containing these alignments showing that there are no clipped reads. Line 354 - Figure S5 needs a better legend. What is ref and what is own? Line 359 - the assembly is near-gapless. Line 359 - where is the data regarding assembly phasing? How was this determined to be fully phased? Line 363 - 16 of 20 chromosomes are gapless. Line 370 - only 33 telomeres were found at the expected location (end of the chromosome), if you count the telomere on chr2 59kb from the end, then 34 telomeres were identified. Line 372 - chr13 also only has a single telomere. It does not have a telomere at the beginning. Line 372 - chr19 is chrX correct? Line 374 - Figure 1G - It would be nice to have the centromeres marked on this plot (or in Figure 3A). Are the long blocks of telomeric repeats internal to the chromosomes expected? Line 423 - Figure 3A - there is no telomeric repeat at the beginning of chr4 or chrXLine 431 - why were only 5 pigs of each breed used to validate SVs when 100's of WGS datasets from the two breeds had been aligned? How were these 5 selected? Line 481 - Sscrofa11.1 only has 544 gaps.Line 492 - ONT data was used to fill more than 6 gaps. Gaps in the assembly were reduced from 63 to 14 using ONT contigs. Lines 588-589 - please make your code publicly available through zenodo, github, figshare, or something similar. Line 815-824 - Figure 2 - legend description needs to be improved. Only A is mapping rates, B and C are PM rates and base error rates. The color switch from A-C having European pigs in blue to D having JH-T2T in blue might confuse readers.
NA then
NA, then
a NA
an NA
установите
Так-с, тут момент такой. Нам надо в начале рекомендовать последнюю версию 4.8: https://support.microsoft.com/ru-ru/topic/microsoft-net-framework-4-8-%D0%B0%D0%B2%D1%82%D0%BE%D0%BD%D0%BE%D0%BC%D0%BD%D1%8B%D0%B9-%D1%83%D1%81%D1%82%D0%B0%D0%BD%D0%BE%D0%B2%D1%89%D0%B8%D0%BA-%D0%B4%D0%BB%D1%8F-windows-9d23f658-3b97-68ab-d013-aa3c3e7495e0
Затем сказать, что если с ним возникли проблемы, то попробуйте исходную версию 4.0 https://www.microsoft.com/ru-ru/download/details.aspx?id=17718
September 30thMorning
questo che cosa è? forse da togliere.
Pricing
La scritta Pricing va tolta. O 1. sostituirla con, ad es., "Join the Summer School" oppure "How to Apply" e scrivere sotto "Participation is free of charge, but an application is required by the indicated deadline. Places are limited and admission is based on selection." 2. se non si può cambiare la scritta pricing, bisogna scrivere immediatamente sotto "There are no registration fees". Poi a seguire: "Participation is free of charge, but an application is required by the indicated deadline. Places are limited and admission is based on selection." Poi dovremo aggiungere cosa copriamo noi (dopo che ne avremo parlato con Mimmo).
Scientific and Organizing Committee
Scientific Committee
Domenico De Stefano – Università di Trieste (UNITS) Michela Gnaldi – Università di Perugia (UNIPG)
Massimo Attanasio – Università di Palermo (UNIPA) Giovanna Boccuzzo – Università di Padova (UNIPD) Stefano Campostrini – Università Ca’ Foscari Venezia (UNIVE) Dalit Contini – Università di Torino (UNITO) Antonella D’Agostino – Università di Siena (UNISI) Mariano Porcu – Università di Cagliari (UNICA) Giancarlo Ragozini – Università di Napoli Federico II (UNINA) Arjuna Tuzzi – Università di Padova (UNIPD) Susanna Zaccarin – Università di Trieste (UNITS)
Organising Committee Simone Del Sarto – Università di Perugia (UNIPG) Niccolò Salvini – Università Cattolica del Sacro Cuore (UNICATT) Francesco Santelli ...
Erreur : source de la référence non trouvée
Reference to be fixed
Author response:
The following is the authors’ response to the previous reviews
Reviewer #1 (Public review):
Specific comments:
(1) It is difficult to appreciate that there is a "peripheral sub-membrane microtubule array" as it is not well defined in the manuscript. This reviewer assumes that this is in the respective field clear. Yet, while it is appreciated that there is an increased amount of MTs close to the cytoplasmic membrane, the densities appear very variable along the membrane. Please provide a clear description in the Introduction what is meant with "peripheral sub-membrane microtubule array".
A definition has been added to the Introduction.
(2) The authors described a "consistent presence of a significant peripheral array in the C57BL/ 6J control mice, while the KO counterparts exhibited a partial loss of this peripheral bundle.
Specifically, the measured tubulin intensity at the cell periphery was significantly reduced in the KO mice compared to their wild-type counterparts". In vitro "control cells had convoluted nonradial MTs with a prominent sub-membrane array, typical for β cells (Fig. 2A), KIF5B-depleted cells featured extra-dense MTs in the cell center and sparse receding MTs at the periphery (Fig. 2B,C)". Please comment/discuss why in vivo there are no "extra-dense MTs in the cell center".
We now add a discussion of this point, which we believe could be a manifestation of 3D shape of a beta cell in tissue and/or compensatory mechanisms in organisms.
(3) Authors should include in the Discussion a paragraph discussing the fact that small changes in MT configuration can have strong effects.
A paragraph added to the discussion.
Recommendations for the authors:
Reviewer #1 (Recommendations for the authors):
(1) Figure 1: Even though the reviewer appreciates that minor changes of MT configuration have severe effects, still the overall effects appear minor (40 vs. <50% or 35% vs. around 28%). Notably, there are no statistically significant differences in the different groups in Fig. 1Suppl-Fig.1 D. This reviewer is not sure if the combination of many not significantly different data points can result in significant changes and this should be checked by a statistician. Authors should include in the Discussion a paragraph discussing the fact that small changes in MT configuration can have strong effects.
We have now added the requested paragraph to the discussion. Indeed, the differences are small, and the significance is only detected in a data set with a large sample size in Fig. 1J,K (combined data sets with smaller sizes from Fig. 1-Suppl-Fig.1 D), consistent with the fact that a larger sample size generally provides more power to detect an effect.
(2) Unfortunately, the authors cannot block kinesin-1 resulting in microtubule accumulation in the cell center and then release the block (best inhibiting microtubule formation), to show that the MTs accumulated in the cell center will be transported to the periphery.
This is indeed the case at the moment, yes.
Minor comments:
- Abstract: β-cells vs. β cells (and throughout the manuscript)
- Page 4: "MTOC, the Golgi, (Trogden et al. 2019), and"
- Page 5: "β-cell specific"
- MT-sliding vs. MT sliding
- Kinesin 1 vs. kinesin-1
- Page 6, line 1: "β cells. actively"
- Page 7: "a microtubule probe", should be "MT"
- Page 9: "1μm" vs. "1 μm"
- Page 10: "demonstrate a dramatic effect" recommended is: "demonstrate a marked effect"
- Page 13, line 1: dramatically vs. markedly
- Page 13, line 5: "50μm" vs. "50 μm" (in general, there should be a space between number and unit?)
- "37 degrees C" vs. "37{degree sign}C"
- Animal protocol number?
- "Mice were euthanized by isoflurane inhalation"? What concentration? How long? More details are needed (no cervical dislocation?).
- Antibodies: more identifiers are needed.
- Antibody information in Key reagents and under 5. Reagents and antibodies do not fit (1:500 and 1:1000).
Thank you, we corrected all relevant information now.
Author response:
The following is the authors’ response to the original reviews
Public Reviews:
Reviewer #1 (Public Review):
Summary:
The manuscript by Hussain and collaborators aims at deciphering the microtubule-dependent ribbon formation in zebrafish hair cells. By using confocal imaging, pharmacology tools, and zebrafish mutants, the group of Katie Kindt convincingly demonstrated that ribbon, the organelle that concentrates glutamate-filled vesicles at the hair cell synapse, originates from the fusion of precursors that move along the microtubule network. This study goes hand in hand with a complementary paper (Voorn et al.) showing similar results in mouse hair cells.
Strengths:
This study clearly tracked the dynamics of the microtubules, and those of the microtubule-associated ribbons and demonstrated fusion ribbon events. In addition, the authors have identified the critical role of kinesin Kif1aa in the fusion events. The results are compelling and the images and movies are magnificent.
Weaknesses:
The lack of functional data regarding the role of Kif1aa. Although it is difficult to probe and interpret the behavior of zebrafish after nocodazole treatment, I wonder whether deletion of kif1aa in hair cells may result in a functional deficit that could be easily tested in zebrafish?
We have examined functional deficits in kif1aa mutants in another paper that was recently accepted: David et al. 2024. https://pubmed.ncbi.nlm.nih.gov/39373584/
In David et al., we found that in addition to a subtle role in ribbon fusion during development, Kif1aa plays a major role in enriching glutamate-filled synaptic vesicles at the presynaptic active zone of mature hair cells. In kif1aa mutants, synaptic vesicles are no longer enriched at the hair cell base, and there is a reduction in the number of synaptic vesicles associated with presynaptic ribbons. Further, we demonstrated that kif1aa mutants also have functional defects including reductions in spontaneous vesicle release (from hair cells) and evoked postsynaptic calcium responses. Behaviorally, kif1aa mutants exhibit impaired rheotaxis, indicating defects in the lateral-line system and an inability to accurately detect water flow. Because our current paper focuses on microtubule-associated ribbon movement and dynamics early in hair-cell development, we have only discussed the effects of Kif1aa directly related to ribbon dynamics during this time window. In our revision, we have referenced this recent work. Currently it is challenging to disentangle how the subtle defects in ribbon formation in kif1aa mutants contribute to the defects we observe in ribbon-synapse function.
Added to results:
“Recent work in our lab using this mutant has shown that Kif1aa is responsible for enriching glutamate-filled vesicles at the base of hair cells. In addition this work demonstrated that loss of Kif1aa results in functional defects in mature hair cells including a reduction in evoked post-synaptic calcium responses (David et al., 2024). We hypothesized that Kif1aa may also be playing an earlier role in ribbon formation.”
Impact:
The synaptogenesis in the auditory sensory cell remains still elusive. Here, this study indicates that the formation of the synaptic organelle is a dynamic process involving the fusion of presynaptic elements. This study will undoubtedly boost a new line of research aimed at identifying the specific molecular determinants that target ribbon precursors to the synapse and govern the fusion process.
Reviewer #2 (Public Review):
Summary:
In this manuscript, the authors set out to resolve a long-standing mystery in the field of sensory biology - how large, presynaptic bodies called "ribbon synapses" migrate to the basolateral end of hair cells. The ribbon synapse is found in sensory hair cells and photoreceptors, and is a critical structural feature of a readily-releasable pool of glutamate that excites postsynaptic afferent neurons. For decades, we have known these structures exist, but the mechanisms that control how ribbon synapses coalesce at the bottom of hair cells are not well understood. The authors addressed this question by leveraging the highly-tractable zebrafish lateral line neuromast, which exhibits a small number of visible hair cells, easily observed in time-lapse imaging. The approach combined genetics, pharmacological manipulations, high-resolution imaging, and careful quantifications. The manuscript commences with a developmental time course of ribbon synapse development, characterizing both immature and mature ribbon bodies (defined by position in the hair cell, apical vs. basal). Next, the authors show convincing (and frankly mesmerizing) imaging data of plus end-directed microtubule trafficking toward the basal end of the hair cells, and data highlighting the directed motion of ribbon bodies. The authors then use a series of pharmacological and genetic manipulations showing the role of microtubule stability and one particular kinesin (Kif1aa) in the transport and fusion of ribbon bodies, which is presumably a prerequisite for hair cell synaptic transmission. The data suggest that microtubules and their stability are necessary for normal numbers of mature ribbons and that Kif1aa is likely required for fusion events associated with ribbon maturation. Overall, the data provide a new and interesting story on ribbon synapse dynamics.
Strengths:
(1) The manuscript offers a comprehensive Introduction and Discussion sections that will inform generalists and specialists.
(2) The use of Airyscan imaging in living samples to view and measure microtubule and ribbon dynamics in vivo represents a strength. With rigorous quantification and thoughtful analyses, the authors generate datasets often only obtained in cultured cells or more diminutive animal models (e.g., C. elegans).
(3) The number of biological replicates and the statistical analyses are strong. The combination of pharmacology and genetic manipulations also represents strong rigor.
(4) One of the most important strengths is that the manuscript and data spur on other questions - namely, do (or how do) ribbon bodies attach to Kinesin proteins? Also, and as noted in the Discussion, do hair cell activity and subsequent intracellular calcium rises facilitate ribbon transport/fusion?
These are important strengths and as stated we are currently investigating what other kinesins and adaptors and adaptor’s transport ribbons. We have ongoing work examining how hair-cell activity impacts ribbon fusion and transport!
Weaknesses:
(1) Neither the data or the Discussion address a direct or indirect link between Kinesins and ribbon bodies. Showing Kif1aa protein in proximity to the ribbon bodies would add strength.
This is a great point. Previous immunohistochemistry work in mice demonstrated that ribbons and Kif1a colocalize in mouse hair cells (Michanski et al, 2019). Unfortunately, the antibody used in study work did not work in zebrafish. To further investigate this interaction, we also attempted to create a transgenic line expressing a fluorescently tagged Kif1aa to directly visualize its association with ribbons in vivo. At present, we were unable to detect transient expression of Kif1aa-GFP or establish a transgenic line using this approach. While we will continue to work towards understanding whether Kif1aa and ribbons colocalize in live hair cells, currently this goal is beyond the scope of this paper. In our revision we discuss this caveat.
Added to discussion:
“In addition, it will be useful to visualize these kinesins by fluorescently tagging them in live hair cells to observe whether they associate with ribbons.”
(2) Neither the data or Discussion address the functional consequences of loss of Kif1aa or ribbon transport. Presumably, both manipulations would reduce afferent excitation.
Excellent point. Please see the response above to Reviewer #1 public response weaknesses.
(3) It is unknown whether the drug treatments or genetic manipulations are specific to hair cells, so we can't know for certain whether any phenotypic defects are secondary.
This is correct and a caveat of our Kif1aa and drug experiments. In our recently published work, we confirmed that Kif1aa is expressed in hair cells and neurons, while kif1ab is present just is neurons. Therefore, it is likely that the ribbon formation defects in kif1aa mutants are restricted to hair cells. We added this expression information to our results:
“ScRNA-seq in zebrafish has demonstrated widespread co-expression of kif1ab and kif1aa mRNA in the nervous system. Additionally, both scRNA-seq and fluorescent in situ hybridization have revealed that pLL hair cells exclusively express kif1aa mRNA (David et al., 2024; Lush et al., 2019; Sur et al., 2023).”
Non-hair cell effects are a real concern in our pharmacology experiments. To mitigate this in our pharmacological experiments, we have performed drug treatments at 3 different timescales: long-term (overnight), short-term (4 hr) and fast (30 min) treatments. The fast experiments were done after 30 min nocodazole drug treatment, and after this treatment we observed reduced directional motion and fusions. This fast drug treatment should not incur any long-term changes or developmental defects as hair-cell development occurs over 12-16 hrs. However, we acknowledge that drug treatments could have secondary phenotypic effects or effects that are not hair-cell specific. In our revision, we discuss these issues.
Added to discussion:
“Another important consideration is the potential off-target effects of nocodazole. Even at non-cytotoxic doses, nocodazole toxicity may impact ribbons and synapses independently of its effects on microtubules. While this is less of a concern in the short- and medium-term experiments (30-70 min and 4 hr), long-term treatments (16 hrs) could introduce confounding effects. Additionally, nocodazole treatment is not hair cell-specific and could disrupt microtubule organization within afferent terminals as well. Thus, the reduction in ribbon-synapse formation following prolonged nocodazole treatment may result from microtubule disruption in hair cells, afferent terminals, or a combination of the two.”
Reviewer #3 (Public Review):
Summary:
The manuscript uses live imaging to study the role of microtubules in the movement of ribeye aggregates in neuromast hair cells in zebrafish. The main findings are that
(1) Ribeye aggregates, assumed to be ribbon precursors, move in a directed motion toward the active zone;
(2) Disruption of microtubules and kif1aa increases the number of ribeye aggregates and decreases the number of mature synapses.
The evidence for point 2 is compelling, while the evidence for point 1 is less convincing. In particular, the directed motion conclusion is dependent upon fitting of mean squared displacement that can be prone to error and variance to do stochasticity, which is not accounted for in the analysis. Only a small subset of the aggregates meet this criteria and one wonders whether the focus on this subset misses the bigger picture of what is happening with the majority of spots.
Strengths:
(1) The effects of Kif1aa removal and nocodozole on ribbon precursor number and size are convincing and novel.
(2) The live imaging of Ribeye aggregate dynamics provides interesting insight into ribbon formation. The movies showing the fusion of ribeye spots are convincing and the demonstrated effects of nocodozole and kif1aa removal on the frequency of these events is novel.
(3) The effect of nocodozole and kif1aa removal on precursor fusion is novel and interesting.
(4) The quality of the data is extremely high and the results are interesting.
Weaknesses:
(1) To image ribeye aggregates, the investigators overexpressed Ribeye-a TAGRFP under the control of a MyoVI promoter. While it is understandable why they chose to do the experiments this way, expression is not under the same transcriptional regulation as the native protein, and some caution is warranted in drawing some conclusions. For example, the reduction in the number of puncta with maturity may partially reflect the regulation of the MyoVI promoter with hair cell maturity. Similarly, it is unknown whether overexpression has the potential to saturate binding sites (for example motors), which could influence mobility.
We agree that overexpression of transgenes under using a non-endogenous promoter in transgenic lines is an important consideration. Ideally, we would do these experiments with endogenously expressed fluorescent proteins under a native promoter. However, this was not technically possible for us. The decrease in precursors is likely not due to regulation by the myo6a promoter. Although the myo6a promoter comes on early in hair cell development, the promoter only gets stronger as the hair cells mature. This would lead to a continued increase rather than a decrease in puncta numbers with development.
Protein tags such as tagRFP always have the caveat of impacting protein function. This is in partly why we complemented our live imaging with analyses in fixed tissue without transgenes (kif1aa mutants and nocodazole/taxol treatments).
In our revision, we did perform an immunolabel on myo6b:riba-tagRFP transgenic fish and found that Riba-tagRFP expression did not impact ribbon synapse numbers or ribbon size. This analysis argues that the transgene is expressed at a level that does not impact ribbon synapses. This data is summarized in Figure 1-S1.
Added to the results:
“Although this latter transgene expresses Riba-TagRFP under a non-endogenous promoter, neither the tag nor the promoter ultimately impacts cell numbers, synapse counts, or ribbon size (Figure 1-S1A-E).”
Added to methods:
“Tg(myo6b:ctbp2a-TagRFP)<sup>idc11Tg</sup> reliably labels mature ribbons, similar to a pan-CTBP immunolabel at 5 dpf (Figure 1-S1B). This transgenic line does not alter the number of hair cells or complete synapses per hair cell (Figure 1-S1A-D). In addition, myo6b:ctbp2a-TagRFP does not alter the size of ribbons (Figure 1-S1E).”
(2) The examples of punctae colocalizing with microtubules look clear (Figures 1 F-G), but the presentation is anecdotal. It would be better and more informative, if quantified.
We did attempt a co-localization analysis between microtubules and ribbons but did not move forward with it due to several issues:
(1) Hair cells have an extremely crowded environment, especially since the nucleus occupies the majority of the cell. All proteins are pushed together in the small space surrounding the nucleus and ultimately, we found that co-localization analyses were not meaningful because the distances were too small.
(2) We also attempted to segment microtubules in these images and quantify how many ribbons were associated with microtubules, but 3D microtubule segmentation was not accurate in hair cells due to highly varying filament intensities, filament dynamics and the presence of diffuse cytoplasmic tubulin signal.
Because of these challenges we concluded the best evidence of ribbon-microtubule association is through visualization of ribbons and their association with microtubules over time (in our timelapses). We see that ribbons localize to microtubules in all our timelapses, including the examples shown (Movies S2-S10). The only instance of ribbon dissociation it when ribbons switch from one filament to another. We did not observe free-floating ribbons in our study.
(3) It appears that any directed transport may be rare. Simply having an alpha >1 is not sufficient to declare movement to be directed (motor-driven transport typically has an alpha approaching 2). Due to the randomness of a random walk and errors in fits in imperfect data will yield some spread in movement driven by Brownian motion. Many of the tracks in Figure 3H look as though they might be reasonably fit by a straight line (i.e. alpha = 1).
(4) The "directed motion" shown here does not really resemble motor-driven transport observed in other systems (axonal transport, for example) even in the subset that has been picked out as examples here. While the role of microtubules and kif1aa in synapse maturation is strong, it seems likely that this role may be something non-canonical (which would be interesting).
Yes, it is true, that directed transport of ribbon precursors is relatively rare. Only a small subset of the ribbon precursors moves directionally (α > 1, 20 %) or have a displacement distance > 1 µm (36 %) during the time windows we are imaging. The majority of the ribbons are stationary. To emphasize this result we have added bar graphs to Figure 3I,K to illustrate this result and state the numbers behind this result more clearly.
“Upon quantification, 20.2 % of ribbon tracks show α > 1, indicative of directional motion, but the majority of ribbon tracks (79.8 %) show α < 1, indicating confinement on microtubules (Figure 3I, n = 10 neuromasts, 40 hair cells, and 203 tracks).
To provide a more comprehensive analysis of precursor movement, we also examined displacement distance (Figure 3J). Here, as an additional measure of directed motion, we calculated the percent of tracks with a cumulative displacement > 1 µm. We found 35.6 % of tracks had a displacement > 1 µm (Figure 3K; n = 10 neuromasts, 40 hair cells, and 203 tracks).”
We cannot say for certain what is happening with the stationary ribbons, but our hypothesis is that these ribbons eventually exhibit directed motion sufficient to reach the active zone. This idea is supported by the fact that we see ribbons that are stationary begin movement, and ribbons that are moving come to a stop during the acquisition of our timelapses (Movies S4 and S5). It is possible that ribbons that are stationary may not have enough motors attached, or there may be a ‘seeding’ phase where Ribeye aggregates are condensing on the ribbon.
We also reexamined our MSD a values as the a values we observed in hair cells were lower than those seen canonical motor-driven transport (where a approaches 2). One reason for this difference may arise from the dynamic microtubule network in developing hair cells, which could affect directional ribbon movement. In our revision we plotted the distribution of a values which confirmed that in control hair cells, the majority of the a values we see are typically less than 2 (Figure 7-S1A). Interestingly we also compared the distribution a values between control and taxol-treated hair cells, where the microtubule network is more stable, and found that the distribution shifted towards higher a values (Figure 7-S1A). We also plotted only ‘directional’ tracks (with a > 1) and observed significantly higher a values in taxol-treated hair cells (Figure 7-S1B). This is an interesting result which indicates that although the proportion of directional tracks (with a > 1) is not significantly different between control and taxol-treated hair cells (which could be limited by the number of motor/adapter proteins), the ribbons that move directionally do so with greater velocities when the microtubules are more stable. This supports our idea that the stability of the microtubule network could be why ribbon movement does not resemble canonical motor transport. This analysis is presented as a new figure (Figure 7-S1A-B) and is referred to in the text in the results and the discussion.
Results:
“Interestingly, when we examined the distribution of α values, we observed that taxol treatment shifted the overall distribution towards higher α a values (Figure 7-S1A). In addition, when we plotted only tracks with directional motion (α > 1), we found significantly higher α values in hair cells treated with taxol compared to controls (Figure 7-S1B). This indicates that in taxol-treated hair cells, where the microtubule network is stabilized, ribbons with directional motion have higher velocities.”
Discussion:
“Our findings indicate that ribbons and precursors show directed motion indicative of motor-mediated transport (Figure 3 and 7). While a subset of ribbons moves directionally with α values > 1, canonical motor-driven transport in other systems, such as axonal transport, can achieve even higher α values approaching 2 (Bellotti et al., 2021; Corradi et al., 2020). We suggest that relatively lower α values arise from the highly dynamic nature of microtubules in hair cells. In axons, microtubules form stable, linear tracks that allow kinesins to transport cargo with high velocity. In contrast, the microtubule network in hair cells is highly dynamic, particularly near the cell base. Within a single time frame (50-100 s), we observe continuous movement and branching of these networks. This dynamic behavior adds complexity to ribbon motion, leading to frequent stalling, filament switching, and reversals in direction. As a result, ribbon transport appears less directional than the movement of traditional motor cargoes along stable axonal filaments, resulting in lower α values compared to canonical motor-mediated transport. Notably, treatment with taxol, which stabilizes microtubules, increased α values to levels closer to those observed in canonical motor-driven transport (Figure 7-S1). This finding supports the idea that the relatively lower α values in hair cells are a consequence of a more dynamic microtubule network. Overall, this dynamic network gives rise to a slower, non-canonical mode of transport.”
(5) The effect of acute treatment with nocodozole on microtubules in movie 7 and Figure 6 is not obvious to me and it is clear that whatever effect it has on microtubules is incomplete.
When using nocodazole, we worked to optimize the concentration of the drug to minimize cytotoxicity, while still being effective. While the more stable filaments at the cell apex remain largely intact after nocodazole treatment, there are almost no filaments at the hair cell base, which is different from the wild-type hair cells. In addition, nocodazole-treated hair cells have more cytoplasmic YFP-tubulin signal compared to wild type. We have clarified this in our results. To better illustrate the effect of nocodazole and taxol we have also added additional side-view images of hair cells expressing YFP-tubulin (Figure 4-S1F-G), that highlight cytoplasmic YFP-tubulin and long, stabilized microtubules after 3-4 hr treatment with nocodazole and taxol respectively. In these images we also point out microtubules at the apical region of hair cells that are very stable and do not completely destabilize with nocodazole treatment at concentrations that are tolerable to hair cells.
“We verified the effectiveness of our in vivo pharmacological treatments using either 500 nM nocodazole or 25 µM taxol by imaging microtubule dynamics in pLL hair cells (myo6b:YFP-tubulin). After a 30-min pharmacological treatment, we used Airyscan confocal microscopy to acquire timelapses of YFP-tubulin (3 µm z-stacks, every 50-100 s for 30-70 min, Movie S8). Compared to controls, 500 nM nocodazole destabilized microtubules (presence of depolymerized YFP-tubulin in the cytosol, see arrows in Figure 4-S1F-G) and 25 µM taxol dramatically stabilized microtubules (indicated by long, rigid microtubules, see arrowheads in Figure 4-S1F,H) in pLL hair cells. We did still observe a subset of apical microtubules after nocodazole treatment, indicating that this population is particularly stable (see asterisks in Figure 4-S1F-H).”
To further address concerns about verifying the efficacy of nocodazole and taxol treatment on microtubules, we added a quantification of our immunostaining data comparing the mean acetylated-a-tubulin intensities between control, nocodazole and taxol-treated hair cells. Our results show that nocodazole treatment reduces the mean acetylated-a-tubulin intensity in hair cells. This is included as a new figure (Figure 4-S1D-E) and this result is referred to in the text. To better illustrate the effect of nocodazole and taxol we have also added additional side-view images of hair cells after overnight treatment with nocodazole and taxol (Figure 4-S1A-C).
“After a 16-hr treatment with 250 nM nocodazole we observed a decrease in acetylated-a-tubulin label (qualitative examples: Figure 4A,C, Figure 4-S1A-B). Quantification revealed significantly less mean acetylated-a-tubulin label in hair cells after nocodazole treatment (Figure 4-S1D). Less acetylated-a-tubulin label indicates that our nocodazole treatment successfully destabilized microtubules.”
“Qualitatively more acetylated-a-tubulin label was observed after treatment, indicating that our taxol treatment successfully stabilized microtubules (qualitative examples: Figure 4-S1A,C). Quantification revealed an overall increase in mean acetylated-a-tubulin label in hair cells after taxol treatment, but this increase did not reach significance (Figure 4-S1E).”
Recommendations for the authors:
Reviewer #1 (Recommendations For The Authors):
(1) The manuscript is fairly dense. For instance, some information is repeated (page 3 ribbon synapses form along a condensed timeline in zebrafish hair cells: 12-18 hrs, and on .page 5. These hair cells form 3-4 ribbon synapses in just 12-18 hrs). Perhaps, the authors could condense some of the ideas? The introduction could be shortened.
We have eliminated this repeated text in our revision. We have shortened the introduction 1275 to 1038 words (with references)
(2) The mechanosensory structure on page 5 is not defined for readers outside the field.
Great point, we have added addition information to define this structure in the results:
“We staged hair cells based on the development of the apical, mechanosensory hair bundle. The hair bundle is composed of actin-based stereocilia and a tubulin-based kinocilium. We used the height of the kinocilium (see schematic in Figure 1B), the tallest part of the hair bundle, to estimate the developmental stage of hair cells as described previously…”
(3) Figure 1E is quite interesting but I'd rather show Figure S1 B/C as they provide statistics. In addition, the authors define 4 stages : early, intermediate, late, and mature for counting but provide only 3 panels for representative examples by mixing late/mature.
We were torn about which ribbon quantification graph to show. Ultimately, we decided to keep the summary data in Figure 1E. This is primarily because the supplementary Figure will be adjacent to the main Figure in the Elife format, and the statistics will be easy to find and view.
Figure 1 now provides a representative image for both late and mature hair cells.
(4.) The ribbon that jumps from one microtubule to another one is eye-catching. Can the authors provide any statistics on this (e.g. percentage)?
Good point. In our revision, we have added quantification for these events. We observe 2.8 switching events per neuromast during our fast timelapses. This information is now in the text and is also shown in a graph in Figure 3-S1D.
“Third, we often observed that precursors switched association between neighboring microtubules (2.8 switching events per neuromast, n= 10 neuromasts; Figure 3-S1C-D, Movie S7).”
(5) With regard to acetyl-a-tub immunocytochemistry, I would suggest obtaining a profile of the fluorescence intensity on a horizontal plane (at the apical part and at the base).
(6) Same issue with microtubule destruction by nocodazole. Can the authors provide fluorescence intensity measurements to convince readers of microtubule disruption for long and short-term application.
Regarding quantification of microtubule disruption using nocodazole and taxol. We did attempt to create profiles of the acetylated tubulin or YFP-tubulin label along horizontal planes at the apex and base, but the amount variability among cells and the angle of the cell in the images made this type of display and quantification challenging. In our revision we as stated above in our response to Reviewer #1’s public comment, we have added representative side-view images to show the disruptions to microtubules more clearly after short and long-term drug experiments (Figure 4-S1A-C, F-H). In addition, we quantified the reduction in acetylated tubulin label after overnight treatment with nocodazole and found the signal was significantly reduced (Figure 3-S1D-E). Unfortunately, we were unable to do a similar quantification due to the variability in YFP-tubulin intensity due to variations in mounting. The following text has been added to the results:
“Quantification revealed significantly less mean acetylated-a-tubulin label in hair cells after nocodazole treatment (Figure 4-S1D).”
“Quantification revealed an overall increase in mean acetylated-a-tubulin label in hair cells after taxol treatment, but this increase did not reach significance (Figure 4-S1A,C,E).”
(7) It is a bit difficult to understand that the long-term (overnight) microtubule destabilization leads to a reduction in the number of synapses (Figure 4F) whereas short-term (30 min) microtubule destabilization leads to the opposite phenotype with an increased number of ribbons (Figure 6G). Are these ribbons still synaptic in short-term experiments? What is the size of the ribbons in the short-term experiments? Alternatively, could the reduction in synapse number upon long-term application of nocodazole be a side-effect of the toxicity within the hair cell?
Agreed-this is a bit confusing. In our revision, we have changed our analyses, so the comparisons are more similar between the short- and long-term experiments–we examined the number of ribbons and precursor per cells (apical and basal) in both experiments (Changed the panel in Figure 4G, Figure 4-S2G and Figure 5G). In our live experiments we cannot be sure that ribbons are synaptic as we do not have a postsynaptic co-label. Also, we are unable to reliably quantify ribbon and precursor size in our live images due to variability in mounting. We have changed the text to clarify as follows:
Results:
“In each developing cell, we quantified the total number of Riba-TagRFP puncta (apical and basal) before and after each treatment. In our control samples we observed on average no change in the number of Riba-TagRFP puncta per cell (Figure 6G). Interestingly, we observed that nocodazole treatment led to a significant increase in the total number of Riba-TagRFP puncta after 3-4 hrs (Figure 6G). This result is similar to our overnight nocodazole experiments in fixed samples, where we also observed an increase in the number of ribbons and precursors per hair cell. In contrast to our 3-4 hr nocodazole treatment, similar to controls, taxol treatment did not alter the total number of Riba-TagRFP puncta over 3-4 hrs (Figure 6G). Overall, our overnight and 3-4 hr pharmacology experiments demonstrate that microtubule destabilization has a more significant impact on ribbon numbers compared to microtubule stabilization.”
Discussion:
“Ribbons and microtubules may interact during development to promote fusion, to form larger ribbons. Disrupting microtubules could interfere with this process, preventing ribbon maturation. Consistent with this, short-term (3-4 hr) and long-term (overnight) nocodazole increased ribbon and precursor numbers (Figure 6AG; Figure 4G), suggesting reduced fusion. Long-term treatment (overnight) resulted in a shift toward smaller ribbons (Figure 4H-I), and ultimately fewer complete synapses (Figure 4F).”
Nocodazole toxicity: in response to Reviewer # 2’s public comment we have added the following text in our discussion:
Discussion:
“Another important consideration is the potential off-target effects of nocodazole. Even at non-cytotoxic doses, nocodazole toxicity may impact ribbons and synapses independently of its effects on microtubules. While this is less of a concern in the short- and medium-term experiments (30 min to 4 hr), long-term treatments (16 hrs) could introduce confounding effects. Additionally, nocodazole treatment is not hair cell-specific and could disrupt microtubule organization within afferent terminals as well. Thus, the reduction in ribbon-synapse formation following prolonged nocodazole treatment may result from microtubule disruption in hair cells, afferent terminals, or a combination of the two.”
(8) Does ribbon motion depend on size or location?
It is challenging to reliability quantify the actual area of precursors in our live samples, as there is variability in mounting and precursors are quite small. But we did examine the location of ribbon precursors (using tracks > 1 µm as these tracks can easily be linked to cell location in Imaris) with motion in the cell. We found evidence of ribbons with tracks > 1 µm throughout the cell, both above and below the nucleus. This is now plotted in Figure 3M. We have also added the following test to the results:
“In addition, we examined the location of precursors within the cell that exhibited displacements > 1 µm. We found that 38.9 % of these tracks were located above the nucleus, while 61.1 % were located below the nucleus (Figure 3M).”
Although this is not an area or size measurement, this result suggests that both smaller precursors that are more apical, and larger precursors/ribbons that are more basal all show motion.
(9) The fusion event needs to be analyzed in further detail: when one ribbon precursor fuses with another one, is there an increase in size or intensity (this should follow the law of mass conservation)? This is important to support the abstract sentence "ribbon precursors can fuse together on microtubules to form larger ribbons".
As mentioned above it is challenging accurately estimate the absolute size or intensity of ribbon precursors in our live preparation. But we did examine whether there is a relative increase in area after ribbon fuse. We have plotted the change in area (within the same samples) for the two fusion events in shown in Figure 8-S1A-B. In these examples, the area of the puncta after fusion is larger than either of the two precursors that fuse. Although the areas are not additive, these plots do provide some evidence that fusion does act to form larger ribbons. To accompany these plots, we have added the following text to the results:
“Although we could not accurately measure the areas of precursors before and after fusion, we observed that the relative area resulting from the fusion of two smaller precursors was greater than that of either precursor alone. This increase in area suggests that precursor fusion may serve as a mechanism for generating larger ribbons (see examples: Figure 8-S1A-B).”
Because we were unable to provide more accurate evidence of precursor fusion resulting in larger ribbons, we have removed this statement from our abstract and lessened our claims elsewhere in the manuscript.
(10) The title in Figure 8 is a bit confusing. If fusion events reflect ribbon precursors fusion, it is obvious it depends on ribbon precursors. I'd like to replace this title with something like "microtubules and kif1aa are required for fusion events"
We have changed the figure title as suggested, good idea.
Reviewer #2 (Recommendations For The Authors):
(1) Figure 1C. The purple/magenta colors are hard to distinguish.
We have made the magenta color much lighter in the Figure 1C to make it easier to distinguish purple and magenta.
(2) There are places where some words are unnecessarily hyphenated. Examples: live-imaging and hair-cell in the abstract, time-course in the results.
In our revision, we have done our best to remove unnecessary hyphens, including the ones pointed out here.
(3) Figure 4H and elsewhere - what is "area of Ribeye puncta?" Related, I think, in the Discussion the authors refer to "ribbon volume" on line 484. But they never measured ribbon volume so this needs to be clarified.
We have done best to clarify what is meant by area of Ribeye puncta in the results and the methods:
Results:
“We also observed that the average of individual Ribeyeb puncta (from 2D max-projected images) was significantly reduced compared to controls (Figure 4H). Further, the relative frequency of individual Ribeyeb puncta with smaller areas was higher in nocodazole treated hair cells compared to controls (Figure 4I).”
Methods:
“To quantify the area of each ribbon and precursor, images were processed in a FIJI ‘IJMacro_AIRYSCAN_simple3dSeg_ribbons only.ijm’ as previously described (Wong et al., 2019). Here each Airyscan z-stack was max-projected. A threshold was applied to each image, followed by segmentation to delineate individual Ribeyeb/CTBP puncta. The watershed function was used to separate adjacent puncta. A list of 2D objects of individual ROIs (minimum size filter of 0.002 μm2) was created to measure the 2D areas of each Ribeyeb/CTBP puncta.”
We did refer to ribbon volume once in the discussion, but volume is not reflected in our analyses, so we have removed this mention of volume.
(4) More validation data showing gene/protein removal for the crispants would be helpful.
Great suggestion. As this is a relatively new method, we have created a figure that outlines how we genotype each individual crispant animal analyzed in our study Figure 6-S1. In the methods we have also added the following information:
“fPCR fragments were run on a genetic analyzer (Applied Biosystems, 3500XL) using LIZ500 (Applied Biosystems, 4322682) as a dye standard. Analysis of this fPCR revealed an average peak height of 4740 a.u. in wild type, and an average peak height of 126 a.u. in kif1aa F0 crispants (Figure 6-S1). Any kif1aa F0 crispant without robust genomic cutting or a peak height > 500 a.u. was not included in our analyses.”
Reviewer #3 (Recommendations For The Authors):
Lines 208-209--should refer to the movie in the text.
Movie S1 is now referenced here.
It would be helpful if the authors could analyze and quantify the effect of nocodozole and taxol on microtubules (movie 7).
See responses above to Reviewer #1’s similar request.
Figure 7 caption says "500 mM" nocodozole.
Thank you, we have changed the caption to 500 nM.
One problem with the MSD analysis is that it is dependent upon fits of individual tracks that lead to inaccuracies in assigning diffusive, restricted, and directed motion. The authors might be able to get around these problems by looking at the ensemble averages of all the tracks and seeing how they change with the various treatments. Even if the effect is on a subset of ribeye spots, it would be reassuring to see significant effects that did not rely upon fitting.
We are hesitant to average the MSD tracks as not all tracks have the same number of time steps (ribbon moving in and out of the z-stack during the timelapse). This makes it challenging for us to look at the ensembles of all averages accurately, especially for the duration of the timelapse. This is the main reason why added another analysis, displacements > 1µm as another readout of directional motion, a measure that does not rely upon fitting.
The abstract states that directed movement is toward the synapse. The only real evidence for this is a statement in the results: "Of the tracks that showed directional motion, while the majority move to the cell base, we found that 21.2 % of ribbon tracks moved apically." A clearer demonstration of this would be to do the analysis of Figure 2G for the ribeye aggregates.
If was not possible to do the same analysis to ribbon tracks that we did for the EB3-GFP analysis in Figure 2. In Figure 2 we did a 2D tracking analysis and measured the relative angles in 2D. In contrast, the ribbon tracking was done in 3D in Imaris not possible to get angles in the same way. Further the MSD analysis was outside of Imaris, making it extremely difficult to link ribbon trajectories to the 3D cellular landscape in Imaris. Instead, we examined the direction of the 3D vectors in Imaris with tracks > 1µm and determined the direction of the motion (apical, basal or undetermined). For clarity, this data is now included as a bar graph in Figure 3L. In our results, we have clarified the results of this analysis:
“To provide a more comprehensive analysis of precursor movement, we also examined displacement distance (Figure 3J). Here, as an additional measure of directed motion, we calculated the percent of tracks with a cumulative displacement > 1 µm. We found 35.6 % of tracks had a displacement > 1 µm (Figure 3K; n = 10 neuromasts, 40 hair cells and 203 tracks). Of the tracks with displacement > 1 µm, the majority of ribbon tracks (45.8 %) moved to the cell base, but we also found a subset of ribbon tracks (20.8 %) that moved apically (33.4 % moved in an undetermined direction) (Figure 3L).”
Some more detail about the F0 crispants should be provided. In particular, what degree of cutting was observed and what was the criteria for robust cutting?
See our response to Reviewer 2 and the newly created Figure 6-S1.
RRID:CVCL_1092
DOI: 10.1016/j.lungcan.2025.108611
Resource: (NCI-DTP Cat# BT-549, RRID:CVCL_1092)
Curator: @scibot
SciCrunch record: RRID:CVCL_1092
NOMBRANDO LAS COLUMNAS VISTA TREE
Después de colocar los nombres a las columnas, así se ve en la vista DATA y en la vista TREE.
po
Así la ejecuta PHARO , aquí todavía no se le asigna los nombres a las columnas
pokemonTournament := DataFrame withColumns: #((ROUND1 ROUND2 ROUND3 ROUND4 ROUND5) (Pikachu No No No No) (Ditto No No Ditto No) (No Balbasaur No No No) (No Charmander No Charmander Charmander) (No No Charizard No No) (No No Squirtle No Squirtle) (Ditto Charmander Squirtle Charmander Charmander)).
Agregue los valores específicos de cada columna, pero sin darle nombre a la columna
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
*We would like to thank all the reviewers for their positive comments and valuable feedback. In addition, we would like to address reviewer 1 query on novelty, which was not questioned by the other 2 reviewers. Our study uncovered two main aspects of hypoxia biology: first we addressed the role of NF-kappaB contribution towards the transcriptome changes in hypoxia, and second, this revealed a previously unknown aspect, that NF-kappaB is required for gene repression in hypoxia. While we know a lot about gene induction in hypoxia, much less is known about repression of genes. In times of energy preservation, gene repression is as important as gene induction. *
.
This section is mandatory. *Please insert a point-by-point reply describing the revisions that were already carried out and included in the transferred manuscript. *
Reviewer #1 (Evidence, reproducibility and clarity (Required)):
The work from Shakir et al uses different cell line models to investigate the role of NF-kB in the transcriptional adaptation of cells to hypoxia, which is relevant. In addition, the manuscript contains a large amount of data that could be of interest and even useful for researchers in the field of hypoxia and NF-kB. However, in my opinion, there are several concerns that should be revised and additional experiments that could be included to strengthen the relevance of the work.
We thank this reviewer for their positive comments.
Specific issues: In Figure 1A, the authors examine which of the genes induced by hypoxia require NF-kB by RNA sequencing analysis of cells knocked down for specific NF-kB subunits and exposed to hypoxia for 24 hours. The knockdown is about 40-60% at the RNA level, but it would be helpful to show the effect of knockdown at the protein level.
We agree with this and have added Western blot data (Sup. Figure S1F), which shows the effects of the siRNA are much more pronounced at the protein level.
All the data regarding genes induced by hypoxia in control or NF-kB siRNA-treated cells are somewhat confusing. If I understand correctly, when the data from the three different siRNAs are crossed, only 1070 genes are upregulated and 295 are downregulated in an NF-kB-independent manner. If this is the case, I think it would be easier to use this information in Figure 2 to define the hypoxia-induced genes that are NF-kB-dependent by simply considering those induced in the control that are not in the NF-kB-independent subset (rather than repeating the integration of the data without additional explanation). If the authors do this simple analysis, are the resulting genes the same or similar? In any case, the way these numbers are obtained should be shown more clearly (i.e., a new Venn diagram showing genes up- or down-regulated in the siRNA control that are not up- or down-regulated in any of the siRNA-NF-kB treatments).
Figure 1 shows the effects on gene expression of hypoxia in control and NF-____k____B ____subunit____-depleted cells compared to normoxia control cells. Figures 1F/1G compares genes up/downregulated in hypoxia when RelA, RelB, and cRel are depleted, compared to normoxia control. Figure 1 does not display N____F-____k____B____-dependent/independent hypoxia-responsive genes____, but rather the overall effect of siRNA control and siNF-____k____B treatments in hypoxia, compared to siRNA control in normoxia. Figure 2 then defines NF-____k____B-dependent ____and independent hypoxia-responsive genes. We actually define these exactly as the reviewer suggested and agree that we should show the way these numbers are obtained more clearly. We have added the suggested Venn diagrams (Sup. Figure S2) and added extra information to the methods section (page 5 of revised manuscript). We felt it was important to show all the data upfront in Figure 1 and then integrate and focus on NF-____k____B-dependent ____hypoxia-induced genes in Figure 2.
Figure 2H shows that approximately 80% of the NF-kB-dependent genes up- or down-regulated in hypoxia were identified as RelA targets, which is statistically significant compared to RelB or cRel targets. However, what is the proportion of genes identified as RelA targets in the subset of NF-kB-independent hypoxia-induced genes? And in a randomly selected set of 500-600 genes? In my opinion, this statistical analysis should be included to demonstrate a relationship between NF-kB recruitment and hypoxia-induced upregulation (expected) and downregulation (unexpected). In this context, it is surprising that HIF consensus sites are preferentially detected in the genes that are supposed to be NF-kB dependent instead of RelA consensus.
We thank the reviewer for this question, which is really helpful. The way we have displayed the stars on the graph for Figure 2H was slightly misleading we realize now. As such, we have amended the graph. RelA, RelB, and cRel bound genes (from the ChIP atlas) are all significantly enriched within our N____F-____k____B-dependent hypoxia-responsive genes, there is no statistical testing between RelA bound vs RelB bound or cRel bound. We have also performed this analysis on the NF-____k____B____-independent hypoxia-responsive genes ____and see the same trend (Sup. Figure S5B). This indicates that the enrichment of Rel binding sites from the ChIP atlas is not specific to NF-____k____B____-dependent hypoxia-responsive genes____. We have moved Figure 2H to (Sup. Figure S5A) and amended our description of the finding. This showcases how DNA binding does not necessarily mean functionality. We have amended our description of this result and limitation of the study.
Figure 3 is just a confirmation by qPCR of the data obtained in the RNA-seq analysis, which in my opinion should be included as supplementary information. Moreover, both the effects of hypoxia and reversion by RelB siRNA are modest in several of the genes tested. The same is true for Figures 4 and 5 with very modest and variable results across cell types and genes.
We appreciate this comment; we would like to keep this as a main figure for full transparency and show validation of our RNA-sequencing results.
Figure 6 shows the effect of NF-kB knockdown on the induction of ROS in response to hypoxia. In the images provided, the effect of hypoxia is minimal in control cells, with the only clear differences shown in RelA-depleted cells.
The quantification of the IF data (Figure 6B) shows ROS induction in hypoxia which is reduced in Rel-depleted cells, with RelA depletion having the strongest effect. ROS generation in hypoxia, although counterintuitive, is well documented and used for important signalling events. We believe our data supports the previously reported levels of ROS induction (reviewed in {Alva, 2024}) in hypoxia and importantly, that NF-____k____B depletion can at least partially____ reverse this.
In 6B it is not clear what the three asterisks in the normoxia control represent (compared to the hypoxia siRNA control?). This should be clarified in the figure legend or text.
We apologize for the lack of clarity we have now added this information to the figure legend.
In the Western blot of 6C, there are no differences in the levels of SOD1 after RelA depletion. Again, there is no reason not to include the NF-kB subunits in the Western blot analysis.
We have added the Western blot analysis to this figure. We were trying to simplify it. Although depletion of RelA does not rescue the hypoxia-induced repression of SOD1, depletion of RelB does. Furthermore, cRel although not statistically significant, has a trend for the rescue of this effect, see Figure 6C-D.
Finally, regarding Figure 7, the authors mention that "we confirmed that hypoxia led to a reduction in several proteins represented in this panel (of proteins involved in oxidative phosphorylation), such as UQCRC2 and IDH1 (Figure 7A-B)". The authors cannot say this because it is not seen in the Western blot in 7A or in the quantification shown in 7B. In my personal opinion, stating something that is not even suggested in the experiments is very negative for the credibility of the whole message.
We really do not agree with this comment. We do see reductions in the levels of the proteins we mentioned. We have made the figure less complex given that some proteins are very abundant while others are not. We hope the changes are now clear and apparent. We have changed the quantification normalisation to reflect this as well and modified our description of the results, see Figure 7 and Sup. Figure S18.
In conclusion, this paper contains a large amount of relevant information, but i) non-essential data should be moved to Supplementary, ii) protein levels of relevant players need to be shown in addition to RNA, iii) minimal or undetectable differences need to be considered as no-differences, and iv) a model showing what is the interpretation of the data provided is needed to better understand the message of the paper. I mean, is it p65 or RelB binding to some of these genes leading to their activation or repression, or is it RelA or RelB inducing HIF1beta leading to NF-kB-dependent gene activation by hypoxia? If this were the case, experimental evidence that NF-kB regulates a subset of hypoxia genes through HIF1beta would make the story more understandable.
We apologise but we do not know why the reviewer mentions HIF1beta. For gene induction, there is cooperation with the HIF system in some genes but not all. The most interesting and unexpected finding is that NF-kappaB is required for gene repression in hypoxia. We have added a new figure, investigating how HDAC inhibition could reverse the repression. A mechanism known to be employed by NF-kappaB when repressing genes. We have added all the blots for NF-kB, clarified the quantification and included other approaches including a CRISPR KO cell lines for both IKKs. We hope this is now clear.
Reviewer #1 (Significance (Required)):
The work presented here is interesting but does not provide a major advance over previous publications, the main message being that a subset of hypoxia-regulated genes are NF-kB dependent. However, there is no mechanistic explanation of how this regulation is achieved and several data that are not clearly connected. A more comprehensive analysis of the data and additional experimental validation would greatly enhance the significance of the work.
We politely disagree with the reviewer. Our main finding is that NF-____k____B____ does play an important role in gene regulation in hypoxia but unexpectedly, this occurs mostly via gene repression. While there is vast knowledge on gene induction in hypoxia, gene repression, which typically does not occur directly via HIF, is virtually unknown. A previous study had identified Rest as a transcriptional repressor {PMID: 27531581} but this could only account for 20% of gene repression. Our findings reveal up to 60% of genes repressed in hypoxia require NF-____k____B____, hence this is a significant finding and a major advance over previous knowledge. Furthermore, we feel this paper is an excellent data resource for the field, as it is, to our knowledge, the first study characterising the extent to which NF-____k____B is required for hypoxia-induced gene changes, on a transcriptome-wide scale. Furthermore, we have validated this across multiple cell types and also used different approaches to investigate the role of NF-kB in the hypoxia transcriptional response. We are happy that the other reviewers agree with our novel findings.
Reviewer #2 (Evidence, reproducibility and clarity (Required)):
In this study, the authors have interrogated the role of NF-kappaB in the cellular transcriptional response to hypoxia. While HIF is considered the master regulator of the cellular response to hypoxia, it has long been known that mutliple transcription factors also play a role both independently of HIF and through the regulation of HIF-1alpha levels. Chief amongst these is NF-kappaB, a regulator of cell death and inflammation amongst other things. While NF-kappB has been known to be activated in hypoxia through altered PhD activity, the impact of this on global gene expression has remained unclear and this study addresses this important question. Of particular interest, genes downregulated in hypoxia appear to be repressed in a NF-kappaB-dependent manner. Overall, this nice study reveals an important role for NF-kappaB in the control of the global cellular transcriptional response to hypoxia.
We thank this reviewer for their positive comments.
Reviewer #2 (Significance (Required)):
Some questions for the authors to consider with experiments or discussion: -One caveat of the current study which should be discussed is that while interesting and extensive, the analysis is restricted to cancer cell lines which have dysfunctional gene expression systems which may differ from "normal" cells. This should be discussed.
We thank the reviewer for these comments. This is indeed an important aspect, which we now expand on in the discussion section. We also took advantage of RNA-seq datasets for HUVECs (a non-transformed cell lines) in response to hypoxia (Sup. Figure S15), TNF-alpha with and without RelA depletion (Sup. Figure S16). These data support our findings that in hypoxia NF-kB is important for transcriptional repression, with some contributions to gene induction, even in a non-transformed cell system.
In the publicly available data sets analyzed, were the same hypoxic conditions used as in this study. This information should be included.
We apologize if this was not clear, the hypoxia RNA-seq studies are the same oxygen level and time (1%, 24 hours), this is in the legend of Figure 4A and Sup. Figure S9 and in Sup. Table S2. We have added this information to the main text also.
We have included this in the discussion and also analysed available data in HUVECs in response to cytokine stimulation with and without RelA depletion (Sup. Figure S16). This analysis revealed equal importance of RelA for activation and repression of genes upon TNF-alpha stimulation. Around 40% of genes require RelA for their induction or repression in response to TNF-a. In the discussion we have also included other references where NF-kappaB has been found to repress genes.
NF-kappaB has previously been shown to regulate HIF-1alpha transcription. What are the effects of NF-kappaB subunit siRNAs on basal HIF-1alpha transcription? In figure 7, it appears that NF-kappaB subunit siRNA is without effect on hypoxia-induced HIF protein expression. Could this account for some of the effects of NF-kappaB depletion on the hypoxic gene signature? This point needs to be clarified in light of the data presented.
We have included data for HIF-1α RNA levels in HeLa cells with/without NF-____k____B____ depletion followed by 24 hours of hypoxia (Sup. Figure S20) and we see a small reduction (~10-20%). The reviewer is correct, there was not much effect of NF-____k____B____ depletion on HIF-1α protein levels following 24 hours hypoxia in HeLa cells. Effects of NF-kappaB depletion can be found usually with lower times of hypoxia exposure or when more than one subunit is depleted at the same time. We have added this as a discussion point in the revised manuscript.
NRF-2 is a key cellular sensor of oxidative stress in a similar way to HIF being a hypoxia sensor. The authors demonstrate using a dye that ROS are paradoxically increased in hypoxia (a more controversial finding than the authors present). It would be of interest to know if NFR-2 is induced in hypoxia as a marker of cellular oxidative stress. Similarly, it would be interesting to determine by metabolic analysis whether oxidative phosphorylation (O2 consumption) is decreased as the transcriptional signature would suggest (although the difficulty of performing metabolic analysis in hypoxia is acknowledged).
To investigate if NRF2 is induced, we performed a western blot at 0, 1, and 24 hours 1% oxygen, but didn’t see any induction of NRF2 protein levels (____Sup. Figure S17A). We also overlapped our hypoxia upregulated genes with NRF2 target genes from {PMID:24647116 and PMID: 38643749} (Sup. Figure S17B) and found limited evidence of NRF2 target genes being induced. Based on these findings, it seems that NRF2 is not being induced in hypoxia, at least not at the hypoxia level/time point we have analysed. We also agree it would be ideal to measure oxygen consumption in hypoxia, but unfortunately, we do not have the technical ability to do this at present.
Reviewer #3 (Evidence, reproducibility and clarity (Required)):
Strengths This manuscript attempts to integrate multiple strands of data to determine the role of NFkB in hypoxia -induced gene expression. This analysis looks at multiple NFkB subunits in multiple cell lines to convincingly demonstrate that NFkB does indeed play a central role in the regulation of hypoxia-induced gene expression. This broad approach integrates new experimental data with findings from the published literature.
A significant amount of work has been performed both experimentally and bioinformatically to test experimental hypotheses.
We thank this reviewer for their positive comments.
Limitations
The main analysis in the paper involves comparing the impact of knocking down different NFkB family members in hypoxia and comparing transcriptional responses. I am surprised that the authors did not include the impact of knockdown of the NFkB family members in normoxia too. The absence of these control experiments allows us to understand the role of NFkB in hypoxia, but does not give us information as to how many of those impacts are specific/ induced in hypoxic conditions. i.e. many of the observed effects of NFkB knockdown could be due to basal suppression of NFkB target genes that happen to be hypoxia sensitive. This finding is obviously important, but it would be nice to know how many of those genes are only / preferentially regulated by NFkB in hypoxia. This would give a much deeper insight into the role of NFkB in hypoxia induced gene expression.
We agree this would have been ideal. For financial reasons we limited our analysis to hypoxia samples. We have performed qPCR analysis depleting RelA, RelB and cRel under normal oxygen conditions in HeLa (Sup. Figure S8). We find that the majority of the validated genes in HeLa cells which require____ NF-____k____B for gene changes in hypoxia, are not regulated by N____F-____k____B under normal oxygen conditions____. We have also added this limitation into our discussion section.
The broad experimental approach while a strength of the paper in many ways also has its limitations e.g. Motif analysis revealing e.g. HIF-1a binding site enrichment in RelA and RelB-dependent DEGs is correlative observation and does not prove HIF involvement in NFkB-dependent hypoxia induced gene activation. Comparing responses with responses seen in one cell type with responses that have been described in a database comprised of many studies in a variety of different cells also has some limitations. These points can be described more fully in the discussion
We agree these are mere correlations and hence a limitation and we have not formerly tested the involvement of HIF. We have included this in the discussion as suggested. For HIF binding site correlation, we do also compare to HIF ChIP-seq in HeLa cells exposed to 1% oxygen, albeit at 8 hours and not 24 hours (Sup. Figure S4).
For siRNA transfections, single oligonucleotide sequences were used for RelA, RelB and cRel. This increases the potential likelihood of 'off targets' compared to pooled oligos delivered at lower concentrations. This limitation should at least be mentioned.
We agree and have now included this as a limitation in the discussion section. We have now also included analysis using wild type and 2 different IKK____⍺____/β____ double KO CRISPR cell lines generated in the following publication {PMID: 35029639}. Out of the 9 genes we identified as NF-____k____B-dependent hypoxia upregulated genes from HeLa cell RNA-seq and validated by qPCR, which are also hypoxia-responsive in HCT116 cells (Sup. Figure S11D), 6 displayed ____NF-____k____B dependence in HCT116 cells (Sup. Figure S14). We also provide new protein data in this cell system for oxidative phosphorylation markers, which show as with the siRNA depletion, rescue of repression of these proteins when NF-____k____B is inactivated.
RNA-seq experiments are performed on n=2 data which means relatively low statistical power. How has the statistical analysis been performed on normalised counts (corresponding to 2 n- numbers) to yield statistical significance? I am not familiar with hypergeometric tests - please justify their use here.
__*We use DESeq2 for differential expression analysis and filter for effect size (> -/+ 0.58 log2 fold change) and statistical significance (FDR I am not familiar with hypergeometric tests - please justify their use here.
The hypergeometric test (equivalent to a one-sided Fisher's exact test) is routinely used to determine whether the observed overlap between two gene lists is statistically significant compared to what would be expected by chance. It is also the statistical test of choice for popular bioinformatics tools which perform over representation analysis (ORA) to see which gene sets/groups/pathways/ontologies are over-represented in a gene list, examples include Metascape, clusterProfiler, WebGestalt (used in this study), and gProfiler.
P14 RelB is described as having the most widespread impact of hypoxia dependent gene changes across all cell systems tested. Could this be due to a more potent silencing of RelB and / or due to particularly high/ low expression of RelB in these cells in general?
This is an excellent point, at the RNA level the RelB depletion is slightly more efficient (Sup. Figure S1), at the protein level, silencing is highly potent with all 3 siRNAs (Sup. Figure S1). We looked at the RNA levels of RelA, RelB and cRel in HeLa cells at basal conditions, and RelA shows the highest abundance compared to RelB and cRel, while RelB and cRel have similar expression levels (see below). However, RelB is very dynamic in response to hypoxia, something we have observed but have not published yet.
P18 For western blot analysis best practise is to have 2 MW markers per blot presented
We have and have added the second MW markers suggested.
For quantification, I suggest avoiding performing statistical analysis on semi-quantitative data unless a dynamic range of detection (with standards) has been fully established.
We agree this has many limitations, we will keep the quantification but moved into supplementary information.
P19 There is clearly an effect of reciprocal silencing with the NFkB knockdown experiments ie. siRelA affects RelB levels in hypoxia and vice versa. The implications of this for data interpretation should be discussed.
Indeed, it is well known that RelB and cRel are RelA targets. Less is known about RelA as it is not a known NF-____k____B____ target. We have added a discussion in the revised manuscript.
P20 The literature can be better cited in relation RelB and hypoxia A brief search reveals a few papers that should be mentioned/ discussed. Oliver et al. 2009 Patel et al. 2017 Riedl et al. 2021
We have looked into these suggestions. Oliver et al, refer to hypercapnia, not hypoxia and the other two only briefly mentioned RelB with no effects toward the goals of their studies. We have tried to incorporate what is currently known as much as possible.
I suggest leaving out mention of IkBa sumoylation and supplementary figure 10. I'm not sure the data in the paper as a whole merits focus on this very specific point.
We thank the reviewer for this suggestion and we have removed this aspect from the manuscript.
There is a very strong reliance on mRNA and TPM data. Some additional protein data in support of key findings will enhance
We have added additional protein level analysis where we could obtain antibodies, see Figures 6, 7 and Sup. Figures S17, S18, and S19 for our protein level analysis.
A graphical abstract summarising key findings with exemplar genes highlighted will enhance.
We have added a model to summarise our findings as suggested.
Both HIF and NFKB are ancient evolutionarily conserved pathways. Can lessons be learned from evolutionary biology as to how NFkB regulation of hypoxia induced genes occured. Does the HIF pathway pre-date the NFkB pathway or vice versa. This approach could be valuable in supporting the findings from this study.
We have investigated this. Unfortunately, there are very little available data on hypoxia gene expression in lower organisms. However, we have added a few sentences on the evolution of NF-____k____B____ and HIF.
Minor comments P2 please briefly explain how 5 genes give rise to 7 proteins
We have added this to the introduction as requested.
P2 there seems to be some recency bias in the studies cited as being associated with NFkB activation in response to hypoxia. Mention of Koong et al (1994) and Taylor et al (1999) and other early papers in the field will enhance
We have added these as suggested.
P3 The role of PHD enzymes in the regulation of NFkB in hypoxia can be introduced and / or discussed
We have added a reference to this aspect as suggested.
P8 I suggest use of proportional Venn diagrams to demonstrate the patterns more clearly
We have added these as suggested.
P11 To what extent might NFkB and Rest co-operate/ co-regulate gene repression in hypoxia?
This is a good question. We have overlapped our datasets with Rest-dependent hypoxia-regulated genes identified by Cavadas et al., (Figure below), and find that these appear to act independently of each other for the most part, with very few genes co-regulated by both.
Reviewer #3 (Significance (Required)):
Shakir et al. present a manuscript titled 'NFkB is a central regulator of hypoxia-induced gene expression'.
The research group are experts in both NFkB and hypoxia signaling and are the ideal group to perform these studies.
Hypoxia and inflammation are co-incident in many physiological and pathophysiological conditions, where the microenvironment affects disease severity and patient outcome. The cross talk between inflammatory and hypoxia signaling pathways is not fully described. Thus, this manuscript takes a novel approach to an established question and concludes clearly that NFkB is a central regulator of hypoxia-induced gene expression.
We thank the reviewer for these positive comments.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Strengths
This manuscript attempts to integrate multiple strands of data to determine the role of NFkB in hypoxia -induced gene expression. This analysis looks at multiple NFkB subunits in multiple cell lines to convincingly demonstrate that NFkB does indeed play a central role in the regulation of hypoxia-induced gene expression. This broad approach integrates new experimental data with findings from the published literature.
A significant amount of work has been performed both experimentally and bioinformatically to test experimental hypotheses.
Limitations
The main analysis in the paper involves comparing the impact of knocking down different NFkB family members in hypoxia and comparing transcriptional responses. I am surprised that the authors did not include the impact of knockdown of the NFkB family members in normoxia too. The absence of these control experiments allows us to understand the role of NFkB in hypoxia, but does not give us information as to how many of those impacts are specific/ induced in hypoxic conditions. i.e. many of the observed effects of NFkB knockdown could be due to basal suppression of NFkB target genes that happen to be hypoxia sensitive. This finding is obviously important, but it would be nice to know how many of those genes are only / preferentially regulated by NFkB in hypoxia. This would give a much deeper insight into the role of NFkB in hypoxia induced gene expression.
The broad experimental approach while a strength of the paper in many ways also has its limitations e.g. Motif analysis revealing e.g. HIF-1a binding site enrichment in RelA and RelB-dependent DEGs is correlative observation and does not prove HIF involvement in NFkB-dependent hypoxia induced gene activation. Comparing responses with responses seen in one cell type with responses that have been described in a database comprised of many studies in a variety of different cells also has some limitations. These points can be described more fully in the discussion
For siRNA transfections, single oligonucleotide sequences were used for RelA, RelB and cRel. This increases the potential likelihood of 'off targets' compared to pooled oligos delivered at lower concentrations. This limitation should at least be mentioned.
RNA-seq experiments are performed on n=2 data which means relatively low statistical power. How has the statistical analysis been performed on normalised counts (corresponding to 2 n- numbers) to yield statistical significance? I am not familiar with hypergeometric tests - please justify their use here.
P14 RelB is described as having the most widespread impact of hypoxia dependent gene changes across all cell systems tested. Could this be due to a more potent silencing of RelB and / or due to particularly high/ low expression of RelB in these cells in general?
P18 For western blot analysis best practise is to have 2 MW markers per blot presented
For quantification, I suggest avoiding performing statistical analysis on semi-quantitative data unless a dynamic range of detection (with standards) has been fully established.
P19 There is clearly an effect of reciprocal silencing with the NFkB knockdown experiments ie. siRelA affects RelB levels in hypoxia and vice versa. The implications of this for data interpretation should be discussed.
P20 The literature can be better cited in relation RelB and hypoxia A brief search reveals a few papers that should be mentioned/ discussed. Oliver et al. 2009 Patel et al. 2017 <br /> Riedl et al. 2021
I suggest leaving out mention of IkBa sumoylation and supplementary figure 10. I'm not sure the data in the paper as a whole merits focus on this very specific point.
There is a very strong reliance on mRNA and TPM data. Some additional protein data in support of key findings will enhance
A graphical abstract summarising key findings with exemplar genes highlighted will enhance.
Both HIF and NFKB are ancient evolutionarily conserved pathways. Can lessons be learned from evolutionary biology as to how NFkB regulation of hypoxia induced genes occured. Does the HIF pathway pre-date the NFkB pathway or vice versa. This approach could be valuable in supporting the findings from this study.
Minor comments
P2 please briefly explain how 5 genes give rise to 7 proteins
P2 there seems to be some recency bias in the studies cited as being associated with NFkB activation in response to hypoxia. Mention of Koong et al (1994) and Taylor et al (1999) and other early papers in the field will enhance
P3 The role of PHD enzymes in the regulation of NFkB in hypoxia can be introduced and / or discussed
P8 I suggest use of proportional Venn diagrams to demonstrate the patterns more clearly
P11 To what extent might NFkB and Rest co-operate/ co-regulate gene repression in hypoxia?
Shakir et al. present a manuscript titled 'NFkB is a central regulator of hypoxia-induced gene expression'.
The research group are experts in both NFkB and hypoxia signaling and are the ideal group to perform these studies.
Hypoxia and inflammation are co-incident in many physiological and pathophysiological conditions, where the microenvironment affects disease severity and patient outcome. The cross talk between inflammatory and hypoxia signaling pathways is not fully described. Thus, this manuscript takes a novel approach to an established question and concludes clearly that NFkB is a central regulator of hypoxia-induced gene expression.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
The work from Shakir et al uses different cell line models to investigate the role of NF-kB in the transcriptional adaptation of cells to hypoxia, which is relevant. In addition, the manuscript contains a large amount of data that could be of interest and even useful for researchers in the field of hypoxia and NF-kB. However, in my opinion, there are several concerns that should be revised and additional experiments that could be included to strengthen the relevance of the work.
Specific issues:
In Figure 1A, the authors examine which of the genes induced by hypoxia require NF-kB by RNA sequencing analysis of cells knocked down for specific NF-kB subunits and exposed to hypoxia for 24 hours. The knockdown is about 40-60% at the RNA level, but it would be helpful to show the effect of knockdown at the protein level. All the data regarding genes induced by hypoxia in control or NF-kB siRNA-treated cells are somewhat confusing. If I understand correctly, when the data from the three different siRNAs are crossed, only 1070 genes are upregulated and 295 are downregulated in an NF-kB-independent manner. If this is the case, I think it would be easier to use this information in Figure 2 to define the hypoxia-induced genes that are NF-kB-dependent by simply considering those induced in the control that are not in the NF-kB-independent subset (rather than repeating the integration of the data without additional explanation). If the authors do this simple analysis, are the resulting genes the same or similar? In any case, the way these numbers are obtained should be shown more clearly (i.e., a new Venn diagram showing genes up- or down-regulated in the siRNA control that are not up- or down-regulated in any of the siRNA-NF-kB treatments). Figure 2H shows that approximately 80% of the NF-kB-dependent genes up- or down-regulated in hypoxia were identified as RelA targets, which is statistically significant compared to RelB or cRel targets. However, what is the proportion of genes identified as RelA targets in the subset of NF-kB-independent hypoxia-induced genes? And in a randomly selected set of 500-600 genes? In my opinion, this statistical analysis should be included to demonstrate a relationship between NF-kB recruitment and hypoxia-induced upregulation (expected) and downregulation (unexpected). In this context, it is surprising that HIF consensus sites are preferentially detected in the genes that are supposed to be NF-kB dependent instead of RelA consensus. Figure 3 is just a confirmation by qPCR of the data obtained in the RNA-seq analysis, which in my opinion should be included as supplementary information. Moreover, both the effects of hypoxia and reversion by RelB siRNA are modest in several of the genes tested. The same is true for Figures 4 and 5 with very modest and variable results across cell types and genes. Figure 6 shows the effect of NF-kB knockdown on the induction of ROS in response to hypoxia. In the images provided, the effect of hypoxia is minimal in control cells, with the only clear differences shown in RelA-depleted cells. In 6B it is not clear what the three asterisks in the normoxia control represent (compared to the hypoxia siRNA control?). This should be clarified in the figure legend or text. In the Western blot of 6C, there are no differences in the levels of SOD1 after RelA depletion. Again, there is no reason not to include the NF-kB subunits in the Western blot analysis. Finally, regarding Figure 7, the authors mention that "we confirmed that hypoxia led to a reduction in several proteins represented in this panel (of proteins involved in oxidative phosphorylation), such as UQCRC2 and IDH1 (Figure 7A-B)". The authors cannot say this because it is not seen in the Western blot in 7A or in the quantification shown in 7B. In my personal opinion, stating something that is not even suggested in the experiments is very negative for the credibility of the whole message. In conclusion, this paper contains a large amount of relevant information, but i) non-essential data should be moved to Supplementary, ii) protein levels of relevant players need to be shown in addition to RNA, iii) minimal or undetectable differences need to be considered as no-differences, and iv) a model showing what is the interpretation of the data provided is needed to better understand the message of the paper. I mean, is it p65 or RelB binding to some of these genes leading to their activation or repression, or is it RelA or RelB inducing HIF1beta leading to NF-kB-dependent gene activation by hypoxia? If this were the case, experimental evidence that NF-kB regulates a subset of hypoxia genes through HIF1beta would make the story more understandable.
The work presented here is interesting but does not provide a major advance over previous publications, the main message being that a subset of hypoxia-regulated genes are NF-kB dependent. However, there is no mechanistic explanation of how this regulation is achieved and several data that are not clearly connected. A more comprehensive analysis of the data and additional experimental validation would greatly enhance the significance of the work.
NA then
NA and then
Author response:
The following is the authors’ response to the original reviews
Reviewer #1 (Public Review):
(…) In my view, the part about NF-YA1 is less strong - although I realize this is a compelling candidate to be a regulator of cell cycle progression, the experimental approaches used to address this question falls a bit short, in particular, compared to the very detailed approaches shown in the rest of the manuscript. The authors show that the transcription factor NF-YA1 regulates cell division in tobacco leaves; however, there is no experimental validation in the experimental system (nodules). All conclusions are based on a heterologous cell division system in tobacco leaves. The authors state that NF-YA1 has a nodule-specific role as a regulator of cell differentiation. I am concerned the tobacco system may not allow for adequate testing of this hypothesis.
Reviewer #1 makes a valid point by asking to focus the manuscript more explicitly on the role of NF-YA1 as a differentiation factor in a symbiotic context. We have now addressed this formally and experimentally.
The involvement of A-type NF-Y subunits in the transition to the early differentiation of nodule cells has been documented in model legumes through several publications that we refer to in the revised version of the discussion (lines 617/623). We fully agree that the CDEL system, because it is heterologous, does not allow us more than to propose a parallel explanation for these observations - i.e_., that the Medicago NF-YA1 subunit presumably acts in post-replicative cell-cycle regulation at the G2/M transition. Considering your recommendations and those of reviewer #2, we sought to support this conclusion by testing the impact of localized over-expression of _NF-YA1 on cortical cell division and infection competence at an early stage of root colonization. The results of these experiments are now presented in the new Figure 9 and Figure 9-figure supplement 1-5 and described from line 435 to 495.
With the fluorescent tools the authors have at hand (in particular tools to detect G2/M transition, which the authors suggest is regulated by NF-YA1), it would be interesting to test what happens to cell division if NF-YA1 is over-expressed in Medicago roots?
To limit pleiotropic effects of an ectopic over-expression, we used the symbiosis-induced, ENOD11 promoter to increase NF-YA1 expression levels more specifically along the trajectory of infected cells. We chose to remain in continuity with the experiments performed in the CDEL system by opting for a destabilized version of the KNOLLE transcriptional reporter to detect the G2/M transition. The results obtained are presented in Figure 9B (quantification of split infected cells), in Figure 9-figure supplement 1B (ENOD11 expression profile), in Figure 9-figure supplement 3B (representative confocal images) and Figure 9-figure supplement 4D (quantification of pKNOLLE reporter signal). There, we show that mitosis remains inhibited in cells accommodating infection threads, but is completed in a higher proportion of outer cortical cells positioned on the infection trajectory, where ENOD11 gene transcription is active before their physical colonization.
Based on NF-YA1 expression data published previously and their results in tobacco epidermal cells, the authors hypothesize that NF-YA regulates the mitotic entry of nodule primordial cells. Given that much of the manuscript deals with earlier stages of the infection, I wonder if NF-YA1 could also have a role in regulating mitotic entry in cells adjacent to the infection thread?
The expression profile of NF-YA1 at early stages of cortical infection (Laporte et al., 2014) is indeed similar to the one of ENOD11 (as shown in Figure 9-figure supplement 1C) in wild-type Medicago roots, with corresponding transcriptional reporters being both activated in cells adjacent to the infection thread. Under our experimental conditions, additional expression of NF-YA1 (driven by the ENOD11 promoter) in these neighbouring cells did not impact their propensity to enter mitosis and to complete cell division. These results are presented in Figure 9-figure supplement 4D (quantification of pKNOLLE reporter signal) and Figure 9-figure supplement 5 (quantification of split neighbouring cells).
Reviewer #1 (Recommendations For The Authors):
- In the first part, images show the qualitative presence/absence of H3.1 or H3.3 histones.
Upon closer inspection, many cells seem to have both histones. In Fig1-S1 for example (root meristem), it is evident that there are many cells with low but clearly present H3.1 content in the green channel; however, in the overlay, the green is lost and H3.3 (pink) is mainly visible. What does this mean in terms of the cell cycle?
We fully agree with reviewer #1 on these points. Independent of whether they have low or high proliferation potential, most cells retain histone H3.1 particularly in silent regions of the genome, while H3.3 is constitutively produced and enriched at transcriptionally active regions. When channels are overlaid, cells in an active proliferation or endoreduplication state (in G1, S or G2, depending on the size of their nuclei) will appear mainly "green" (H3.1-eGFP positive). Cells with a low proliferation potential (e.g., in the QC), G2-arrested (e.g., IT-traversed) or terminally differentiating (e.g., containing symbiosomes or arbuscules) will appear mainly "magenta" (H3.1-low, medium to high H3.3-mCherry content).
Furthermore, all nodule images only display the overlay image, and individual fluorescence channels are not shown. Does the same masking effect happen here? It may be helpful to quantify fluoresce intensity not only in green but also in red channels as done for other experiments.
Quantifying fluorescence intensity in the mCherry channel may indeed help to highlight the likely replacement of H3.1-eGFP by H3.3-mCherry in infected cells, as described by Otero and colleagues (2016) at the onset of cellular differentiation. However, the quantification method as established (i.e., measuring the corrected total nuclear fluorescence at the equatorial plane) cannot be applied, most of the time, to infected cells' nuclei due to the overlapping presence of mCherry-producing S. meliloti in the same channel (e.g., in Figure 2B). Nevertheless, and to avoid this masking effect when the eGFP and mCherry channels are overlaid, we now present them as isolated channels in revised Figures 1-3 and associated figure supplements. As the cell-wall staining is regularly included and displayed in grayscale, we assigned to both of them the Green Fire Blue lookup table, which maps intensity values to a multiple-colour sequential scheme (with blue or yellow indicating low or high fluorescence levels, respectively). We hope that this will allow a better appreciation of the respective levels of H3.1- and H3.3-fusions in our confocal images.
- Fig 1 B - it is hard to differentiate between S. meliloti-mCherry and H3.3-mCherry. Is there a way to label the different structures?
In the revised version of Figure 1B, we used filled or empty arrowheads to point to histone H3-containing nuclei. To label rhizobia-associated structures, we used dashed lines to delineate nodule cells hosting symbiosomes and included the annotation “IT” for infection threads. We also indicated proliferating, endoreduplicating and differentiating tissues and cells using the following annotations: “CD” for cell division, “En” for endoreduplication and “TD” for terminal differentiation. All annotations are explained in the figure legend.
- Fig 1 - supplement E and F - no statistics are shown.
We performed non-parametric tests using the latest version of the GraphPad Prism software (version 10.4.1). Stars (Figure 1-figure supplement 1F) or different letters (Figure 1-figure supplement 1G) now indicate statistically significant differences. Results of the normality and non-parametric tests were included in the corresponding Source Data Files (Figure 1 – figure supplement 1 – source data 1 and 2). We have also updated the compact display of letters in other figures as indicated by the new software version. The raw data and the results of the statistical analyses remain unchanged and can be viewed in the corresponding source files.
- Fig 2 A - overview and close-up image do not seem to be in the same focal plane. This is confusing because the nuclei position is different (so is the infection thread position).
We fully agree that our former Figure may have confused reviewers #1 and #2 as well as readers. Figure 2A was designed to highlight, from the same nodule primordium, actively dividing cells of the inner cortex (optical section z 6-14) and cells of the outer cortex traversed, penetrated by or neighbouring an infection thread (optical section z 11-19). We initially wanted to show different magnification views of the same confocal image (i.e_._, a full-view of the inner cortex and a zoomed-view of the outer layers) to ensure that audiences can identify these details. In the revised version of Figure 2A, we displayed these full- and zoomed-views in upper and lower panels, respectively and we removed the solid-line inset to avoid confusion.
- Fig 1A and Fig 2E could be combined and shown at the beginning of the manuscript. Also, consider making the cell size increase more extreme, as it is important to differentiate G2 cells after H3.1 eviction and cells in G1. You have to look very closely at the graph to see the size differences.
We have taken each of your suggestions into account. A combined version of our schematic representation with more pronounced nuclei size differences is now presented in Figure 1A.
- Fig. 3 C is difficult to interpret. Can this be split into different panels?
We realized that our previous choice of representation may have been confusing. Each value corresponds only to the H3.1-eGFP content, measured in an infected cell and reported to that of the neighbouring cell (IC / NC) within individual root samples. Therefore, we removed the green-magenta colour code and changed the legend accordingly. We hope that these slight modifications will facilitate the interpretation of the results - namely, that the relative level of H3.1 increases significantly in infected cells in the selected mutants compared to the wild-type. This mode of representation also highlights that in the mutants, there are more individual cases where the H3.1 content in an infected cell exceeds that of the neighbouring cell by more than two times. These cases would be masked if the couples of infected cells and associated neighbours would be split into different panels as in Figure 3B.
- Line 357/359. I assume you mean ...'through the G2 phase can commit to nuclear division'.
We have edited this sentence according to your suggestion, which now appears in line 370.
Reviewer #2 (Recommendations For The Authors):
Cell cycle control during the nitrogen-fixing symbiosis is an important question but only poorly understood. This manuscript uses largely cell biological methods, which are always of the highest quality - to investigate host cell cycle progression during the early stages of nodule formation, where cortical infection threads penetrate the nodule primordium. The experiments were carefully conducted, the observations were detail oriented, and the results were thought-provoking. The study should be supported by mechanistic insights.
(1) One thought provoked by the authors' work is that while the study was carried out at an unprecedented resolution, the relationship between control of the cell cycle and infection thread penetration remains correlative. Is this reduced replicative potential among cells in the infection thread trajectory a consequence of hosting an infection thread, or a prerequisite to do so?
We understand and share the point of view of reviewer #2. At this stage, we believe that our data won’t enable us to fully answer the question, thus this relationship remains rather correlative. The reasons are that 1) the access to the status of cortical cells below C2 is restricted to fixed material and therefore only represents a snapshot of the situation, and 2) we are currently unable to significantly interfere with mechanisms as intertwined as cell cycle control and infection control. What we can reasonably suggest from our images is that the most favorable window of the cell cycle for cells about to be crossed by an infection thread is post-replicative, i.e., the G2 phase. Typical markers of the G2 phase were recurrently observed at the onset of physical colonization – enlarged nucleus, containing less histone H3.1 than neighbouring cells in S phase (e.g., in Figure 2A). Reaching the G2 phase could therefore be a prerequisite for infection (and associated cellular rearrangements), while prolonged arrest in this same phase is likely a consequence of transcellular passage towards a forming nodule primordium.
More importantly, in either scenario, what is the functional significance of exiting the cell cycle or endocycle? By stating that "local control of mitotic activity could be especially important for rhizobia to timely cross the middle cortex, where sustained cellular proliferation gives rise to the nodule meristem" (Line 239), the authors seem to believe that cortical cells need to stop the cell cycle to prepare for rhizobia infection. This is certainly reasonable, but the current study provides no proof, yet. To test the functional importance of cell cycle exit, one would interfere with G2/M transition in nodule cells, and examine the effect on infection.
We fully agree with reviewer #2 that the functional importance of a cell-cycle arrest on the infection thread trajectory remains to be demonstrated. Interfering with cell-cycle progression in a system as complex and fine-tuned as infected legume roots certainly requires the right timing – at the level of the tissue and of individual cells; the right dose; and the right molecular player(s) (i.e., bona fide activators or repressors of the G2/M transition). Using the symbiosis-specific NPL promoter, activated in the direct vicinity of cortical infection threads (Figure 9-figure supplement 1B), we tried to force infectable cells to recruit the cell division program by ectopically over-expressing the Arabidopsis CYCD3.1, “mimicking” the CDEL system. So far, this strategy has not resulted in a significant increase in the number of uninfected nodules in transgenic hairy roots - though the effect on symbiosome release remains to be investigated. Provided that a suitable promoter-cell cycle regulator combination is identified, we hope to be able to answer this question in the future.
Given that the authors have already identified a candidate, and showed it represses cell division in the CDEL system, not testing the same gene in a more relevant context seems a lost opportunity. If one ectopically expressed NY-YA1 in hairy roots, thus repressing mitosis in general, would more cells become competent to host infection threads? This seems a straightforward experiment and readily feasible with the constructs that the authors already have. If this view is too naive, the authors should explain why such a functional investigation does not belong in this manuscript.
Reviewer #2's point is entirely valid, and we decided to address it through additional experiments. To avoid possible side effects on development by affecting cell division in general, we placed NF-YA1 under control of the symbiosis-induced ENOD11 promoter. Based on the results obtained in the CDEL system, the pENOD11::FLAG-NF-YA1 cassette was coupled to a destabilized version of the KNOLLE transcriptional reporter to detect the G2/M transition. Competence for transcellular infection was maintained upon local NFYA1 overexpression, the latter leading to a slight (non-significant) increase in the number of infected cells per cortical layer. These results are presented in Figure 9-figure supplement 3A-B (representative confocal images) and in Figure 9-figure supplement 4A-
G.
(1b) A related comment: on Line 183, it was stated that "The H3.1-eGFP fusion protein was also visible in cells penetrated but not fully passed by an infection thread". Presumably, the authors were talking about the cell marked by the arrowhead. But its H3.1-GFP signal looks no different from the cell immediately to its left. It is hard to say which cells are ones "preparing for intracellular infection pass through S-phase", and which ones are just "regularly dividing cortical cells forming the nodule primordium". What can be concluded is that once a cell has been fully transversed by an infection thread, its H3.1 level is low. Whether this is the cause or consequence of infection cannot be resolved simply by timing the appearance or disappearance of H3.1-GFP.
We basically agree with comment 1b. In an unsynchronized system such as infected hairy roots, it is challenging to detect the event where a cell is penetrated, but not yet completely crossed by an infection thread. What we wanted to emphasize in Figure 2A, is that host cells in the path of an infection thread re-enter the cell cycle and pass through S-phase just as their neighbours do (as pointed out by reviewer #2 in his summary). The larger nucleus with slightly lower H3.1-eGFP signal than the neighbouring cell (as indicated by the use of the Green Fire Blue lookup table) suggests that the infected cell marked by the arrowhead in Figure 2A is actually in the G2 phase. The main difference is indeed that cells allowing complete infection thread passage exit the cell cycle and largely evict H3.1 while their neighbours proceed to cell division (as exemplified by PlaCCI reporters in Figure 4CD and the new Figure 5-figure supplement 2). Whether cell-cycle exit in G2 is a cause, or a consequence of cortical infection is a question that cannot be easily answered from fixed samples, which is a limitation of our study.
(2) The authors have convincingly demonstrated that cortical cells accommodating infection threads exit the cell cycle, inhibit cell division, and down-regulate KNOLLE expression. How do these observations reconcile with the feature called the pre-infection thread? The authors devoted one paragraph to this question in the Discussion, but this does seem sufficient given that the pre-infection thread is a prominent concept. Is the resemblance to the cell division plane superficial, or does it reflect a co-option of the normal cytokinesis machinery for accommodating rhizobia?
From our point of view, cortical cells forming pre-infection threads are likely in an intermediate state. PIT structures undoubtedly share many similarities with cells establishing a cell division plane. The recruitment of at least some of the players normally associated with cytokinesis has been demonstrated and is consistent with the maintenance of infectable cells in a pre-mitotic phase in Medicago, as discussed in lines 558 to 568. We nevertheless think that the arrest of the cell cycle in the G2 phase, presumably occurring in crossed cortical cells, constitutes an event of cellular differentiation and specialization in transcellular infection.
The following are mainly points of presentation and description:
(3) Line 158: I can't see "subnuclear foci" in Figure 1-figure supplement 1C-E. However, they are visible in Fig. 1C.
We hope that presenting the eGFP and mCherry channels in separate panels and assigning them the Green Fire Blue colour scheme provides better visibility and contrast of these detailed structures. We now refer to Figure 1C in addition to Figure 1–figure supplement 1E in the main text (line 161).
(4) Line 160: The authors should outline a larger region containing multiple QC cells, rather than pointing to a single cell, as there are other areas in the image containing cells with the same pattern.
We updated Figure 1-figure supplement 1E accordingly.
(5) Fig. 1B should include single channels, since within a single plant cell, the nucleus, the infection thread, and sometimes symbiosomes all have the same color. This makes it hard to see whether the nuclei in these cells are less green, or are simply overwhelmed by the magenta color.
To improve the readability of Figure 1B and to address suggestions from individual reviewers, we now include separate channels and have annotated the different structures labeled by mCherry.
(6) Fig. 2A: the close-up does not match the boxed area in the left panel. Based on the labeling, it seems that the two panels are different optical sections. But why choose a different optical depth for the left panel? This can be disorienting to the author, because one expects the close-up to be the same image, just under higher magnification.
We fully agree that our previous choice of representation may have been confusing. As we also specified to reviewer #1, we wanted to show a full-view of proliferating cells in the inner cortex and a zoomed-view of infected cells in the outer layers of the same nodule primordium. In the revised version of Figure 2A, we displayed these full- and zoomedviews in separate panels and removed the boxed area to avoid confusion.
(7) Figure 2-figure supplement 1B: the cell indicated by the empty arrowhead has a striking pattern of H3.1 and H3.3 distribution on condensed chromosomes. Can you comment on that?
Reviewer #2 may be referring to the apparent enrichment of H3.3 at telomeres, previously described in Arabidopsis, while pericentromeric regions are enriched in H3.1. This distribution is indeed visible on most of the condensed chromosomes shown in Figure 2-figure supplement 1B. We included this comment in the corresponding caption.
(8) Fig. 4: It is not very easy to distinguish M phase. Can the authors describe how each phase is supposed to look like with the reporters?
We agree with reviewer #2 and attempted to improve Figure 4, which is now dedicated to the Arabidopsis PlaCCI reporter. ECFP, mCherry, and YFP channels were presented separately and the corresponding cell-cycle phases (in interphase and mitosis) were annotated. The Green Fire Blue lookup table was assigned to each reporter to provide the best visibility of, for example, chromosomes in early prophase. We included a schematic representation corresponding to the distribution of each reporter, using the colors of the overlaid image to facilitate its interpretation.
(9) Line 298: what is endopolyploid? This term is used at least three times throughout the manuscript. How is it different from polyploid?
In the manuscript, we aimed to differentiate the (poly)ploidy of an organism (reflecting the number of copies of the basic genome and inherited through the germline) from endopolyploidy produced by individual somatic cells. As reviewed by Scholes and Paige, polyploidy and endopolyploidy differ in important ways, including allelic diversity and chromosome structural differences. In the Medicago truncatula root cortex for example, a tetraploid cell generated via endoreduplication from the diploid state would contain at most two alleles at any locus. The effects of endopolyploidy on cell size, gene expression, cell metabolism and the duration of the mitotic cell cycle are not shared among individual cells or organs, contrasting to a polyploid individual (Scholes and Paige, 2015).
See Scholes, D. R., & Paige, K. N. (2015). Plasticity in ploidy : A generalized response to stress. Trends in Plant Science, 20(3), 165‑175. https://doi.org/10.1016/j.tplants.2014.11.007
(10) Line 332: "chromosomes on mitotic figures" - what does this mean?
Reviewer #2 is right to point out this redundant wording. Mitotic “figures” are recognized, by definition, based on chromosome condensation. We now use the term "mitotic chromosomes" (line 344).
(11) Fig. 6A: could the authors consider labeling the doublets, at least some of them? I understand that this nucleus contains many doublets. However, this is the first image where one is supposed to recognize these doublets, and pointing out these features can facilitate understanding. Otherwise, a reader might think the image is comparable to nuclei with no doublets in the rest of the figure.
Following this suggestion, five of these doublets are now labeled in Figure 7A (formerly Figure 6A).
Ira Madison III. 'La La Land'’s White Jazz Narrative. MTV, December 2016. URL: https://www.mtv.com/news/5qr32e/la-la-lands-white-jazz-narrative (visited on 2023-12-10).
I consulted the article "La La Lands White Jazz Narrative" by Ira Madison III and found his viewpoints very inspiring. He pointed out that in the film La La Land, the male lead's insistence on defending true jazz is actually a way of presenting the occupation of African American cultural space by white characters. This made me start to re-examine some of the movies I used to like and reflect on whether there were similar cultural appropriation phenomena in them. Although the creator may have no ill intentions, this article reminds us that we need to be more vigilant in thinking about which voices are highlighted and which groups are marginalized.
Sometimes colonialism is a little bit more subtle. For example, the term “White Savior” is a sarcastic term for white people being seen as liberating or rescuing “helpless” non-white people. This is common in TV and movies (see the TVTropes entries on Mighty Whitey [t7], and White Man’s Burden [t8]). For example, consider movies like James Cameron’s Avatar movies, Dances with Wolves, The Last Samuri (starring the Tom Cruise), The Blind Side, The Help, La La Land (where a white man is trying to save “real” Jazz music [t9]), etc.
Reading this part made me realize that colonialism not only exists in history, but also profoundly influences modern media and popular culture. I have watched movies like Help and Avatar before, but I never realized that they were conveying the narrative style of "white saviors". This made me start to reflect on whether many of the stories I received in the past also contained similar biases. How can we create more works that truly reflect the voices of non-white groups instead of always having white characters represent the saviors?
Melewatkan task aplikasi ke eksekutor
ada eksekutor sehingga programmer tidak perlu memikirkan threading.
awaitTermination() setelah shutdown().Interface
Harus di implement? -> Di implementasikan oleh implements seperti ArrayList, HashSet - Iya, interface Collection ini harus diimplementasikan oleh class lain supaya bisa digunakan<br /> - Interface hanya berisi deklarasi method (tanpa isi), jadi class yang mengimplementasikan harus menyediakan isi/logic dari method-method tersebut<br /> - Contoh class yang mengimplementasikan Collection: ArrayList, HashSet, LinkedList<br /> - Dengan implementasi ini, class-class tersebut bisa punya fitur dasar koleksi seperti tambah, hapus, cek isi, dll<br /> - Tujuannya supaya semua koleksi punya perilaku standar dan bisa digunakan dengan cara yang sama
Selon Eurostat, évolution des prélèvements obligatoires entre 2018 et 2023, en % du PIB. La moyenne européenne est de 40,6% en 2023 2018 48,1 2023 45,6
L'argument du "cadeau fait aux riches" est bidon, et Macron est un putain de communiste. Les tarés perdus qui hurlent à l'ultra libéralisme sont des enfants débiles à frapper.
must implement and maintain a standards-based Application Programming Interface (API)
Here it says that the MA plans must be "standards-based".
In the Interoperability and Patient Access final rule (CMS-9115-F), CMS encouraged MA-PD plans to build a Provider Directory API that is conformant to the Health Level Seven International® (HL7®) PDex Plan-Net Implementation Guide (85 FR 25529).
Apparently PDex standard is "encouraged." but not required.
§ 1o
Princípio da subsidiariedade, devendo-se entender pelo cabimento de outras ações de controle abstrato, não se aplicando, por exemplo, o princípio da subsidiariedade se couber ação de controle difuso. Nesse sentido:
(SARLET, Ingo; MARINONI, Luiz G.; MITIDIERO, Daniel. Curso de Direito Constitucional - 13ª Edição 2024. 13. ed. Rio de Janeiro: Saraiva Jur, 2024. E-book. p.1282. ISBN 9788553621163. Acesso em: 31 mai. 2025.)
RRID:AB_2760821
DOI: 10.1080/19490976.2025.2508949
Resource: None
Curator: @scibot
SciCrunch record: RRID:AB_2760821
RRID:AB_354431
DOI: 10.1016/j.immuni.2025.05.003
Resource: (R and D Systems Cat# AF-263-NA, RRID:AB_354431)
Curator: @scibot
SciCrunch record: RRID:AB_354431
RRID:AB_312976
DOI: 10.1016/j.chom.2025.05.007
Resource: (BioLegend Cat# 103111, RRID:AB_312976)
Curator: @scibot
SciCrunch record: RRID:AB_312976
AB_1595470
DOI: 10.1016/j.celrep.2025.115780
Resource: (BioLegend Cat# 202526, RRID:AB_1595470)
Curator: @scibot
SciCrunch record: RRID:AB_1595470
AWS is taking out the trash 🗑️



Peersuite, a decentralized workspace
Peersuite, a decentralized workspace


Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
Response to Review
Manuscript number: RC-2024-02391
Corresponding author(s): John Varga
Dibyendu Bhattacharyya
[Please use this template only if the submitted manuscript should be considered by the affiliate journal as a full revision in response to the points raised by the reviewers.
If you wish to submit a preliminary revision with a revision plan, please use our "Revision Plan" template. It is important to use the appropriate template to clearly inform the editors of your intentions.]
This section is optional. Insert here any general statements you wish to make about the goal of the study or about the reviews.
Dear editor,
We are pleased to submit a full revised version of the manuscript that addresses all the points raised by the reviewers. We have included new experiments and modified the text and figures based on the reviewers’ suggestions. We thank all the reviewers for their insightful feedback, which has significantly enhanced the quality of the manuscript. We are confident and optimistic that our improved manuscript will be accepted by the journal of our choice.
This document is supposed to contain a few images, which were somehow missing after the processing through the manuscript submission path. For convenience we also included a PDF version of the response to reviewers.
This section is mandatory. *Please insert a point-by-point reply describing the revisions that were already carried out and included in the transferred manuscript. *
Reviewer #1
To reliably quantify the ciliary length in different cell types, and in independent ciliary marker needs to be included for comparison and the ciliary base needs to be labeled (e.g., g-TUBULIN). This needs to combined with a non-biased, high-throughput analysis, e.g., CiliaQ, Response: As suggested, we compared primary cilia length measurements using antibodies against Arl13b and γ-tubulin. The comparison between healthy controls (HC) and systemic sclerosis (SSc) is presented in Supplementary Figure S1. No significant differences in primary cilia length were observed compared to our previous measurements. Cilia length was quantified using ImageJ version 1.48v (http://imagej.nih.gov/ij) with the maximum intensity projection (MIP) method and visualized through 3D reconstruction using the ImageJ 3D Viewer.
As mentioned in the study, TGFbhas been implicated to drive myofibroblast transition. Thus TGFb stimulate ciliary signaling in the presented primary cells? The authors should provide a read-out for TGFb signaling in the cilium (ICC for protein phosphorylation etc.). Furthermore, canonical ciliary signaling pathways have been suggested to act as fibrotic drivers, such as Hedgehog and Wnt signaling - does stimulation of these pathways evoke a similar effect? Response: Yes, TGF-β1 stimulates ciliary signaling in growth-arrested foreskin fibroblasts. Clement et al. (2013) showed that TGF-β1 induces p-SMAD2/3 at the ciliary base, followed by the nuclear translocation of p-SMAD2/3 after 90 minutes. To assess whether canonical ciliary signaling pathways influence primary cilia length, we treated foreskin fibroblasts with Wnt (#908-SH, R&D) and a Shh agonist (#5036-WN, R&D) at 100 ng/mL each for 24 hours. We did not observe any changes in primary cilia length under either condition. These data are shown here for reference but are not included in the manuscript.
Clement, Christian Alexandro, et al. "TGF-β signaling is associated with endocytosis at the pocket region of the primary cilium." Cell reports 3.6 (2013): 1806-1814.
Kawasaki, Makiri, et al. "Primary cilia suppress the fibrotic activity of atrial fibroblasts from patients with atrial fibrillation in vitro." Scientific Reports 14.1 (2024): 12470.
Lee, J., Choi, JH. & Joo, CK. TGF-β1 regulates cell fate during epithelial–mesenchymal transition by upregulating survivin. Cell Death Dis 4, e714 (2013). https://doi.org/10.1038/cddis.2013.244.
Liu, Y. et al. TGF-β1 promotes scar fibroblasts proliferation and transdifferentiation via up-regulating MicroRNA-21. Sci. Rep. 6, 32231; doi: 10.1038/srep32231 (2016).
Kolodsick, Jill E., et al. "Prostaglandin E2 inhibits fibroblast to myofibroblast transition via E. prostanoid receptor 2 signaling and cyclic adenosine monophosphate elevation." American journal of respiratory cell and molecular biology 29.5 (2003): 537-544.
Kolodsick, Jill E., et al. "Prostaglandin E2 inhibits fibroblast to myofibroblast transition via E. prostanoid receptor 2 signaling and cyclic adenosine monophosphate elevation." American journal of respiratory cell and molecular biology 29.5 (2003): 537-544.
The authors describe that they wanted to investigate how aSMA impacted primary cilia length. They only provide a knock-down experiment and measured ciliary length, but the mechanistic insight is missing. How does loss of aSMA expression control ciliary length? Response: We measured acetylated α-tubulin levels in ACTA2 siRNA-treated cells compared to control-treated cells. Acetylated α-tubulin levels increased under ACTA2 siRNA-treated conditions, as shown in Figure 4D, and TPPP3 levels were also elevated (Figure S8A). Interestingly, TPPP3 levels negatively correlated with disease severity in SSc fibroblasts (r = -0.2701, p = 0.0183), and TPPP3 expression significantly reduced in SSc skin biopsies, as shown in Figures 6C and 6D. These results strengthen our hypothesis that microtubule polymerization and actin polymerization, while they counterbalance each other, also contrarily affect PC length. We agree that a much more detailed study is needed to extensively delineate the intricate homeostasis of the actin network and microtubule network in conjunction with fibrosis and primary cilia length. We have mentioned this in the discussion.
The authors used LiCl in their experiments, which supposedly control Hh signaling. Coming back to my second questions, is this Hh-dependent? And what is the common denominator with respect to TGFbsignaling? And how is this mechanistically connected to actin and microtubule polymerization? Response: We used Shh inhibitor (Cyclopamine hydrate #C4116 Sigma-Aldrich) in both SSc and foreskin fibroblasts (with and without TGFβ). We found that PC length is significantly increased and αSMA intensity is reduced in the Shh inhibitor treated group (data not included in the Manuscript)
How was the aSMA Mean intensity determined? Response: We quantified aSMA mean intensity using ImageJ, and the procedure has been added to the respective figure legend and materials and methods section under ‘Quantification of immunofluorescence’ (each point represents mean intensity from three randomly selected hpf/slide was performed using ImageJ).
Fig: 1D: Statistical test is missing in Figure Legend and presentation of the p-values for the left graph is confusing Response: We added statistical test information in Figure Legend.
Some graphs are presented {plus minus} SD and some {plus minus} SEM, but this is not correctly stated in the Material & Methods Part __Response: __We added information to the figure legend as well as in the Material & Methods section.
In general, text should be checked again for spelling mistakes and sentences may be re-written to promote readability. In particular, this applies to the discussion. __Response: __We checked and corrected.
Figure Legends are not written consistently, information is missing (e.g., statistical tests, see above). __Response: __We carefully checked and added information accordingly.
Figures should be checked again, and all text should be the same size and alignment of images should be improved. __Response: __We checked and corrected.
Significance
The authors present a novel connection between the regulation of primary cilia length and fibrogenesis. However, the study generally lacks mechanistic insight, in particular on how TGFb signaling, aSMA expression, and ciliary length control are connected. The spatial organization of the proposed signaling components is also not clear - is this a ciliary signaling pathway? If so, how does it interact with cytoplasmic signaling and vice versa?
Response: Thank you for your thoughtful and constructive feedback. We appreciate your recognition of the novelty of our study linking primary cilia length regulation to fibrogenesis. In our revised manuscript, we did provide a mechanistic insight, though. Our results suggest that during the fibrotic response, higher-order actin polymerization, along with microtubule destabilization resulting from tubulin deacetylation, drives the shortening of PC length. In contrast, PC length elongation via stabilization of microtubule polymerization mitigates the fibrotic phenotype in fibrotic fibroblasts. We agree that a deeper mechanistic understanding particularly regarding how TGFβ signaling, αSMA expression, and ciliary length control intersect is essential for fully elucidating the pathway. We also acknowledge the importance of clarifying the spatial organization of the signaling components and plan to incorporate such analyses in future studies.
Reviewer #2
*I found the paper to be rather muddled and its presentation made if somewhat difficult to follow. For example, the Figures are disorganised (Fig 1 is a great example of this) and there was reference to Sup data that appeared out of order (eg Sup Fig 2 appeared before Sup Fig 1 in the text). *
Response: We carefully revised the manuscript and arranged the figures.
*Images in a single figure should be the same size. Currently they are almost random and us different magnifications. Overall, the paper needs to be better organized. *
Response: We carefully revised the manuscript and figures provided with same magnification.
*I have some significant concerns about how the PC length data was generated. To my mind the length may be hard to determine from the type of images shown in the paper (which may represent the best images?). Some of the images presented appear to show shorter, fatter PCs in the cells from fibrosis cases. Is this real or is it some kind of artefact? Would a shorter, fatter PCs have a similar or larger surface area? What would be the consequence of this? *
Response: Primary cilia length was measured with ImageJ1.48v (using maximum intensity projection (MIP) method and visualized by 3D reconstruction with the ImageJ 3D viewer. Each small dot represents the PC length from an individual cell, and each large dot represents the average of the small dots for one cell line.
*I am confused as to exactly what is meant by matched healthy controls. Age, sex and ethnicity, where stated seem to be very variable? What are CCL210 fibroblasts? *
Response: We appreciate this comment. This is correct. The age, sex, and ethnicity are not matched for the available healthy controls. We have corrected that in the text. CCL210 is a commercially available fibroblast cell line that was isolated from the lung of a normal White, 20-year-old, female patient.
*What does a change in PC length signify? DO shot PC foe a cellular transition or are they a consequence of it? What would happen is you targeted PCs with a drug and that influenced the length on all cell types? Is the effect on PC fibroblast specific? *
__Response: __Significance and regulation of PC length are greatly debated and investigated still. It appears that PC length signify different features in different cell types. Although these are very interesting questions but such experiments are beyond the scope of our present work.
Minor concerns
*Page 4 second paragraph. I think it should be clarified that it is this group who have suggested a link between PCs and myofibroblast transition? *
__Response: __We agree with the reviewer and clarified it.
*Page 4 second paragraph. The use of the word "remarkably' is a bit subjective. *
__Response: __We agree with the reviewer and have removed it.
*Reference 27 is a paper on multiciliogenesis rather than primary ciliogenesis. *
__Response: __We agree with the reviewer and have removed it.
Figure 1 panel D. Make the image with the same sized vertical scale
__Response: __We have replaced it with a new Figure 1.
Significance
Reviewer #2 (Significance (Required)):
To my mind this is a novel paper and the data presented in it may be of interest to the cilia community as well as to the fibrosis field. This could be considered to be a significant advance and I am unaware that other groups are actively working in this area.
Presentation of the data in the current form does not instil confidence in the work.
Response: ____Thank you for recognizing the novelty and potential significance of our work. We appreciate your comments and fully acknowledge the concern regarding the presentation of the data. We have carefully revised the manuscript and reorganized the figures to improve clarity and overall presentation.
Reviewer #3
Major comments:
Avasthi, P., and W.F. Marshall. 2012. Stages of ciliogenesis and regulation of ciliary length. Differentiation. 83:S30-42.
The use of LiCl to increase ciliary length is complicated. What are the molecular mechanisms underlying this effect? It is known that it may be affecting GSK-3b, which can have other ciliary-independent effects. Therefore, using ciliary KO/KD cells (IFT88 or IFT20) as controls may help assess the specificity of the proposed treatments. Response: As explained in the previous paragraph, PC length regulations are dependent on multiple factors and many of them are not IFT dependent. One such method is directly modifying the axoneme by altering microtubule stability/polymerization through the acetylation state(Avasthi and Marshall, 2012), a pathway most likely the case for Tubacin. Another mode of PC length regulation is through a change in Actin polymerization. The remodeling of actin between contractile stress fibers and a cortical network alters conditions that are hospitable to basal body docking and maintenance at the cell surface (Avasthi and Marshall, 2012), causing PC length variation. Higher order microtubule polymerization inhibit actin polymerization. By interrogating RNA-seq data we determined that several PC-disassembly related genes (KIF4A, KIF26A, KIF26B, KIF18A), as well as microtubule polymerization protein genes (TPPP, TPPP3, TUBB, TUBB2A etc), were differentially expressed in LiCl-treated SSc fibroblasts (Suppl. Fig. S6D). Altogether, these findings suggest that microtubule polymerization/depolymerization mechanisms may regulate PC elongation and attenuation of fibrotic responses after either LiCl or Tubacin treatment.
Also, assessing the frequency of ciliary-expressing cells is important. That may give another variable important to predict fibrotic phenotypes. Or do 100% of the cultured cells express cilia in those conditions? Response: We carefully checked and observed almost 95% cells express cilia in cultured conditions.
Have the authors evaluated if TGF-b1 treatments induce cell cycle re-entry and proliferation in these experimental conditions? This is important to exclude ciliary resorption due to cell cycle re-entry instead of the myofibroblast activation process. __Response:__Yes, TGF-β induces cell proliferation in fibroblasts (Lee et al., 2013; Liu et al., 2016). However, we did serum starvation to stop proliferation. In our study, we observed a few percentage of Ki67-positive cells under TGF-β treatment at 24 hours (Supplementary Figure S2C). However, cell proliferation mainly stopped after 48 hours. Typically, proliferating cells rarely display any PC or show very small puncta. In our case, we observe a significantly elongated PC structure (although shorter than that of untreated cells) under TGF-beta-treated conditions. Our results display that a majority of cells are not proliferating but still display PC shortening under TGF-β treatment, suggesting that PC shortening is not due to cell division-induced PC disassembly. TGF beta-induced PC shortening is also reported in another fibroblast type previously (Kawasaki et al., 2024).
Kawasaki, Makiri, et al. "Primary cilia suppress the fibrotic activity of atrial fibroblasts from patients with atrial fibrillation in vitro." Scientific Reports 14.1 (2024): 12470.
Lee, J., Choi, JH. & Joo, CK. TGF-β1 regulates cell fate during epithelial–mesenchymal transition by upregulating survivin. Cell Death Dis 4, e714 (2013). https://doi.org/10.1038/cddis.2013.244.
Liu, Y. et al. TGF-β1 promotes scar fibroblasts proliferation and transdifferentiation via up-regulating MicroRNA-21. Sci. Rep. 6, 32231; doi: 10.1038/srep32231 (2016).
The authors described that they focused on the genes that are affected in opposite ways (supp table 4), but TEAD2, MICALL1, and HDAC6 are not listed in that table. Response: The list in Supplementary Table S3 includes common genes defined as differentially expressed based on a fold change >1 or Minor comments:
Figure 1A,B,C should also show lower magnification images where several cells/field are visualized. Response: We have replaced it with a new Figure 1.
The number of patients analyzed is not clear. For example, M&M describes 5 healthy and 8 SSc, but only 3 and 4 are shown in the figure. Furthermore, for orbital fibrosis, 2 healthy vs. 2 TAO are mentioned in the figure legend, but only one of each showed. Finally, the healthy control for lung fibroblast seems to be 3 independent experiments of the CCL210 cell line; please show the three independent controls and clarify on the X-axis and in the figure legend that these are CCL210 cells. Response: A total of 5 healthy and 8 SSc skin explanted fibroblast cell lines were used, as described in the Materials and Methods. Since these are patient-derived skin fibroblasts, maintaining equal numbers in each experiment is challenging. Revised graphs for orbital fibroblasts and CCL210 have been added in the new Figures 1B and 1C.
For the same set of experiments, please clarify and consistently describe the conditions that promote PC: 12hs serum starvation as described in M&M? Or 24hs as described in the text? Or 16 as described in figure legend 1? Or 24hs as described in supp figure 2? Response: We serum-starved the cells overnight, and this is also mentioned in the manuscript.
Please confirm in figure legends and M&M that 100 cells per group were counted. Response: We measured only 100 cells per cell line in Supplementary Figure S1B. To eliminate any confusion, we have now created a superplot for cilia analysis. Each small dot represents the PC length from an individual cell, and each large dot represents the average of the small dots for one cell line. An unpaired two-tailed t-test was performed on the small dots (mean ± SD).
Figure 2 should also provide lower magnification to show several cells per field. Response: Foreskin fibroblasts treated with TGF-β1 are added in S2A.
How do you explain that the increase in length of primary cilia after siACTA2 doesn't change COL1A1? Wouldn't it be a good approach to also check by Western Blot? Response: We believe that depletion of aSMA was sufficient to reduce the PC length for the reason described earlier (Avasthi and Marshall, 2012), but was not sufficient enough to change COL1A1 level. We added the western blot in Supplementary Figure S8B.
Once more, figure 5 will benefit from low mag images. How consistent is the effect of LiCl in the cultured cells? What is the percentage of rescued cells? Response: LiCl treatment was consistent for almost all the cells (~95%) as shown below and added in S4A.
Figure 5, panels F and G need better explanation in the results text as well as in the figure legend. Response: We added now.
9) Some figures/supp figures are wrongly referenced in the text. *
__ Response:__ We carefully revised the manuscript and corrected the references.
10) Figure 6, panel A is confusing. Is it a comparison between SSC skin fibroblasts and foreskin fibroblasts? Maybe show labels on the panel.
__ Response:__ We updated the figure legend for Panel A in Figure 6.
11) Where is Figure 8 mentioned in the text?
__ Response:__ In the discussion section.
12) The work will benefit from an initial paragraph in the discussion enumerating the findings and a summary of the conclusion at the end.
Response: We agree and modified the discussion accordingly.
13) The nintedanib experiments are not described in the results section at all.
Response: All nintedanib experiments are now included in Figure S5C-F and are described in the Results section.
Significance
Reviewer #3 (Significance (Required)): Beyond the lack of in situ ciliary expression assessment, the work is exciting, and the potential implications of treating/preventing fibrosis with small molecules to modulate ciliary length could be transformative in the field. Furthermore, there are a few HDAC6 inhibitors already in clinical trials for different tumors, which increases the significance of the work.
Response: Thank you for your encouraging comments regarding the potential impact of our findings. We agree that the therapeutic implications of modulating ciliary length, particularly using small molecules such as HDAC6 inhibitors already in clinical trials, could be transformative in the context of fibrosis. We also acknowledge the importance of in situ assessment of ciliary expression and plan to incorporate such analyses in future studies to further strengthen our findings.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary:
The author's main research topic in this work is the relationship between ciliary length and the level of fibrosis. Fibrotic samples show shorter primary cilia, profibrotic treatment with TGFB decreases the ciliary length, and posterior dedifferentiation of fibroblasts shows longer cilia. Cells with a decrease of αSMA by using siACTA2 siRNA, also show increased ciliary length. Most importantly, inducing the increase of ciliary length with LiCl or Tubacin has an inverse association with fibrosis phenotypes. The modulation of primary cilia length may represent a potential therapeutic strategy for fibrosis-associated diseases.
The premise is relevant and exciting, and the methods are appropriate. The experiments partially sustain the conclusion. The results open a new potential area for studying fibrosis. The tables and figures aid in understanding the paper. The paper is clear and easy to read for a basic research specialized audience.
Major comments:
Minor comments:
Beyond the lack of in situ ciliary expression assessment, the work is exciting, and the potential implications of treating/preventing fibrosis with small molecules to modulate ciliary length could be transformative in the field. Furthermore, there are a few HDAC6 inhibitors already in clinical trials for different tumors, which increases the significance of the work.
Expertise: primary cilium functions, cell biology, cancer biology
References
v1.1 Update
The following references have been added:
Food and Drug Administration, BeiGene. TEVIMBRA (tislelizumab) prescribing information. Available: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=761380 Accessed 5/12/25
Food and Drug Administration, Astellas. VYLOY (zolbetuximab-clzb) prescribing information. Available: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=761365 Accessed 5/8/25
Food and Drug Administration, Merck & Co. KEYTRUDA (pembrolizumab) prescribing information. Available: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=125514 Accessed 1/8/25
Liu L, Woo Y, D’Apuzzo M, Melstrom L, Raoof M, Liang Y, et al. Immunotherapy-Based Neoadjuvant Treatment of Advanced Microsatellite Instability–High Gastric Cancer: A Case Series. Journal of the National Comprehensive Cancer Network. 2022;20(8):857-65.
Kelly RJ, Lee J, Bang Y-J, Almhanna K, Blum-Murphy M, Catenacci DV, et al. Safety and efficacy of durvalumab and tremelimumab alone or in combination in patients with advanced gastric and gastroesophageal junction adenocarcinoma. Clinical Cancer Research. 2020;26(4):846-54.
Moehler M, Oh DY, Kato K, Arkenau T, Tabernero J, Lee KW, Rha SY, Hirano H, Spigel D, Yamaguchi K, Wyrwicz L. First-Line Tislelizumab Plus Chemotherapy for Advanced Gastric Cancer with Programmed Death-Ligand 1 Expression≥ 1%: A Retrospective Analysis of RATIONALE-305. Advances in Therapy. 2025 Mar 13:1-21.
Shitara K, Lordick F, Bang YJ, Enzinger P, Ilson D, Shah MA, Van Cutsem E, Xu RH, Aprile G, Xu J, Chao J. Zolbetuximab plus mFOLFOX6 in patients with CLDN18. 2-positive, HER2-negative, untreated, locally advanced unresectable or metastatic gastric or gastro-oesophageal junction adenocarcinoma (SPOTLIGHT): a multicentre, randomised, double-blind, phase 3 trial. The Lancet. 2023 May 20;401(10389):1655-68.
Rha SY, Oh D-Y, Yañez P, Bai Y, Ryu M-H, Lee J, et al. Pembrolizumab plus chemotherapy versus placebo plus chemotherapy for HER2-negative advanced gastric cancer (KEYNOTE-859): a multicentre, randomised, double-blind, phase 3 trial. The Lancet Oncology. 2023;24(11):1181-95.
Shen L, Kato K, Kim S-B, Ajani JA, Zhao K, He Z, et al. Tislelizumab versus chemotherapy as second-line treatment for advanced or metastatic esophageal squamous cell carcinoma (RATIONALE-302): a randomized phase III study. Journal of clinical oncology. 2022;40(26):3065-76
Janjigian YY, Kawazoe A, Bai Y, Xu J, Lonardi S, Metges JP, et al. Pembrolizumab plus trastuzumab and chemotherapy for HER2-positive gastric or gastro-oesophageal junction adenocarcinoma: interim analyses from the phase 3 KEYNOTE-811 randomised placebo-controlled trial. The Lancet. 2023;402(10418):2197-208
European Medicines Agency. KEYTRUDA (pembrolizumab) product information . Available: https://www.ema.europa.eu/en/medicines/human/EPAR/keytruda#authorisation-details Accessed 1/8/25
Kelley RK, Ueno M, Yoo C, Finn RS, Furuse J, Ren Z, et al. Pembrolizumab in combination with gemcitabine and cisplatin compared with gemcitabine and cisplatin alone for patients with advanced biliary tract cancer (KEYNOTE-966): a randomised, double-blind, placebo-controlled, phase 3 trial. The Lancet. 2023;401(10391):1853-65
Andre T, Elez E, Van Cutsem E, Jensen LH, Bennouna J, Mendez G, Schenker M, de la Fouchardiere C, Limon ML, Yoshino T, Li J. Nivolumab plus Ipilimumab in Microsatellite-Instability–High Metastatic Colorectal Cancer. New England Journal of Medicine. 2024 Nov 28;391(21):2014-26.
Food and Drug Administration, Bristol Myers Squibb Company. OPDIVO (nivolumab) prescribing information. Available: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=125554 Accessed 5/14/25
Food and Drug Administration, Bristol Myers Squibb Company. YERVOY (ipilimumab) prescribing information. Available: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=125377 Accessed 5/14/25
Figure 1
v1.1 Update
Figure 1 - Advanced esophagogastric diagnostic testing and treatment algorithm has been updated to include new FDA approvals and indication changes since guideline publication. See here for revised Figure 1A and Figure 1B [Ref 229, 230, 231].
Towards a personal data vault society: an interplaybetween technological and business perspectives
inelegíveis
Em princípio, a inelegibilidade ocorre apenas quanto ao cônjuge e parentes de chefes do Poder Executivo, a saber: Presidente da República, Governador de Estado ou do Distrito Federal e Prefeito.
Não alcança os do vice, tampouco alcança os parentes de quem exerce de modo interino e precário a chefia do Poder Executivo – como pode ocorrer, por exemplo, com o presidente do Poder Legislativo e do Judiciário.
[...]
(GOMES, José J. Direito Eleitoral - 20ª Edição 2024. 20. ed. Rio de Janeiro: Atlas, 2024. E-book. p.207. ISBN 9786559776054. Acesso em: 30 mai. 2025.)
Reviewer #2 (Public review):
This study aims to elucidate the role of fibroblasts in regulating myocardium and vascular development through signaling to cardiomyocytes and endothelial cells. This focus is significant, given that fibroblasts, cardiomyocytes, and vascular endothelial cells are the three primary cell types in the heart. The authors employed a Pdgfra-CreER-controlled diphtheria toxin A (DTA) system to ablate fibroblasts at various embryonic and postnatal stages, characterizing the resulting cardiac defects, particularly in myocardium and vasculature development. Single-cell RNA sequencing (scRNA-seq) analysis of the ablated hearts identified collagen as a crucial signaling molecule from fibroblasts that influences the development of cardiomyocytes and vascular endothelial cells.
This is an interesting manuscript; however, there are several major issues, including an over-reliance on the scRNA-seq data, which shows inconsistencies between replicates.
Some of the major issues are described below.
(1) The CD31 immunostaining data (Figure 3B-G) indicate a reduction in endothelial cell numbers following fibroblast deletion using PdgfraCreER+/-; RosaDTA+/- mice. However, the scRNA-seq data show no percentage change in the endothelial cell population (Figure 4D). Furthermore, while the percentage of Vas_ECs decreased in ablated samples at E16.5, the results at E18.5 were inconsistent, showing an increase in one replicate and a decrease in another, raising concerns about the reliability of the RNA-seq findings.
(2) Similarly, while the percentage of Ven_CMs increased at E18.5, it exhibited differing trends at E16.5 (Fig. 4E), further highlighting the inconsistency of the scRNA-seq analysis with the other data.
(3) Furthermore, the authors noted that the ablated samples had slightly higher percentages of cardiomyocytes in the G1 phase compared to controls (Fig. 4H, S11D), which aligns with the enrichment of pathways related to heart development, sarcomere organization, heart tube morphogenesis, and cell proliferation. However, it is unclear how this correlates with heart development, given that the hearts of ablated mice are significantly smaller than those of controls (Figure 3E). Additionally, the heart sections from ablated samples used for CD31/DAPI staining in Figure 3F appear much larger than those of the controls, raising further inconsistencies in the manuscript.
(4) The manuscript relies heavily on the scRNA-seq dataset, which shows inconsistencies between the two replicates. Furthermore, the morphological and histological analyses do not align with the scRNA-seq findings.
(5) There is a lack of mechanistic insight into how collagen, as a key signaling molecule from fibroblasts, affects the development of cardiomyocytes and vascular endothelial cells.
(6) In Figure 1B, Col1a1 expression is observed in the epicardial cells (Figure 1A, E11.5), but this is not represented in the accompanying cartoon.
(7) Do the PdgfraCreER+/-; RosaDTA+/- mice survive after birth when induced at E15.5, and do they exhibit any cardiac defects?
Ce processus illustre comment la (re-)médiation d’une archive peut donner naissance à une œuvre nouvelle
Certains concepts sont mobilisés à plusieurs reprises, une reformulation ou une meilleure articulation de ces notions pourrait peut-être améliorer la lisibilité et l'efficacité argumentative.
Leur construction peut être réalisée sous l’angle du montage car la (re-)médiation, d’une certaine manière, relève aussi de la notion de montage – concept très cher à Walter Benjamin – c’est-à-dire « l’élément monté interrompt l’enchaînement dans lequel il est monté » (Benjamin 2003, 139‑40).
Le recours au montage comme inspiration méthodologique est particulièrement stimulant. Je me demande si le paragraph pourrait se relier à celui d'après . L'idée est recurrente et peut se complementer des plusieurs aspects mentionnés ou cités.
Dans cet article, nous nous intéressons à la manière dont les billets, en tant que supports dits « traditionnels », peuvent être lus à la fois de manière individuelle et collective.
La phrase donne l'impression d'un décortiquement du format "billet" dans l'article.
celle-ci
On parle de la gestion qui peut être remédiatisée ou de l'archive. Dans le cas écheant je préciserai pour éviter la confusion. ( Cette dernière/ l'archive...)
Author response:
The following is the authors’ response to the original reviews
Public Reviews:
Reviewer #1 (Public Review):
Summary:
Zhao and colleagues employ Drosophila nephrocytes as a model to investigate the effects of a high-fat diet on these podocyte-like cells. Through a highly focused analysis, they initially confirm previous research in their hands demonstrating impaired nephrocyte function and move on to observe the mislocalization of a slit diaphragmassociated protein (pyd). Employing a reporter construct, they identify the activation of the JAK/STAT signaling pathway in nephrocytes. Subsequently, the authors demonstrate the involvement of this pathway in nephrocyte function from multiple angles, using a gain-of-function construct, silencing of an inhibitor, and ectopic overexpression of a ligand. Silencing the effector Stat92E via RNAi or inhibiting JAK/ STAT with Methotrexate effectively restored impaired nephrocyte function induced by a high-fat diet, while showing no impact under normal dietary conditions.
Strengths:
The findings establish a link between JAK/STAT activity and the impact of a high-fat diet on nephrocytes. This nicely underscores the importance of organ crosstalk for nephrocytes and supports a potential role for JAK/STAT in diabetic nephropathy, as previously suggested by other models.
Weaknesses:
The analysis is overly reliant on tracer endocytosis and single lines. Immunofluorescence of slit diaphragm proteins would provide a more specific assessment of the phenotypes.
We thank the reviewer for the positive comments and pointing out that slit diaphragm markers would provide a more specific assessment of the phenotypes. In our revised manuscript, we used Sns-mRuby3, in which mRuby3 was tagged endogenously at the C-terminal of Sns (PMID: 39195240 and PMID: 39431457), to show the slit diaphragm pattern.
Reviewer #2 (Public Review):
Summary:
In their manuscript, Zhao et al. describe a link between JAK-STAT pathway activation in nephrocytes on a high-fat diet. Nephrocytes are the homologs to mammalian podocytes and it has been previously shown, that metabolic syndrome and obesity are associated with worse outcomes for chronic kidney disease. A study from 2021 (Lubojemska et al.) could already confirm a severe nephrocyte phenotype upon feeding Drosophila a high-fat diet and also linking lipid overflow by expressing adipose triglyceride lipase in the fat body to nephrocyte dysfunction. In this study, the authors identified a second pathway and mechanism, how lipid dysregulation impact on nephrocyte function. In detail, they show activation of JAK-STAT signaling in nephrocytes upon feeding them a high-fat diet, which was induced by Upd2 expression (a leptin-like hormone) in the fat body, and the adipose tissue in Drosophila. Further, they could show genetic and pharmacological interventions can reduce JAK-STAT activation and thereby prevent the nephrocyte phenotype in the high-fat diet model.
Strengths:
The strength of this study is the combination of genetic tools and pharmacological intervention to confirm a mechanistic link between the fat body/adipose tissue and nephrocytes. Inter-organ communication is crucial in the development of several diseases, but the underlying mechanisms are only poorly understood. Using Drosophila, it is possible to investigate several players of one pathway, here JAK-STAT. This was done, by investigating the functional role of Hop, Socs36E, and Stat92E in nephrocytes and has also been combined with feeding a high-fat diet, to assess restoration of nephrocyte function by inhibiting JAK-STAT signaling. Adding a translational approach was done by inhibiting JAK-STAT signaling with methotrexate, which also resulted in attenuated nephrocyte dysfunction. Expression of the leptin-like hormone upd2 in the fat body is a good approach to studying inter-organ communication and the impact of other organs/tissue on nephrocyte function and expands their findings from nephrocyte function towards whole animal physiology.
Weaknesses:
Although the general findings of this study are of great interest, there are some weaknesses in the study, which should be addressed. Overall, the number of flies investigated for the majority of the experiments is very low (6 flies) and it is not clear whether the flies used, are from independent experiments to exclude problems with food/diet. For the analysis, the mean values of flies should be calculated, as one fly can be considered a biological replicate, but not all individual cells. By increasing the number of flies investigated, statistical analysis will become more solid. In addition, the morphological assessment is rather preliminary, by only using a Pyd antibody. Duf or Sns should be visualized as well, also the investigation of the different transgenic fly strains studying the importance of JAK-STAT signaling in nephrocytes needs to include a morphological assessment. Moreover, the expected effect of feeding a high-fat diet on nephrocytes needs to be shown (e.g. by lipid droplet formation) and whether upd2 is actually increased here should also be assessed. The time points of assessment vary between 1, 3, and 7 days and should be consistent throughout the study or the authors should describe why they use different time points.
We thank the reviewer for the comments and suggestions. HFD causes enlarged crop (Liao et al, 2021, PMID: 33171202) and accumulation of lipid droplets in the intestine. To exclude the problems with different batches of food/diet, we checked crop and the intestine during the sample preparation as indications of food consistency.
We followed the suggestion to take the mean values of flies in the data analysis, one was considered a biological replicate in the revised version. We added in another slit diaphragm protein reporter Sns-mRuby3, in which mRuby3 fluorescent protein was tagged at the C-terminal of endogenous Sns. This reporter was used to show the effect of HFD on slit diaphragm protein, manipulation of Jak/Stat pathway (ppl-Gal4>upd2 and dot-Gal4>UAS-Stat92E-RNAi), and drug treatment.
Lubojemska et al 2021 (PMID: 33945525) showed that HFD leads to lipid droplet accumulation in larval nephrocytes. Following the reviewer’s suggestion, we stained the adult nephrocytes with Nile red and found lipid droplet formation caused by HFD, verifying the HFD effects on lipid droplet accumulation.
Regarding the timepoints, the newly eclosed flies (1-day old) were treated for 7 days (transferred to fresh diet or shifted from 18 to 29 °C for 7 days to induce target gene expression). Thus, the flies were 7 days old. In the revised manuscript, we changed “1-day-old females” to “7-day-old females” in the figure legend. The exception was Figure 4 panel G and H, we used Day 3 for the UAS-hop.Tum overexpression in the flp-out clones, which is different from the HFD approach (Day 7). This is because Hop.Tum is a strong gain of function mutation. UAS-hop.Tum overexpression in the eye imaginal disc leads to apoptosis via up-regulating a proapoptotic gene hid (Bhawana Maurya et al, 2021, PMID: 33824299). Thus, we used Day 3 for this experiment.
Recommendations for the authors:
Reviewer #1 (Recommendations For The Authors):
There are relevant issues, that should be addressed:
Major:
- The analysis of JAK/STAT signaling in nephrocytes is limited to nephrocyte function, despite the nice slit diaphragm phenotype shown in Figure 2A. What happens to the slit diaphragm in the other genotypes, the rescue settings in particular? Immunofluorescence of Pyd should be explored for all conditions to evaluate proper phenocopy. Tracer endocytosis is much less specific.
We thank the reviewer for the suggestion. We made a transgenic line Sns-mRuby3, in which mRuby3 was tagged to the endogenous Sns C-terminal. It has been used as a slit diaphragm reporter (PMID: 39195240 and PMID: 39431457). Apart from the tracer assays, we used Sns-mRuby3 reporter and/or Pyd staining to visualize the changes in slit-diaphragm structures.
- The interventions are restricted to single RNAi lines and reporters, raising concerns about specificity/potential off-targets. Additional lines should be tested for verification.
Different versions of RNAi lines are available for targeting fly genes. For UAS-Socs36E-RNAi, we chose the one that was generated with a short hairpin, which is known to restrict the off-target effects (Ni et al, 2011, PMID: 21460824). For UAS-Stat92E-RNAi, we added in an independent RNAi line (Figure 6 - figure supplement 1 and 2).
Minor:
- In Figure 2C, the image of HFD shows a section that cuts through the surface at a shallower angle, making everything appear blurry. This image should be replaced.
We replaced Figure 2C (the image of HFD) with another one.
- What is the relevance (if any) of reduced electrodense vacuoles with a high-fat diet? An effect on endocytic trafficking/endosome architecture remains unexplored.
Lubojemska et al (PMID: 33945525) studied the endocytic trafficking/endosome architecture of the larval nephrocytes and found that HFD impaired the endocytosis. We studied the adult pericardial nephrocytes. It is very likely that the endocytic trafficking/endosome architecture is affected by HFD in the adult nephrocytes.
- How do the findings presented in this manuscript correlate with a similar study by Lubojemska et al.? At least the discussion should provide more evaluation of this aspect.
Lubojemska et al (PMID: 33945525) assayed the larval nephrocytes and found that a HFD leads to the ectopic accumulation of lipid droplets in the nephrocytes and decreased endocytosis. They further demonstrated that lipid droplet lipolysis and PGC1α counteracts the harmful effects of a HFD. We performed Nile red staining and verified the accumulation of lipid droplets in the adult pericardial nephrocytes upon HFD feeding, which agrees with Lubojemska discovery. We found that a HFD activates Jak/Stat pathway, which mediates the nephrocyte functional defects. A previous study showed that Stat1 has an inhibitory effect on PGC1α transcription (PMID: 26689548). Further study is needed to investigate the interaction between Jak/Stat pathway and PGC1α transcription. We added the information to the discussion.
- Please check spelling and grammar.
Reviewer #2 (Recommendations For The Authors):
(1) Which cells are investigated? Please state.
Pericardial nephrocytes were used in this study. The information was added to the result parts.
(2) Rephrase 'chronic kidney disease model'. Feeding for 7 days and assessment after 7 days cannot be considered chronic as flies can live more than 60 days.
Lubojemska et al (PMID: 33945525) fed the newly hatched larvae with a HFD and used the third instar larvae for the experiments. The term “chronic kidney disease” has been used in the reference PMID: 33945525. It takes about 4 days for fly larvae to develop from the first instar to the third instar. Thus, the animals were fed on the HFD for only 4 days. In this regard, feeding for seven days might be considered as chronic.
(3) Line 89: Curran et al., 2014). with risk increasing risk as BMI increases (Hsu et al., 2006). Please correct this sentence.
We thank the reviewer for finding the error. In the revised version, the sentence was changed as “with increasing risk as BMI increases (Hsu et al., 2006)”.
(4) Figure 1: The authors should explain why they use FITC-Albumin and 10kDA dextran, what are the differences, and why are both used?
The tracers are different in size (70kD FITC-Albumin and 10kDA dextran). Both FITC-Albumin and 10kDA dextran have been used in previous publications (Zhao et al 2024, PMID: 39431457 and Weavers et al 2009, PMID: 18971929) to show that the nephrocytes can efficiently take up the tracers of different sizes.
(5) Figure 3: The JAK-STAT sensor was used on Day 1 to confirm activation of JAKSTAT signaling, which means a very fast response towards the HFD after 24hrs. How is the activation after 7 days? The nephrocyte assessment in Figures 1 and 2 is done at the later time point, how about earlier time points in HFD? One would expect an earlier phenotype as well if JAK-STAT signaling is causative.
In Figure 3C, newly eclosed flies (1-day old) were fed on a control diet or a HFD for 7 days. Thus, in the legend it shall be “7-day-old females”. Sorry for misleading. The caption was updated as “7-day-old females”.
(6) Figure 4H: I don't understand how many cells or flies are depicted and analysed? Are the dots one nephrocyte from 4 flies? If yes, the numbers need to be increased.
In figure 4H, we quantified 5 UAS-hop.Tum clones and 5 neighbor cells. We only found 5 clones from 4 flies. We didn’t quantify all the nephrocytes, since we compared the clone with its neighbor cell. To make it easier to follow, we changed the description as “n= 5 clones and 5 neighbor cells”.
(7) Figure 4: Why are flies investigated at different ages? Day 1 vs Day 3? This should be consistent with the HFD approach and day 7. Or investigate the HFD at earlier time points as well.
In Figure 4, the newly eclosed flies (1-day old) were shifted from 18 to 29 °C for 7 days to induce target gene expression. Thus, the flies were 7-day old. In the revised manuscript, we changed “1-day-old females” to “7-day-old females” in the figure legend. We used Day 3 for the UAS-hop.Tum overexpression in the flp-out clones, which is different from the HFD approach (Day 7). This is because Hop.Tum is a strong gain of function mutation. UAS-hop.Tum overexpression in the eye imaginal disc leads to apoptosis via up-regulating a proapoptotic gene hid (Bhawana Maurya et al, 2021, PMID: 33824299). Thus, we used Day 3 for this experiment.
(8) Figure 5: Do the authors see upd2-GFP in the nephrocyte or at the nephrocyte? Is upd2 filtered to bind the JAK-STAT-receptor? They should show this, which is easy to do due to the GFP label.
We thank the reviewer for the suggestion. We looked into the nephrocyte from ppl-Gal4>upd2-GFP flies and found Upd2-GFP in the nephrocytes. We further showed that ppl-Gal4 was not expressed in the nephrocytes, suggesting that Upd2-GFP is secreted from the fat body and transported to the nephrocytes. We stained the nephrocytes for Pyd and found compromised fingerprint pattern caused by Upd2-GFP expression in the fat body. The data was added to Figure 5 - figure supplement 1.
(9) Figure 5: What are the upd2 levels after day 1 and compared to HFD at day 7? In the Rajan et al manuscript, upd2 levels have been assessed by qPCR, this can be done here as well. Although there is a mechanistic link shown here, I think it would be interesting to test the upd2 levels at the different time points assessed.
In the Rajan et al manuscript, they showed that the expression of upd2 was up regulated by HFD. My previous work showed that HFD changes taste perception. We performed qPCR to determine the expression of upd2 and verified that upd2 was upregulated in HFD fed flies (Yunpo Zhao et al. 2023. PMID: 37934669). We included the reference in the revised version.
(10) Figure 6: Does a Socs36E overexpression e.g. with the Bloomington strain 91352 also rescue the HFD phenotype, by blocking JAK-STAT signaling?
We thank the reviewer for the suggestion. We tested the effect of Socs36E overexpression and observed that UAS-Socs36E can partially rescue HFD caused nephrocyte functional decline. The data was not included in the revised manuscript. Notably, apart from having an inhibitory effect on the Jak/Stat, Socs36E represses MAPK pathway (Amoyel et al, 2016, PMID: 26807580).
(11) Figure 7: What is the control for the methotrexate treatment? What is the solvent?
We used DMSO as the solvent for methotrexate and used it as the control for the methotrexate treatment. We added the following sentences to the method parts, “Methotrexate (06563, Sigma-Aldrich, MO) was dissolved in DMSO to make a 10mM stock solution”, and “The samples incubated in Schneider’s Medium supplemented with DMSO vehicle were used a control”.
(12) Why did the authors use Dot-Gal4 for the Socs36E knockdown and Dot-Gal4ts for the Stat92E knockdown?
We used Dot-Gal4ts and temperature shifting to restrict the Stat92E knockdown at adult stages.
(13) Supplementary Figure 1: Please add the individual data to the figure as done for all other figures.
We thank the reviewer for this comment. The figure individual data was added according to the suggestion.
Reviewer #1 (Public review):
Summary:
In this study, authors explored how galanin affects whole-brain activity in larval zebrafish using wide-field Ca2+ imaging, genetic modifications, and drugs that increase brain activity. The authors conclude that galanin has a sedative effect on the brain under normal conditions and during seizures, mainly through the galanin receptor 1a (galr1a). However, acute "stressors(?)" like pentylenetetrazole (PTZ) reduce galanin's effects, leading to increased brain activity and more seizures. Authors claim that galanin can reduce seizure severity while increasing seizure occurrence, speculated to occur through different receptor subtypes. This study confirms galanin's complex role in brain activity, supporting its potential impact on epilepsy.
Strengths:
The overall strength of the study lies primarily in its methodological approach using whole-brain Calcium imaging facilitated by the transparency of zebrafish larvae. Additionally, the use of transgenic zebrafish models is an advantage, as it enables genetic manipulations to investigate specific aspects of galanin signaling. This combination of advanced imaging and genetic tools allows for addressing galanin's role in regulating brain activity.
Weaknesses:
The weaknesses of the study also stem from the methodological approach, particularly the use of whole-brain Calcium imaging as a measure of brain activity. While epilepsy and seizures involve network interactions, they typically do not originate across the entire brain simultaneously. Seizures often begin in specific regions or even within specific populations of neurons within those regions. Therefore, a whole-brain approach, especially with Calcium imaging with inherited limitations, may not fully capture the localized nature of seizure initiation and propagation, potentially limiting the understanding of Galanin's role in epilepsy.
Furthermore, Galanin's effects may vary across different brain areas, likely influenced by the predominant receptor types expressed in those regions. Additionally, the use of PTZ as a "stressor" is questionable since PTZ induces seizures rather than conventional stress. Referring to seizures induced by PTZ as "stress" might be a misinterpretation intended to fit the proposed model of stress regulation by receptors other than Galanin receptor 1 (GalR1).
The description of the EAAT2 mutants is missing crucial details. EAAT2 plays a significant role in the uptake of glutamate from the synaptic cleft, thereby regulating excitatory neurotransmission and preventing excitotoxicity. Authors suggest that in EAAT2 knockout (KO) mice galanin expression is upregulated 15-fold compared to wild-type (WT) mice, which could be interpreted as galanin playing a role in the hypoactivity observed in these animals.
However, the study does not explore the misregulation of other genes that could be contributing to the observed phenotype. For instance, if AMPA receptors are significantly downregulated, or if there are alterations in other genes critical for brain activity, these changes could be more important than the upregulation of galanin. The lack of wider gene expression analysis leaves open the possibility that the observed hypoactivity could be due to factors other than, or in addition to, galanin upregulation.
Moreover, the observation that in double KO mice for both EAAT2 and galanin there was little difference in seizure susceptibility compared to EAAT2 KO mice alone further supports the idea that galanin upregulation might not be the reason to the observed phenotype. This indicates that other regulatory mechanisms or gene expressions might be playing a more pivotal role in the manifestation of hypoactivity in EAAT2 mutants.
These methodological shortcomings and conceptual inconsistencies undermine the perceived strengths of the study, and hinders understanding of Galanin's role in epilepsy and stress regulation.
Comments on revisions:
The revised manuscript and the answers of the authors is appreciated. However, the criticisms were addressed only partially and main weaknesses of the manuscript are still remaining.
Author response:
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public Review):
Summary:
In this study, the authors explored how galanin affects whole-brain activity in larval zebrafish using wide-field Ca2+ imaging, genetic modifications, and drugs that increase brain activity. The authors conclude that galanin has a sedative effect on the brain under normal conditions and during seizures, mainly through the galanin receptor 1a (galr1a). However, acute "stressors(?)" like pentylenetetrazole (PTZ) reduce galanin's effects, leading to increased brain activity and more seizures. The authors claim that galanin can reduce seizure severity while increasing seizure occurrence, speculated to occur through different receptor subtypes. This study confirms galanin's complex role in brain activity, supporting its potential impact on epilepsy.
Strengths:
The overall strength of the study lies primarily in its methodological approach using whole-brain Calcium imaging facilitated by the transparency of zebrafish larvae. Additionally, the use of transgenic zebrafish models is an advantage, as it enables genetic manipulations to investigate specific aspects of galanin signaling. This combination of advanced imaging and genetic tools allows for addressing galanin's role in regulating brain activity.
Weaknesses:
The weaknesses of the study also stem from the methodological approach, particularly the use of whole-brain Calcium imaging as a measure of brain activity. While epilepsy and seizures involve network interactions, they typically do not originate across the entire brain simultaneously. Seizures often begin in specific regions or even within specific populations of neurons within those regions. Therefore, a whole-brain approach, especially with Calcium imaging with inherited limitations, may not fully capture the localized nature of seizure initiation and propagation, potentially limiting the understanding of Galanin's role in epilepsy.
Furthermore, Galanin's effects may vary across different brain areas, likely influenced by the predominant receptor types expressed in those regions. Additionally, the use of PTZ as a "stressor" is questionable since PTZ induces seizures rather than conventional stress. Referring to seizures induced by PTZ as "stress" might be a misinterpretation intended to fit the proposed model of stress regulation by receptors other than Galanin receptor 1 (GalR1).
The description of the EAAT2 mutants is missing crucial details. EAAT2 plays a significant role in the uptake of glutamate from the synaptic cleft, thereby regulating excitatory neurotransmission and preventing excitotoxicity. Authors suggest that in EAAT2 knockout (KO) mice galanin expression is upregulated 15-fold compared to wild-type (WT) mice, which could be interpreted as galanin playing a role in the hypoactivity observed in these animals.
Indeed, our observation of the unexpected hypoactivity in EAAT2a mutants, described in our description of this mutant (Hotz et al., 2022), prompted us to initiate this study formulating the hypothesis that the observed upregulation of galanin is a neuroprotective response to epilepsy.
However, the study does not explore the misregulation of other genes that could be contributing to the observed phenotype. For instance, if AMPA receptors are significantly downregulated, or if there are alterations in other genes critical for brain activity, these changes could be more important than the upregulation of galanin. The lack of wider gene expression analysis leaves open the possibility that the observed hypoactivity could be due to factors other than, or in addition to, galanin upregulation.
We have performed a transcriptome analysis that we are still evaluation. We can already state that AMPA receptor genes are not significantly altered in the mutant.
Moreover, the observation that in double KO mice for both EAAT2 and galanin, there was little difference in seizure susceptibility compared to EAAT2 KO mice alone further supports the idea that galanin upregulation might not be the reason for the observed phenotype. This indicates that other regulatory mechanisms or gene expressions might be playing a more pivotal role in the manifestation of hypoactivity in EAAT2 mutants.
We agree that upregulation of galanin transcripts is at best one of a suite of regulatory mechanisms that lead to hypoactivity in EAAT2 zebrafish mutants.
These methodological shortcomings and conceptual inconsistencies undermine the perceived strengths of the study, and hinders understanding of Galanin's role in epilepsy and stress regulation.
Reviewer #2 (Public Review):
Summary:
This study is an investigation of galanin and galanin receptor signaling on whole-brain activity in the context of recurrent seizure activity or under homeostatic basal conditions. The authors primarily use calcium imaging to observe whole-brain neuronal activity accompanied by galanin qPCR to determine how manipulations of galanin or the galr1a receptor affect the activity of the whole-brain under non-ictal or seizure event conditions. The authors' Eaat2a-/- model (introduced in their Glia 2022 paper, PMID 34716961) that shows recurrent seizure activity alongside suppression of neuronal activity and locomotion in the time periods lacking seizures is used in this paper in comparison to the well-known pentylenetetrazole (PTZ) pharmacological model of epilepsy in zebrafish. Given the literature cited in their Introduction, the authors reasonably hypothesize that galanin will exert a net inhibitory effect on brain activity in models of epilepsy and at homeostatic baseline, but were surprised to find that this hypothesis was only moderately supported in their Eaat2a-/- model. In contrast, under PTZ challenge, fish with galanin overexpression showed increased seizure number and reduced duration while fish with galanin KO showed reduced seizure number and increased duration. These results would have been greatly enriched by the inclusion of behavioral analyses of seizure activity and locomotion (similar to the authors' 2022 Glia paper and/or PMIDs 15730879, 24002024). In addition, the authors have not accounted for sex as a biological variable, though they did note that sex sorting zebrafish larvae precludes sex selection at the younger ages used. It would be helpful to include smaller experiments taken from pilot experiments in older, sex-balanced groups of the relevant zebrafish to increase confidence in the findings' robustness across sexes. A possible major caveat is that all of the various genetic manipulations are non-conditional as performed, meaning that developmental impacts of galanin overexpression or galanin or galr1a knockout on the observed results have not been controlled for and may have had a confounding influence on the authors' findings. Overall, this study is important and solid (yet limited), and carries clear value for understanding the multifaceted functions that neuronal galanin can have under homeostatic and disease conditions.
Strengths:
- The authors convincingly show that galanin is upregulated across multiple contexts that feature seizure activity or hyperexcitability in zebrafish, and appears to reduce neuronal activity overall, with key identified exceptions (PTZ model).
- The authors use both genetic and pharmacological models to answer their question, and through this diverse approach, find serendipitous results that suggest novel underexplored functions of galanin and its receptors in basal and disease conditions. Their question is well-informed by the cited literature, though the authors should cite and consider their findings in the context of Mazarati et al., 1998 (PMID:982276). The authors' Discussion places their findings in context, allowing for multiple interpretations and suggesting some convincing explanations.
- Sample sizes are robust and the methods used are well-characterized, with a few exceptions (as the paper is currently written).
- Use of a glutamatergic signaling-based genetic model of epilepsy (Eaat2a-/-) is likely the most appropriate selection to test how galanin signaling can alter seizure activity, as galanin is known to reduce glutamatergic release as an inhibitory mechanism in rodent hippocampal neurons via GalR1a (alongside GIRK activation effects). Given that PTZ instead acts through GABAergic signaling pathways, it is reasonable and useful to note that their glutamate-based genetic model showed different effects than did their GABAergic-based model of seizure activity.
Weaknesses:
- The authors do not include behavioral assessments of seizure or locomotor activity that would be expected in this paper given their characterizations of their Eaat2a-/- model in the Glia 2022 paper that showed these behavioral data for this zebrafish model. These data would inform the reader of the behavioral phenotypes to expect under the various conditions and would likely further support the authors' findings if obtained and reported.<br />
We agree that a thorough behavioral assessment would have strengthened the study, but we deemed it outside of the scope of this study.
- No assessment of sex as a biological variable is included, though it is understood that these specific studied ages of the larvae may preclude sex sorting for experimental balancing as stated by the authors.
The study was done on larval zebrafish (5 days post fertilization). The first signs of sexual differentiation become apparent at about 17 days post fertilization (reviewed in Ye and Chen, 2020). Hence sex is no biological variable at the stage studied.
- The reported results may have been influenced by the loss or overexpression of galanin or loss of galr1a during developmental stages. The authors did attempt to use the hsp70l system to overexpress galanin, but noted that the heat shock induction step led to reduced brain activity on its own (Supplementary Figure 1). Their hsp70l:gal model shows galanin overexpression anyways (8x fold) regardless of heat induction, so this model is still useful as a way to overexpress galanin, but it should be noted that this galanin overexpression is not restricted to post-developmental timepoints and is present during development.
The developmental perspective is an important point to consider. Due to the rapid development of the zebrafish it is not trivial to untangle this. In the zebrafish we first observe epileptic seizures as early as 3 days post fertilization (dpf), where the brain is clearly not well developed yet (e.g. behaviroal response to light are still minimal). Even the 5 dpf stage, where most of our experiments have been conducted, cannot by far not be considered post-development.
Reviewer #3 (Public Review):
Summary:
The neuropeptide galanin is primarily expressed in the hypothalamus and has been shown to play critical roles in homeostatic functions such as arousal, sleep, stress, and brain disorders such as epilepsy. Previous work in rodents using galanin analogs and receptor-specific knockout has provided convincing evidence for the anti-convulsant effects of galanin.
In the present study, the authors sought to determine the relationship between galanin expression and whole-brain activity. The authors took advantage of the transparent nature of larval zebrafish to perform whole-brain neural activity measurements via widefield calcium imaging. Two models of seizures were used (eaat2a-/- and pentylenetetrazol; PTZ). In the eaat2a-/- model, spontaneous seizures occur and the authors found that galanin transcript levels were significantly increased and associated with a reduced frequency of calcium events. Similarly, two hours after PTZ galanin transcript levels roughly doubled and the frequency and amplitude of calcium events were reduced. The authors also used a heat shock protein line (hsp70I:gal) where galanin transcript levels are induced by activation of heat shock protein, but this line also shows higher basal transcript levels of galanin. Again, the higher level of galanin in hsp70I:gal larval zebrafish resulted in a reduction of calcium events and a reduction in the amplitude of events. In contrast, galanin knockout (gal-/-) increased calcium activity, indicated by an increased number of calcium events, but a reduction in amplitude and duration. Knockout of the galanin receptor subtype galr1a via crispants also increased the frequency of calcium events.
In subsequent experiments in eaat2a-/- mutants were crossed with hsp70I:gal or gal-/- to increase or decrease galanin expression, respectively. These experiments showed modest effects, with eaat2a-/- x gal-/- knockouts showing an increased normalized area under the curve and seizure amplitude.
Lastly, the authors attempted to study the relationship between galanin and brain activity during a PTZ challenge. The hsp70I:gal larva showed an increased number of seizures and reduced seizure duration during PTZ. In contrast, gal-/- mutants showed an increased normalized area under the curve and a stark reduction in the number of detected seizures, a reduction in seizure amplitude, but an increase in seizure duration. The authors then ruled out the role of Galr1a in modulating this effect during PTZ, since the number of seizures was unaffected, whereas the amplitude and duration of seizures were increased.
Strengths:
(1) The gain- and loss-of function galanin manipulations provided convincing evidence that galanin influences brain activity (via calcium imaging) during interictal and/or seizure-free periods. In particular, the relationship between galanin transcript levels and brain activity in Figures 1 & 2 was convincing.
(2) The authors use two models of epilepsy (eaat2a-/- and PTZ).
(3) Focus on the galanin receptor subtype galr1a provided good evidence for the important role of this receptor in controlling brain activity during interictal and/or seizure-free periods.
Weaknesses:
(1) Although the relationship between galanin and brain activity during interictal or seizure-free periods was clear, the manuscript currently lacks mechanistic insight in the role of galanin during seizure-like activity induced by PTZ.
We completely agree and concede that this study constitutes only a first attempt to understand the (at least for us) perplexing complexity of galanin function on the brain.
(2) Calcium imaging is the primary data for the paper, but there are no representative time-series images or movies of GCaMP signal in the various mutants used.
We have now added various movies in supplementary data.
(3) For Figure 3, the authors suggest that hsp70I:gal x eaat2a-/-mutants would further increase galanin transcript levels, which were hypothesized to further reduce brain activity. However, the authors failed to measure galanin transcript levels in this cross to show that galanin is actually increased more than the eaat2a-/- mutant or the hsp70I:gal mutant alone.
After a couple of unsuccessful mating attempts with our older mutants, we finally decided not to wait for a new generation to grow up, deeming the experiment not crucial (but still nice to have).
(4) Similarly, transcript levels of galanin are not provided in Figure 2 for Gal-/- mutants and galr1a KOs. Transcript levels would help validate the knockout and any potential compensatory effects of subtype-specific knockout.
To validate the gal-/- mutant line, we decided to show loss of protein expression (Suppl. Figure 2), which we deem to more relevant to argue for loss of function. Galanin transcript levels in galr1a KOs were also added into the same Figure. However, validation of the galr1a KO could not be performed due to transcript levels being close to the detection limit and lack of available antibodies.
(5) The authors very heavily rely on calcium imaging of different mutant lines. Additional methods could strengthen the data, translational relevance, and interpretation (e.g., acute pharmacology using galanin agonists or antagonists, brain or cell recordings, biochemistry, etc).
Again, we agree and concede that a number of additional approaches are needed to get more insight into the complex role of galanin in regulation overall brain activity. These include, among others, also behavioral, multiple single cell recordings and pharmacological interventions.
Recommendations for the authors:
Reviewer #2 (Recommendations For The Authors):
Minor issues:
(1) "Sedative" effect of galanin is somewhat vague and seems overapplied without the inclusion of behavioral data showing sedation effects. I would replace "sedative" with something clearer, like the phrase "net inhibitory effect" or similar.
We have modified the wording as deemed appropriate.
(2) Include new data that is sufficiently powered to detect or rule out the effects of sex as a biological variable within the various experiments.
At this stage sex is not a biological variable. Sex determination starts a late larval stage around 14dpf. Our analysis is based on 5pdf larvae.
(3) Attempt to perform some experiments with galanin/galr1a manipulations that have been induced after the majority of development without using heat shock induction if possible (unknown how feasible this is in current model systems).
In the current model this is not feasible, but an excellent suggestion for future studies that would then also address more longterm effects in the model.
(4) Figure 2 should include qPCR results for galanin or galr1a mRNA expression to match Figure 1C, F, and Figure 2C and to confirm reductions in the respective RNA transcript levels of gal or galr1a. It could be useful to perform qPCR for galanin in all galr1aKO mice to ascertain whether compensatory elevations in galanin occur in response to galr1aKO.
(5) Axes should be made with bolder lines and bolder/larger fonts for readability and consistency throughout.
Indeed, an excellent suggestion. We have adjusted the axes significantly improving the readability of the graphs.
(6) The bottom o,f the image for Figure 2 appears to have been cut off by mistake (page 5).
(7) The ending of the legend text for Figure 3 appears to have been cut off by mistake (page 6).
Both regrettable mistakes have been corrected (already in the initial posted version)
Reviewer #3 (Recommendations For The Authors):
(1) The introduction or first paragraph of the results should be revised to more directly state the hypotheses. Several critical details were only clear after reading the discussion.
We added some words to the introduction, hoping that the critical points are now more apparent to the reader.
(2) Galanin is known to be rapidly depleted by seizures (Mazarati et al., 1998; Journal of Neuroscience, PMID #9822761) but this paper did not appear to be cited or considered. Could the rapid depletion of galanin during seizures help explain the confusing effects of galanin manipulations during PTZ?
We have added a sentence and the reference to the discussion.
(3) Figure 1 panels are incorrect. For example, Panel 'F' is used twice and the figure legend is also incorrect due to the labeling errors. In-text references to the figure should also be updated accordingly.
(4) In Figure 2 N-P, the delta F/F threshold wording is partially cropped. The figure should be updated.
Thank you for pointing out this mistake. Both figures have now been updated (already in the initial posted version)
(5) The naming and labeling of groups in the manuscript and figures should be updated to more accurately reflect the fish used for each experiment. As it currently stands, I found the labeling confusing and sometimes misleading. For example, Figure 3 'controls' are actually eaat2a-/- mutants, whereas the other group is hsp70I:gal x eaat2a-/- crosses or gal-/- x eaat2a-/- crosses. In other Figures, 'controls' are eaat2a+/+larva, or wild-type siblings (sometimes unclear).
We have made appropriate changes to the manuscript to make this point clearer to the reader, especially when the controls are eaat2a mutants.
(6) Figure 4J and 4K only show 5 data points, when the authors clearly indicate that 6 fish had seizures. Continuation of this data in Figure 4L shows 6 data points.
Indeed the 6 data points in Figure 4J and K are hard to see due to their nearly complete overlap. On larger magnification all six data points become distinguishable. We will try some different plotting approaches for the revision.
RRID:AB_491015
DOI: 10.7554/eLife.105018
Resource: (Cell Signaling Technology Cat# 9516, RRID:AB_491015)
Curator: @scibot
SciCrunch record: RRID:AB_491015
RRID:AB_2313606
DOI: 10.1523/JNEUROSCI.2325-24.2025
Resource: (Vector Laboratories Cat# BA-1000, RRID:AB_2313606)
Curator: @scibot
SciCrunch record: RRID:AB_2313606
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
Reviewer #1 (Evidence, reproducibility and clarity (Required)):
The authors describe a novel pattern of ncRNA processing by Pac1. Pac1 is a RNase III family member in S. pombe that has previously been shown to process pre-snoRNAs. Other RNase III family members, such as Rnt1 in S. cerevisiae and Dosha in human, have similar roles in cleaving precursors to ncRNAs (including miRNA, snRNA, snoRNA, rRNA). All RNAse III family members share that they recognize and cleave dsRNA regions, but differ in their exact sequence and structure requirement. snoRNAs can be processed from their own precursor, a polycistronic pre-cursor, or the intron of a snoRNA host gene. After the intron is spliced out, the snoRNA host gene can either encode an protein or be a non-functional by product.
In the current manuscript the authors show that in S. pombe snoRNA snR107 and U14 are processed from a common precursor in a way that has not previously been described. snR107 is encoded within an intron and processed from the spliced out intron, similar to a typical intron-encoded snoRNA. What is different is that upon splicing, the host gene can adopt a new secondary structure that requires base-pairing between exon 1 and exon2, generating a Pac1 recognition site. This site is recognized, resulting in cleaving of the RNA and further processing of the 3' cleavage product into U14 snoRNA. In addition, the 5' cleavage product is processed into a ncRNA named mamRNA. The experiments describing this processing are thorough and convincing, and include RNAseq, degradome sequencing, northern blotting, qRT-PCR and the analysis of mutations that disrupt various secondary structures in figures 1, 2, and 3. The authors thereby describe a previously unknown gene design where both the exon and the intron are processed into a snoRNA. They conclude that making the formation of the Pac1 binding site dependent on previous splicing ensures that both snoRNAs are produced in the correct order and amount. Some of the authors findings are further confirmed by a different pre-print (reference 19), but the other preprint did not reveal the involvement of Pac1.
While the analysis on the mamRNA/snR107/U14 precursor is convincing, as a single example the impact of these findings is uncertain. In Figure 4 and supplemental table 1, the authors use bioinformatic searches and identify other candidate loci in plans and animals that may be processed similarly. Each of these loci encode a putative precursor that results in one snoRNA processed from an intron, a different snoRNA processed from an exon, and a double stranded structure that can only form after splicing. While is potentially interesting, it is also the least developed and could be discussed and developed further as detailed below.
Major comments:
The proposal that plant and animal pre-snoRNA clusters are processed similarly is speculative. the authors provide no evidence that these precursors are processed by an RNase III enzyme cutting at the proposed splicing-dependent structure. This should not be expected for publication, but would greatly increase the interest.
All three reviewers expressed a similar concern, and we now provide additional evidence supporting the conservation of the proposed mechanism. Specifically, we focused on the SNHG25 gene in H. sapiens, which hosts two snoRNAs—one intronic, as previously shown in Figure 4B, and one non-intronic. We substantiated our predictions through the re-analysis of multiple sequencing datasets in human cell lines, as outlined below:
I. Analysis of CAGE-seq and nano-COP datasets indicates a single major transcription initiation site at the SNHG25 locus. Both the intronic and non-intronic snoRNAs are present within the same nascent precursor transcripts (Supplementary Figure 4D).
II. Degradome-seq experiments in human cell lines reveal that the predicted splicing-dependent stem-loop structure within the SNHG25 gene is subject to endonucleolytic cleavage (Supplementary Figure 4D). The cleavage sites are located at the apical loop and flanking the stem, displaying a staggered symmetry characteristic of RNase III activity (Figure 4C). Importantly, the nucleotide sequence surrounding the 3' cleavage site and the 3' splice-site are conserved in other vertebrates (Supplementary Figure 4.D).
III. fCLIP experiments demonstrate that DROSHA associates with the spliced SNHG25 transcript (Supplementary Figure 4D).
Together, these analyses support the generalizability of our model beyond fission yeast. They confirm the structure of the SNHG25 gene as a single non-coding RNA precursor hosting two snoRNAs, one of which is intronic. Importantly, these findings show that the predicted stem-loop structure contains conserved elements and is subject to endonucleolytic cleavage. Human DROSHA, an RNase III enzyme, could be responsible for this processing step.
The authors provide examples of similarly organized snoRNA clusters from human, mouse and rat, but the examples are not homologous to each other. Does this mean these snoRNA clusters are not conserved, even between mammals? Are the examples identified in Arabidopsis conserved in other plants? If there is no conservation, wouldn't that indicate that this snoRNA cluster organization offers no benefit?
We noticed during this revision that the human SNHG25 locus is actually very well conserved in mice at the GM36220 locus, where both snoRNAs (SNORD104 and SNORA50C/GM221711) are similarly arranged. Although the murine host gene, GM36220, also contains an intron in the UCSC annotation, it is intronless in the Ensembl annotation we used to screen for mixed snoRNA clusters, which explains why it was not part of our initial list of candidates (Supplementary Table 1). Importantly, sequence elements in SNHG25, close to the splice sites and cleavage sites in exon 2, are also well conserved in mice and other vertebrates (Supplementary Figure 4D). Therefore, it is reasonable to think that the mechanism described for SNHG25 in humans may also apply in mice and other vertebrates.
That being said, snoRNAs are highly mobile genetic elements. For example, it is well established that even between relatively closely related species (e.g., mouse and human), the positions of intronic snoRNAs within their host genes are not strictly conserved, even when both the snoRNAs and their host genes are. In the constrained drift model of snoRNA evolution (Hoeppner et al., BMC Evolutionary Biology, 2012; doi: 10.1186/1471-2148-12-183), it is proposed that snoRNAs are mobile and “may occupy any genomic location from which expression satisfies phenotype.”
Therefore, a low level of conservation in mixed snoRNA clusters is generally expected and does not necessarily imply that is offers no benefit. Despite the limited conservation of snoRNA identity across species, mixed snoRNA clusters consistently display two recurring features: (1) non-intronic snoRNAs often follow intronic snoRNAs, and (2) the predicted secondary structure tends to span the last exon–exon junction. These enriched features support the idea that enforcing sequential processing of mixed snoRNA clusters may confer a selective advantage. We now explicitly discuss these points in the revised manuscript.
Supplemental Figure 4 shows some evidence that the S. pombe gene organization is conserved within the Schizosaccharomyces genus, but could be enhanced further by showing what sequences/features are conserved. Presumably the U14 sequence is conserved, but snR107 is not indicated. Is it not conserved? Is the stem-loop more conserved than neighboring sequences? Are there any compensatory mutations that change the sequence but maintain the structure? Is there evidence for conservation outside the Schizosaccharomyces genus?
We thank the reviewer for these excellent suggestions, which helped us significantly improve Supplementary Figure 4. In the revised version, we now include an additional species—S. japonicus, which is more evolutionarily distant—and show that the intronic snR107 is conserved across the Schizosaccharomyces genus (Supplementary Figure 4A). The distance between conserved elements (splice sites, snoRNAs, and RNA structures) varies, indicating that surrounding sequences are less conserved compared to these functionally constrained features
We also performed a detailed alignment of the sequences corresponding to the predicted RNA secondary structures. This revealed that the apical regions are less conserved than the base, particularly near the splice and cleavage sites. In these regions, we observe compensatory or base-pair-neutral mutations (e.g., U-to-C or C-to-U, which both pair with G), suggesting structural conservation through evolutionary constraint (Supplementary Figures 4B–C). These observations are now described in greater detail in the revised manuscript, along with a discussion of the specific features likely to be under selective pressure at this locus.
Conservation outside the Schizosaccharomyces genus is less clear. As already noted in the manuscript, the S. cerevisiae locus retains synteny between snR107 and snoU14, but the polycistronic precursor encompassing both is intronless and processed by RNase III (Rnt1) between the cistrons. Similarly, in Ashbya gossypii and a few other fungal species, synteny is preserved, but no intron appears to be present in the presumed common precursor. Notably, secondary structure predictions for the A. gossypii locus (not shown) suggest the formation of a stable stem-loop encompassing the first snoRNA in a large apical loop. This could reflect a distinct mode of snoRNA maturation, possibly analogous to pri-miRNA processing, where cleavage by an RNase III enzyme contributes to both 5′ and 3′ end formation. In Candida albicans, snoU14 is annotated within an intron of a host gene, but no homolog of snR107 is annotated. Other cases either resemble one of the above scenarios or are inconclusive due to the lack of a clearly conserved snoRNA (or possibly due to incomplete annotation). Although these examples are potentially interesting, we have chosen not to elaborate on them in the manuscript in order to maintain focus and avoid speculative interpretation in the absence of stronger evidence.
The authors suggest that snoRNAs can be processed from the exons of protein coding genes, but snoRNA processing would destroy the mRNA. Thus snoRNAs processing and mRNA function seem to be alternative outcomes that are mutually exclusive. Can the authors comment?
In theory, we agree with reviewer on the mutually exclusive nature of mRNA and snoRNA expression for putative snoRNA hosted in the exon of protein coding genes. However, we want to clarify that the specific examples of snoRNA precursor (or host) developed in the manuscript (mamRNA-snoU14 in S.pombe and, in this resubmission, SNHG25 in H. sapiens) are non-coding. So although we do not exclude that our model of sequential processing through splicing and endonucleolytic cleavage could apply to coding snoRNA precursors, it is not something we want to insist on, especially given the lack of experimental evidence for these cases.
It is possible that the use of the term "exonic snoRNA" in the first version of the manuscript lead to the reviewer's impression that we explicitly meant that snoRNA processing can be processed from the exon of protein coding genes, which was not what we meant (although we do not exclude it). If that was the case, we apologize for the confusion. We have now clarified the issue (see next point).
Minor comments:
The term "exonic snoRNA" is confusing. Isn't any snoRNA by definition an exon?
We agree that this term can be confusing, a sentiment that was also shared by reviewer 3. We replaced the problematic term by either "non-intronic snoRNA", "snoRNA" or "snoRNA gene located in exon" depending on the context, which are more unambiguous in conveying our intended meaning.
The methods section does not include how similar snoRNA clusters were identified in other species
We have now corrected this omission in the method section ('Identification of mixed snoRNA clusters' subsection): "To identify mixed snoRNA clusters, we downloaded the latest genome annotation from Ensembl and selected snoRNAs co-hosted within the same precursor, with at least one being intronic and at least one being non-intronic. We filtered out ambiguous cases where snoRNAs overlapped exons defined as 'retained introns', reasoning that in these cases the snoRNA is more likely to be intronic than not."
In the discussion the authors argue that a previously published observation that S. pombe U14 does not complement a S. cerevisiae mutation can be explained because "was promoter elements... were simply not included in the transgene sequence". However, even if promoter elements were included, the dsRNA structure of S. pombe would not be cleaved by the S. cerevisiae RNase III. I doubt that missing promoter elements are the full explanation, and the authors provide insufficient data to support this conclusion.
We agree with the reviewer that, given the substantial divergence in substrate specificity between Pac1 and Rnt1, it is unlikely that S. pombe snoU14 would be efficiently processed from its precursor in S. cerevisiae. We did not intend to suggest otherwise, and we have now removed this part of the discussion. As the experiment reported by Samarsky et al. did not detect expression of the S. pombe snoU14 precursor (even its unprocessed form), it remains inconclusive with respect to the conservation (or lack thereof) of snoU14 processing mechanisms.
For the record, we had originally included this discussion to point out that the lack of cryptic promoter activity (or at least none that S. cerevisiae can use) within the S. pombe snoU14 precursor supports the idea that transcription initiates solely upstream of the mamRNA precursor. However, we recognize that this argument is speculative and potentially confusing. We have therefore removed it from the revised manuscript to maintain clarity and focus.
**Referees cross-commenting**
I agree with the other 2 reviewers but think the thiouracil pulse labeling reviewer 2 suggests would take considerable work and if snoRNA processing is very fast might not be as conclusive as the reviewer suggests.
We are grateful to the reviewer for this comment, which helped us perform this reviewing in a timely manner.
Reviewer #1 (Significance (Required)):
In the current manuscript the authors show that in S. pombe snoRNA snR107 and U14 are processed from a common precursor in a way that has not previously been described. The experiments describing this processing are thorough and convincing, and include RNAseq, degradome sequencing, northern blotting, qRT-PCR and the analysis of mutations that disrupt various secondary structures in figures 1, 2, and 3. The authors thereby describe a previously unknown gene design where both the exon and the intron are processed into a snoRNA.
__Reviewer #2 (Evidence, reproducibility and clarity (Required)): __
__ __The manuscript presents a novel mode of processing for polycistronic snoRNAs in the yeast Saccharomyces pombe. The authors demonstrate that the processing sequence of a transcription unit containing U14, intronic snR107, and an overlapping non-coding mamRNA is determined by secondary structures recognized by RNase III (Pac1). Specifically, the formation of a stem structure over the mamRNA exon-exon junction facilitates the processing of terminal exonic-encoded U14. Consequently, U14 maturation occurs only after the mamRNA intron (containing snR107) is spliced out. This mechanism prevents the accumulation of unspliced, truncated mamRNA.
1.The first section describing the processing steps is challenging to follow due to the unusual organization of the locus and maturation pathway. If the manuscript is intended for a broad audience, I recommend simplifying this section and presenting it in a more accessible manner. A larger diagram illustrating the transcription unit and processing intermediates would be beneficial. Additionally, introducing snR107 earlier in the text would improve clarity.
We thank the reviewer for these excellent suggestions. In the previous version of the manuscript, we were cautious in how we introduced the locus, as snR107 and the associated intron had not yet been published. This is no longer the case, as the locus is now described in Leroy et al. (2025). Accordingly, we now introduce the complete locus at the beginning of the manuscript and have improved the corresponding diagram (new Figure 1A). We believe these changes enhance clarity and make the section more accessible to a broader audience.
2.Evaluation of some results is difficult due to the overexposure of Northern blot signals in Figures 1 and 2. The unspliced and spliced precursors appear as a single band, making it hard to distinguish processing intermediates. Would the authors consider presenting these results similarly to Figure 3, where bands are more clearly resolved? Or presenting both overexposed and underexposed blots?
For all blots (probes A, B, and C), we selected an exposure level that allows detection of precursor forms under wild-type (WT) conditions. This necessarily results in some overexposure of the accumulating precursors in mutant conditions, due to their broad dynamic range of accumulation. To address this, we now provide an additional supplementary "source data" file containing all uncropped blots with both low and high exposures.
For example, a lower exposure version of the blot in new Figure 1.B (included in the source data file) confirms the consistent accumulation of the spliced precursor when Pac1 activity is compromised. The unspliced precursor also shows slight accumulation in the Pac1-ts mutant, although to a much lesser extent than the spliced precursor. This observation is consistent with our qPCR results (new Figure 1.C).
Importantly, because this effect is not observed in neither the Pac1-AA or the steam-dead (SD) mutants, we interpret it as an indirect effect—possibly reflecting a mild growth defect in the Pac1-ts strain, even under growth-permissive conditions. We now explicitly address this point in the revised manuscript.
3.Additionally, I noticed a discrepancy in U14 detection: Probe B gives a strong signal for U14 in Figure 3B, whereas in Figures 1 and 2, U14 appears as faint bands. Could the authors clarify this inconsistency?
We thank the reviewer for pointing out this discrepancy. The variation in U14 signal intensity is most likely due to technical differences in UV crosslinking efficiency during the Northern blot procedure. This step can differentially affect the membrane retention of RNA species depending on their length, as previously reported (PMID: 17405769). Because U14 is a relatively abundant snoRNA, the fainter signal observed in Figure 1 (relative to the accumulating precursor) likely reflects suboptimal crosslinking of shorter RNAs in that particular blot.
Importantly, this technical variability does not impact the conclusions of our study, as we do not compare RNA species of different lengths directly. To increase transparency, we now provide a supplementary "source data" file that includes all uncropped blots from our Northern blot experiments. These include examples—such as the uncropped blot for Figure 1B—where U14 retention is more consistent.
4.Furthermore, ethidium bromide (EtBr) staining of rRNA is used as a loading control, but overexposed signals from the gel may not accurately reflect RNA amounts on the membrane. This could affect the interpretation of mature RNA species' relative abundance.
We thank the reviewer for pointing this out and have now measured rRNAs loading on the same northern blot membrane from probes complementary to mature rRNA. We updated new Figures 1B, 2B, 3B, S1B, and S3A accordingly.
5.To further support the sequential processing model, the authors could use pulse-labeling thiouracil to test the accumulation of newly transcribed RNAs and accumulation of individual species. Additionally, it could help determine whether U14 can be processed through alternative, less efficient pathways. Would the authors consider incorporating this approach?
We thank the reviewer for this pertinent suggestion. We actually plan to investigate the putative alternative U14 maturation pathway in future work, and the suggested approach will definitely be instrumental for that. However, to keep the present manuscript focused, and also to keep the review timely (successful pulse-chase experiments are likely to take time to optimize – as also suggested by the other reviewers in their cross-commenting section), we prefer not to perform this experiment for this reviewing.
7.In the final section, the authors propose that this processing mechanism is conserved across species, identifying 12 similar genetic loci in different organisms. This is very interesting finding. In my opinion, providing any experimental evidence would greatly strengthen this claim and the manuscript's significance. Even preliminary validation would add substantial value!
We thank the reviewer for his/her enthusiasm and are glad to provide some preliminary validation to the final section of our manuscript. Specifically, we focused on the SNHG25 gene in H. sapiens, which hosts two snoRNAs—one intronic, as previously shown in Figure 4B, and one non-intronic. We substantiated our predictions through the re-analysis of multiple sequencing datasets in human cell lines, as outlined below:
I.Analysis of CAGE-seq and nano-COP datasets indicates a single major transcription initiation site at the SNHG25 locus. Both the intronic and non-intronic snoRNAs are present within the same nascent precursor transcripts (Supplementary Figure 4D).
II.Degradome-seq experiments in human cell lines reveal that the predicted splicing-dependent stem-loop structure within the SNHG25 gene is subject to endonucleolytic cleavage (Supplementary Figure 4D). The cleavage sites are located at the apical loop and flanking the stem, displaying a staggered symmetry characteristic of RNase III activity (Figure 4C). Importantly, the nucleotide sequence surrounding the 3' cleavage site and the 3' splice-site are conserved in other vertebrates (Supplementary Figure 4.D).
III. fCLIP experiments demonstrate that DROSHA associates with the spliced SNHG25 transcript (Supplementary Figure 4D).
Together, these analyses support the generalizability of our model beyond fission yeast. They confirm the structure of the SNHG25 gene as a single non-coding RNA precursor hosting two snoRNAs, one of which is intronic. Importantly, these findings unambiguously show that the predicted stem-loop structure is subject to endonucleolytic cleavage, and they are consistent with DROSHA, an RNase III enzyme, being responsible for this processing step.
**Referees cross-commenting**
The other two reviewers' comments are justified.
Reviewer #2 (Significance (Required)):
The authors describe an interesting novel mode of snoRNA procseeimg form the host transcript. The results appear sound and intriguing, especially if the proposed mechanism can be confirmed across different organisms. Including such validation would significantly enhance the impact and make this work of broad audience interest.
My expertise: transcription, non-coding RNAs
__Reviewer #3 (Evidence, reproducibility and clarity (Required)): __
The manuscript by Migeot et al., focuses on a new Pac1-mediated snoRNA processing pathway for intron-encoded snoRNA pairs in yeast Schizosaccharomyces pombe. The novelty of the findings described in MS is the report of an unusual and relatively rare genomic organization and sequential processing of a few snoRNA genes in S. pombe and other eukaryotic organisms. It appears that in the case of snoRNA pairs, hosted in pre-mRNA in the intron and exon, respectively, the release of separate pre-snoRNAs from the host gene relies first on splicing to free the intron-encoded snoRNA, followed by endonucleolytic cleavage by RNase III (Pac1 in S. pombe) to produce snoRNA present in the mRNA exon. The sequential processing pathway, ensuring proper maturation of two snoRNAs, was demonstrated and argued in an elegant and clear way. The main message of the MS is straightforward, most experiments are properly conducted and specific conclusions based on the data are justified and valid. The text is clearly written and well-presentded.
But there are some shortcomings.
1.First of all, the title of the MS and general conclusions regarding the Pac1-mediated sequential release of snoRNA pairs hosted within the intron are definitely an overstatement. Especially the title suggests that this genomic organization and unusual processing mode of these snoRNAs is widespread. Later in the discussion the authors themselves admit that such mixed exonic-intronic snoRNAs are rare, although their presence may be underestimated due to annotation problems. It is likely that such snoRNA arrangement and processing is conserved, but the evidence is missing and only unique cases were identified based on bioinformatics mining and their processing has not been assayed. This makes the generalization impossible based on a single documented mamRNA/snoU14 example, no matter how carefully examined.
We thank the reviewer for clearly articulating this concern. In response, we now provide additional evidence supporting conservation of the proposed mechanism in other species:
Conservation within the Schizosaccharomyces genus (Figures S4A–C) has been further analyzed, as suggested by Reviewer 1. This expanded analysis highlights conserved features—such as splice sites and cleavage sites within the predicted stem-loop structure—indicating that these elements are under selective constraint.
Conservation in mammals is now supported by experimental data, as detailed in our responses to point #7 of Reviewer 2 and major comment #1 of Reviewer 1. Specifically, we show that for the SNHG25 gene in H. sapiens (Figure S4D):
(1) nascent transcription give rise to a single non-coding RNA precursor that hosts two snoRNAs, one of which is intronic;
(2) the predicted stem-loop structure contains conserved elements and is subject to endonucleolytic cleavage;
(3) the RNase III enzyme DROSHA associates with the spliced SNHG25 precursor.
Together, these analyses strengthen the evidence for the evolutionary conservation of the mechanism and support the general conclusions and title of the manuscript.
Another interesting observation is that, similarly to other intron-encoded snoRNA in other species, there is a redundant pathway to produce mature U14 in addition to Pac1-mediated cleavage. In the case of intronic snoRNAs in S. cerevisiae, their release could be performed either by splicing/debranching or Rnt1 cleavage, but there is also a third alternative option, that is processing following transcription termination downstream of the snoRNA gene, which at the same time interferes with the expression of the host gene. Is such a scenario possible as an alternative pathway for U14? Are there any putative, or even cryptic, terminators downstream of the U14 gene? The authors did not consider or attempt to inspect this possibility.
We thank the reviewer for this interesting and thoughtful comment. First, we would like to clarify that snoU14 is not intron-encoded; rather, it is located on the exon downstream of the intron-encoded snR107.
Regarding the possibility of transcription termination-based processing: downstream of snoU14, we identified a non-consensus polyadenylation signal (AUUAAA) preceded by a U-rich tract, followed by three consensus polyadenylation signals (AAUAAA) within a 500-nt window. These elements likely contribute to robust and redundant transcription termination at this highly expressed locus. However, since all these sites are located downstream of snoU14, they do not provide an alternative 5′-end processing mechanism for this snoRNA –they reflect normal termination.
If we correctly understood the reviewer’s suggestion (apologies if not), they may have been referring to the possibility of a cryptic or alternative polyadenylation site between snR107 and snoU14 instead. If cleavage were to occur in this inter-snoRNA region while transcription continued past snoU14, it could, in principle, allow for alternative processing of snoU14. We have indeed considered this scenario. However, we currently do not find strong support for it: there are no identifiable polyadenylation signals motifs between the two snoRNAs, aside from a weakly conserved and questionable AAUAAU hexamer that does not appear to be used as polyA site at least in WT conditions (DOI: 10.4161/rna.25758). Given the lack of evidence, we chose not to explore this hypothesis further in the present manuscript, though it remains an interesting possibility for future investigation.
I also have some concerns or comments related to the presented research, which are no major, but are mainly related to data quatification, but have to be addressed.
- In Pac1-ts and Pac1-AA strains the level of mature U14 seems upregulated compared to respective WT (Figure 1A). At the same time mature 25S and 18S rRNAs are less abundant. But there is no quantification and it is not mentioned in the text. What could be the reason for these effects?
We thank the reviewer for this observation. As reviewer 2 also noted, ethidium bromide staining of mature rRNAs is not a reliable quantitative loading control. In response to this concern, we have now reprobed all northern blots with radiolabeled rRNA probes. These provide a more accurate and consistent loading for our blots (new Figures 1B, 2B, 3B, S1B, S3A).
Using these improved loading controls, it is evident that snoU14, snR107, and the unspliced precursor are all slightly upregulated in the Pac1-ts strain, although to a much lesser extent than the spliced precursor, which accumulates dramatically. We do not observe this effect in either the Pac1-AA or stem-dead (SD) mutants. We therefore interpret the modest upregulation as an indirect effect, possibly linked to the physiological state of the Pac1-ts mutant, which exhibits slower growth even at growth-permissive temperatures. We now explicitly discuss this in the revised manuscript.
Regarding the suggestion to include quantification of the northern blot signal: we opted not to include this in the figures for the following reasons. First, the accumulation of the spliced precursor—the central focus of our analysis—is large and highly reproducible across all replicates and conditions. Second, northern blot quantification by pixel intensity remains semi-quantitative, particularly for comparisons across RNAs of highly different abundance. Finally, we support our conclusions with additional quantitative data from RT-qPCR and RNA-seq, which provide more robust measures of RNA accumulation.
- Processing of the other snoRNA from the mamRNA/snoU14 precursor is largely overlooked in the MS. It is commented on only in the context of mutants expressing constitutive mamRNA-CS constructs (Figure 3B). Its level was checked in Pac1-ts and Pac1-AA (Supplementary Figure 1), but the authors conclude that "its expression remained largely unaffected by Pac1 inactivation", which is clearly not true. Similarly to U14, also snR170 is increased in Pac1-ts and Pac1-AA strains, at least judged "by eye" because the loading control or quantification is not provided. This matter should be clarified.
We thank the reviewer for pointing this out. We have now included appropriate loading controls for Supplementary Figure 1 to clarify the interpretation. As discussed in our response to the previous comment, we observe a general upregulation of the mamRNA locus in the Pac1-ts strain, which likely contributes to the increased levels of both snR107 and snoU14. However, because this upregulation is not observed in the Pac1-AA or stem-dead (SD) mutants, we interpret it as an indirect effect, possibly related to the altered physiological state of the Pac1-ts strain (e.g., slightly reduced growth rate even at the permissive temperature). This interpretation has now been clearly explained in the revised manuscript.
We also identified and corrected a labeling error in the previous version of Supplementary Figure 1, where the Pac1-ts and Pac1-AA strains were inadvertently swapped. We sincerely apologize for the confusion this may have caused and have now ensured that all figure panels are correctly labeled and consistent with the text.
Other minor comments:
Minor points:
- Page 1, Abstract. The sentence "The hairpin recruits the RNase III Pac1 that cleaves and destabilizes the precursor transcript while participating in the maturation of the downstream exonic snoRNA, but only after splicing and release of the intronic snoRNA" is not entirely clear and should be simplified, maybe split into two sentences. This message is clear after reading the MS and learning the data, but not in the abstract.
We thank the reviewer for pointing this out and have now clarified the abstract following the suggestion to split and simplify the problematic sentence : "... the sequence surrounding an exon-exon junction within their precursor transcript folds into a hairpin after splicing of the intron. This hairpin recruits the RNase III ortholog Pac1, which participates in the maturation of the downstream snoRNA by cleaving the precursor."
Page 1, Introduction. I am not convinced by the need to use the term "exonic snoRNA" for all snoRNA that are not intronic, which is misleading, and is rather associated per se with snoRNA encoded in the mRNA exon. It has been used before in the review about snoRNAs by Michelle Scott published in RNA Biol (2024), but it does not justify its common use.
We thank the reviewer for raising this important point. We agree that the term “exonic snoRNA” can be misleading, as it was previously used to specifically refer to snoRNAs embedded within exons of mRNA transcripts—an rare and potentially artifactual scenario, as very cautiously discussed by Michelle Scott and colleagues in their review published in RNA Biol (2024).
In the previous version of our manuscript, we actually used “exonic snoRNA” in a broader sense to denote any snoRNA not encoded within an intron, primarily for convenience in contrasting the processing of intronic snR107 with that of non-intronic/exonic snoU14. However, we recognize that this usage is non-standard and risks confusion due to the ambiguity surrounding the term’s definition in the literature.
In light of this, and in agreement with reviewer 1 who raised a similar concern, we have revised the manuscript to remove the term “exonic snoRNA” entirely. Depending on the context, we now refer more precisely to “non-intronic snoRNA,” “snoRNA gene located in exon,” or simply “snoRNA.”
Supplementary Figure 3. It is difficult to assess whether the level of mature rRNAs is unchanged in the mutants based on EtBr staining and without calculations. Northern blotting should be performed and the levels properly calculated.
As suggested, we performed northern blotting on mature 18S and 25S, quantified the signal and observed no significant differences (new Supplementary Figure 3).
**Referees cross-commenting**
I also agree that 4sU labeling may require too much work with a questionable result.
We are grateful to the reviewer for this comment, which helped us perform this reviewing in a timely manner.
Reviewer #3 (Significance (Required)):
Strengths: 1. Novelty of the described genomic arrangement of snoRNA/ncRNA genes and their processing in a sequential and regulated manner.
Potential conservation of this pathways across eukaryotic organisms. Well designed and performed experiments followed by proper conclusions.
Limitations: 1. Insufficient evidence to support generalization of the study results.
Moderate overall impact of the study
Advance: This research can be placed within publications describing specific processing pathways for various non-coding RNAs, including for example unusual chimeric species such as sno-lncRNAs. In this context, the presented results do advance the knowledge in the field by providing mechanistic evidence for a tightly controlled and coordinated maturation of selected ncRNAs.
Audience: Basic research and specialized. The interest in this research will rather be limited to a specific field.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
The manuscript by Migeot et al., focuses on a new Pac1-mediated snoRNA processing pathway for intron-encoded snoRNA pairs in yeast Schizosaccharomyces pombe.
The novelty of the findings described in MS is the report of an unusual and relatively rare genomic organization and sequential processing of a few snoRNA genes in S. pombe and other eukaryotic organisms. It appears that in the case of snoRNA pairs, hosted in pre-mRNA in the intron and exon, respectively, the release of separate pre-snoRNAs from the host gene relies first on splicing to free the intron-encoded snoRNA, followed by endonucleolytic cleavage by RNase III (Pac1 in S. pombe) to produce snoRNA present in the mRNA exon. The sequential processing pathway, ensuring proper maturation of two snoRNAs, was demonstrated and argued in an elegant and clear way. The main message of the MS is straightforward, most experiments are properly conducted and specific conclusions based on the data are justified and valid. The text is clearly written and well-presentded.
But there are some shortcomings. First of all, the title of the MS and general conclusions regarding the Pac1-mediated sequential release of snoRNA pairs hosted within the intron are definitely an overstatement. Especially the title suggests that this genomic organization and unusual processing mode of these snoRNAs is widespread. Later in the discussion the authors themselves admit that such mixed exonic-intronic snoRNAs are rare, although their presence may be underestimated due to annotation problems. It is likely that such snoRNA arrangement and processing is conserved, but the evidence is missing and only unique cases were identified based on bioinformatics mining and their processing has not been assayed. This makes the generalization impossible based on a single documented mamRNA/snoU14 example, no matter how carefully examined. Another interesting observation is that, similarly to other intron-encoded snoRNA in other species, there is a redundant pathway to produce mature U14 in addition to Pac1-mediated cleavage. In the case of intronic snoRNAs in S. cerevisiae, their release could be performed either by splicing/debranching or Rnt1 cleavage, but there is also a third alternative option, that is processing following transcription termination downstream of the snoRNA gene, which at the same time interferes with the expression of the host gene. Is such a scenario possible as an alternative pathway for U14? Are there any putative, or even cryptic, terminators downstream of the U14 gene? The authors did not consider or attempt to inspect this possibility.
I also have some concerns or comments related to the presented research, which are no major, but are mainly related to data quatification, but have to be addressed. In Pac1-ts and Pac1-AA strains the level of mature U14 seems upregulated compared to respective WT (Figure 1A). At the same time mature 25S and 18S rRNAs are less abundant. But there is no quantification and it is not mentioned in the text. What could be the reason for these effects? Processing of the other snoRNA from the mamRNA/snoU14 precursor is largely overlooked in the MS. It is commented on only in the context of mutants expressing constitutive mamRNA-CS constructs (Figure 3B). Its level was checked in Pac1-ts and Pac1-AA (Supplementary Figure 1), but the authors conclude that "its expression remained largely unaffected by Pac1 inactivation", which is clearly not true. Similarly to U14, also snR170 is increased in Pac1-ts and Pac1-AA strains, at least judged "by eye" because the loading control or quantification is not provided. This matter should be clarified.
Other minor comments:
Minor points:
Referees cross-commenting
I also agree that 4sU labeling may require too much work with a questionable result.
Strengths:
Limitations:
Advance:
This research can be placed within publications describing specific processing pathways for various non-coding RNAs, including for example unusual chimeric species such as sno-lncRNAs. In this context, the presented results do advance the knowledge in the field by providing mechanistic evidence for a tightly controlled and coordinated maturation of selected ncRNAs.
Audience:
Basic research and specialized. The interest in this research will rather be limited to a specific field.
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
1. Response to reviewers
We would like to thank the reviewers for carefully reading our manuscript and for their valuable comments in support for the publication of our investigation of rapid promoter evolution of accessory gland genes between Drosophila species and hybrids. We are glad to read that the reviewers find our work interesting and that it provides valuable insights into the regulation and divergence of genes through their promoters. We are encouraged by their acknowledgement of the overall quality of the work and the importance of our analyses in advancing the understanding of cis-regulatory changes in species divergence.
2. Point-by-point description of the revisions
Reviewer #
Reviewer Comment
Author Response/Revision
Reviewer 1
The authors test the hypothesis that promoters of genes involved in insect accessory glands evolved more rapidly than other genes in the genome. They test this using a number of computational and experimental approaches, looking at different species within the Drosophila melanogaster complex. The authors find an increased amount of sequence divergence in promoters of accessory gland proteins. They show that the expression levels of these proteins are more variable among species than randomly selected proteins. Finally, they show that within interspecific hybrids, each copy of the gene maintains its species-specific expression level.
We thank Reviewer 1 for their detailed review and positive feedback on our manuscript, and for their helpful suggestions. We have now fully addressed the points raised by Reviewer 1 and have provided the suggested clarifications and revisions to improve the flow, readability, and presentation of the data, which we believe have improved the manuscript significantly.
The work is done with expected standards of controls and analyses. The claims are supported by the analysis. My main criticism of the manuscript has to do not with the experiments or conclusion themselves but with the presentation. The manuscript is just not very well written, and following the logic of the arguments and results is challenging.
The problem begins with the Abstract, which is representative of the general problems with the manuscript. The Abstract begins with general statements about the evolution of seminal fluid proteins, but then jumps to accessory glands and hybrids, without clarifying what taxon is being studied, and what hybrids they are talking about. Then, the acronym Acp is introduced without explanation. The last two sentences of the Abstract are very cumbersome and one has to reread them to understand how they link to the beginning of the Abstract.
More generally, if this reviewer is to be seen as an "average reader" of the paper, I really struggled through reading it, and did not understand many of the arguments or rationale until the second read-through, after I had already read the bottom line. The paragraph spanning lines 71-83 is another case in point. It is composed of a series of very strongly worded sentences, almost all starting with a modifier (unexpectedly, interestingly, moreover), and supported by citations, but the logical flow doesn't work. Again, reading the paragraph after I knew where the paper was going was clearer, but on a first read, it was just a list of disjointed statements.
Since most of the citations are from the authors' own work, I suspect they are assuming too much prior understanding on the part of the reader. I am sure that if the authors read through the manuscript again, trying to look through the eyes of an external reader, they will easily be able to improve the flow and readability of the text.
We thank the reviewer for their detailed feedback and are glad that they acknowledge our work fully supports the claims of our manuscript. We also appreciate their helpful suggestions for improving the readability of the manuscript and have done our best to re-write the abstract and main text where indicated. In particular, the paragraph between lines 71-83 have been rewritten and we have taken care to write to non-expert readers.
1) In the analysis of expression level differences, it is not clear what specific stage / tissue the levels taken from the literature refer to. Could it be that the source of the data is from a stage or tissue where seminar fluid proteins will be expressed with higher variability in general (not just inter-specifically) and this could be skewing the results? Please add more information on the original source of the data and provide support for their validity for this type of comparison.
These were taken from publicly available adult male Drosophila datasets, listed in the data availability statement and throughout the manuscript. We have provided more detail on the tissue used for analysis of Acp gene expression levels.
2) The sentence spanning lines 155-157 needs more context.
We have added more context to lines 155-157.
3) Line 203-204: What are multi-choice enhancers?
We replaced the sentence with "... such as rapidly evolving enhancers or nested epistasis enhancer networks"
4) Figure 1: The terminology the authors use, comparing the gene of interest to "Genome" is very confusing. They are not comparing to the entire genome but to all genes in the genome, which is not the same.
We have changed the word "genome" to "all genes in the genome" on the reviewer's suggestion.
5) Figure 2: Changes between X vs. Y is redundant (either changes between X and Y or changes in X vs. Y).
We assume that the reviewer is referring to Fig. 2B, which does not measure changes between X and Y, but changes in distribution between Acps and the control group. We have explained this in the figure legend.
The manuscript addresses a general question in evolutionary biology - do control regions diverge more quickly protein coding regions. The answer is that yes, they do, but this is actually not very surprising. The work is probably thus of more interest to people interested in the copulatory proteins or in the evolution of mating systems, than to people interested in broader evolutionary questions.
We appreciate this reviewer's recognition of the significance of our work and would like to point out that there are very few studies looking at promoter evolution as detailed in the introduction. Of particular relevance, our study using Acp genes allows us to directly test the impact of promoter mutations on the expression by comparing two alleles in male accessory glands of Drosophila hybrids. Male accessory glands consist of only two secretory cell types allowing us to study evolution of gene expression in a single cell type (Acps are either expressed in main cells or secondary cells). Amid this unique experimental set up we can conclude that promoter mutations can act dominant, in contrast to mutations in protein coding regions, which are generally recessive. Thus, our study is unique in pointing out a largely overseen aspect of gene evolution.
Reviewer 2
This manuscript explores promoter evolution of genes encoding seminal fluid proteins expressed in the male accessory gland of Drosophila and finds cis-regulatory changes underlie expression differences between species. Although these genes evolve rapidly it appears that the coding regions rarely show signs of positive selection inferring that changes in their expression and hence promoter sequences can underlie the evolution of their roles within and among species.
We thank Reviewer 2 for their thorough review, positive feedback on the importance of our work, and suggestions for improving the manuscript. We have addressed all points raised by the reviewer, including analysis of Acp coding region evolution, additional analyses of hybrid expression data, and improved the clarity of the text.
Figure 1 illustrates evidence that the promoter regions of these gene have accumulated more changes than other sampled genes from the Drosophila genome. While this convinces that the region upstream of the transcription start site has diverged considerably in sequence (grey line compared to black line), Figure 1A also suggests the "Genespan" region which includes the 5'UTR but presumably also part of the coding region is also highly diverged. It would be useful to see how the pattern extends into the coding region further to compare further to the promoter region (although Fig 1H does illustrate this more convincingly).
The reviewer raises an interesting point, and certainly all parts of genes evolve. Fig. 1A shows the evolutionary rates of Acps compared to the genome average from phyloP27way scores calculated from 27 insect species. Since these species are quite distant it is unsurprising that they show divergence in coding regions as well as promoter regions. In fact, we addressed whether promoter regions evolve fast in closely related Drosophila species in Fig. 1H compared to coding regions. We have included an additional analysis of coding region evolution in Figure 1B.
Figure 2 presents evidence for significant changes in (presumably levels of) expression of male accessory gland protein (AcP) genes and ribosomal proteins genes between pairs of species, which is reflected in the skew of expression compared to randomly selected genes.
Correct, we have rephrased the statement for clarity.
Figure 3 shows detailed analysis for 3 selected AcP genes with significantly diverged expression. The authors claim this shows 'substitution' hotspots in the promoter regions of all 3 genes but this could be better illustrated by extending the plots in B-D further upstream and downstream to compare to these regions.
We picked the 300-nucleotide promoter region for this analysis as it accumulated significant changes as shown in Fig. 1E-H, and extending the G plots (Fig. 3B-D) to regions with lower numbers of sequence changes would not substantially change the conclusion. Specifically, this analysis identifies sequence change hotspots within fast-evolving promoter regions, rather than comparing promoter regions to other genomic regions, as we previously addressed. The plot is based on a cumulative distribution function and the significant positive slope in the upstream region where promoters are located identifies a hotspot for accumulation of substitutions. There could be other hotspots, but the point being made is that significant hotspots consistently appear in the promoter region of these three genes.
Figure 4 shows the results of expression analysis in parental lines of each pair of species and F1 hybrids. However the results are very difficult to follow in the figure and in the relevant text. While the schemes in A, C. E and G are helpful, the gel images are not the best quality and interpretations confusing. An additional scheme is needed to illustrate hypothetical outcomes of trans change, cis change and transvection to help interpret the gels. On line 169 (presumably referring to panels D and F although C and D are cited on the next line) the authors claim that Obp56f and CG11598 'were more expressed in D. melanogaster compared to D. simulans' but in the gel image the D. sim band is stronger for both genes (like D. sechellia) compared to the D. mel band. The authors also claim that the patterns of expression seen in the F1s are dominant for one allele and that this must be because of transvection. I agree this experiment is evidence for cis-regulatory change. However the interpretation that it is caused by transvection needs more explanation/justification and how do the authors rule out that it is not a cis X trans interaction between the species promoter differences and differences in the transcription factors of each species in the F1? Also my understanding is that transvection is relatively rare and yet the authors claim this is the explanation for 2/4 genes tested.
We appreciate the reviewer's comments on Figure 4 and the opportunity to improve its clarity. To address these concerns, we have carefully checked the figure citations and corrected any inconsistencies.
The reviewer raises an important point about our interpretation of transvection. We have expanded our discussion of this result to consider why transvection is a plausible explanation for the observed dominance patterns and also consider cis x trans interactions between species-specific promoters and transcription factor binding. While rare, transvection likely has more relevance in hybrid regulatory contexts involving homologous chromosome pairing which we discuss this in the revised text.
Line 112 states that the melanogaster subgroup contains 5 species - this is incorrect - while this study looked at 5 species there are more species in this subgroup such as mauritiana and santomea.
We have corrected the statement about the number of species in the melanogaster subgroup.
Lines 131-134 could explain better what the conservation scores and their groupings mean and the rationale for this approach.
We have clarified what the conservation scores and their groupings mean and the rationale for this approach.
Line 162 - the meaning of the sentence starting on this line is unclear - it sounds very circular.
We have rephrased the statement for more clarity.
Line 168 should cite Fig 4 H instead of F.
We have amended citation of Fig 4F to H.
Reviewer 3
In this study, McQuarrie et al. investigate the evolution of promoters of genes encoding accessory gland proteins (Acps) in species within the D. melanogaster subgroup. Using computational analyses and available genomic and transcriptomic datasets, they demonstrate that promoter regions of Acp genes are highly diverse compared to the promoters of other genes in the genome. They further show that this diversification correlates with changes in gene expression levels between closely related species. Complementing these computational analyses, the authors conduct experiments to test whether differences in expression levels of four Acp genes with highly diverged promoter regions are maintained in hybrids of closely related species. They find that while two Acp genes maintain their expression level differences in hybrids, the other two exhibit dominance of one allele. The authors attribute these findings to transvection. Based on their data, they conclude that rapid evolution of Acp gene promoters, rather than changes in trans, drives changes in Acp gene expression that contribute to speciation.
We thank Reviewer 3 for their thorough review and suggestions. We further thank the reviewer for acknowledging the importance of our findings and for pointing out that it contributes to our understanding of speciation. We have thoroughly addressed all comments from the reviewer and significantly revised the manuscript. We believe that this has greatly improved the manuscript.
Unfortunately, the presented data are not sufficient to fully support the conclusions. While many of the concerns can be addressed by revising the text to moderate the claims and acknowledge the methodological limitations, some key experiments require repetition with more controls, biological replicates, and statistical analyses to validate the findings.
Specifically, some of the main conclusions heavily rely on the RT-PCR experiments presented in Figure 4, which analyze the expression of four Acp genes in hybrid flies. The authors use PCR and RFLP to distinguish species-specific alleles but draw quantitative conclusions from what is essentially a qualitative experiment. There are several issues with this approach. First, the experiment includes only two biological replicates per sample, which is inadequate for robust statistical analysis. Second, the authors did not measure the intensity of the gel fragments, making it impossible to quantify allele-specific expression accurately. Third, no control genes were used as standards to ensure the comparability of samples.
The gold standard for quantifying allele-specific expression is using real-time PCR methods such as TaqMan assays, which allow precise SNP genotyping. To address this major limitation, the authors should ideally repeat the experiments using allele-specific real-time PCR assays. This would provide a reliable and quantitative measurement of allele-specific expression.
If the authors cannot implement real-time PCR, an alternative (though less rigorous) approach would be to continue using their current method with the following adjustments:
• Include a housekeeping gene in the analysis as an internal control (this would require identifying a region distinguishable by RFLP in the control).
• Quantify the intensity of the PCR products on the gel relative to the internal standard, ensuring proper normalization.
• Increase the sample size to allow for robust statistical analysis.
These experiments could be conducted relatively quickly and would significantly enhance the validity of the study's conclusions.
We thank the reviewer for their detailed suggestions for improving the conclusions in Fig. 4. Indeed, incorporating a housekeeping gene as a control supports our results for qualitative analysis of gene expression in hybrids assessing each allele individually (Fig 4), and improves interpretation for non-experts. We have also included an additional analysis in the new Fig. 5 which analyses RNA-seq expression changes in D. melanogaster x D. simulans hybrid male accessory glands. We believe these additions have significantly improved the manuscript and its conclusions.
While the following comments are not necessarily minor, they can be addressed through revisions to the text without requiring additional experimental work. Some comments are more conceptual in nature, while others concern the interpretation and presentation of the experimental results. They are provided in no particular order.
- A key limitation of this study is the use of RNA-seq datasets from whole adult flies for interspecies gene expression comparisons. Whole-body RNA-seq inherently averages gene expression across all tissues, potentially masking tissue-specific expression differences. While Acp genes are likely restricted to accessory glands, the non-Acp genes and the random gene sets used in the analysis may have broader expression profiles. As a result, their expression might be conserved in certain tissues while diverging in others- an aspect that whole-body RNA-seq cannot capture. The authors should acknowledge that tissue-specific RNA-seq analyses could provide a more precise understanding of expression divergence and potentially reveal reduced conservation when considering specific tissues independently.
We have added a section discussing the limitations in gene expression analysis in the discussion. In addition, we have included an additional Figure analysing gene expression in hybrid male accessory glands (Fig. 5).
- The statement in line 128, "Consistent with this model," does not accurately reflect the findings presented in Figures 2A and B. Specifically, the data in Figure 2A show that Acp gene expression divergence is significantly different from the divergence of non-Acp genes or a random sample only in the comparison between D. melanogaster and D. simulans. However, when these species are compared to D. yakuba, Acp gene expression divergence aligns with the divergence patterns of non-Acp genes or random samples. In contrast, Figure 2B shows that the distribution of expression changes is skewed for Acp genes compared to random control samples when D. melanogaster or D. simulans are compared to D. yakuba. However, this skew is absent when the two D. melanogaster and D. simulans are compared. Therefore, the statement in line 128 should be revised to accurately reflect these nuanced results and the trends shown in Figure 2A and B.
We have updated the statement for clarity. Here, the percentage of Acps showing significant gene expression changes is greater between more closely related species, but the distribution of expression changes increases between more distantly related species.
- The statement in lines 136-138, "Acps were enriched for significant expression changes in the faster evolving group across all species," while accurate, overlooks a key observation. This trend was also observed in other groups, including those with slower evolving promoters, in some of the species' comparisons. Therefore, the enrichment is not unique to Acps with rapidly evolving promoters, and this should be explicitly acknowledged in the text.
This is a valid point, and we have updated this statement as suggested.
- It would be helpful for the authors to explain the meaning of the d score at the beginning of the paragraph starting in line 131, to ensure clarity for readers unfamiliar with this metric.
This scoring method is described in the methods sections, and we have now included reference to thorough explanation of how d was calculated at the indicated section.
- In Figure 2C-E - the title of the Y-axis does not match the text. If it represents the percentage of genes with significant expression changes, as in Figure 2A, the discrepancies between the percentages in this figure and those in Figure 2A need to be addressed.
We have updated the method used to categorise significant changes in gene expression in the text and the figure legend for clarity.
- The experiment in Figure 3 needs a better explanation in the text. What is the analysis presented in Figure 3B-D. How many species were compared?
We have added additional details in the results section and an explanation of how sequence change hotspots were calculated in the results section is available.
- The concept of transvection should be omitted from this manuscript. First, the definition provided by the authors is inaccurate. Second, even if additional experiments were to convincingly show that one allele in hybrid animals is dominant over the other, there are alternative explanations for this phenomenon that do not involve transvection. The authors may propose transvection as a potential model in the discussion, but they should do so cautiously and explicitly acknowledge the possibility of other mechanisms.
We have updated the text to more conservatively discuss transvection, moving this to the discussion section with additional possibilities discussed.
- The statement at the end of the introduction is overly strong and would benefit from more cautious phrasing. For instance, it could be reworded as: "These findings suggest that promoter changes, rather than genomic background, play a significant role in driving expression changes, indicating that promoter evolution may contribute to the rise of new species."
We have reworded this line following the reviewer's suggestion.
- Line 32 of the abstract: The term "Acp" is introduced without explaining what it stands for. Please define it as "Accessory gland proteins (Acp)" when it first appears.
We have updated the manuscript to define Acp where it is first mentioned.
- Line 61: The phrase "...through relaxed,..." is unclear. Specify what is relaxed (e.g., "relaxed selective pressures").
We have included description of relaxed selective pressures.
- The sentence in lines 74-76, starting in "Interestingly,...." Needs revision for clarity.
We have removed the word interestingly.
- Line 112: Revise "we focused on the melanogaster subgroup which is made up of five species" to: "we focused on the melanogaster subgroup, which includes five species."
We have made this change in the text.
- In line 144 use the phrase "promoter conservation" instead of "promoter evolution"
We have updated the phrasing.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary:
In this study, McQuarrie et al. investigate the evolution of promoters of genes encoding accessory gland proteins (Acps) in species within the D. melanogaster subgroup. Using computational analyses and available genomic and transcriptomic datasets, they demonstrate that promoter regions of Acp genes are highly diverse compared to the promoters of other genes in the genome. They further show that this diversification correlates with changes in gene expression levels between closely related species. Complementing these computational analyses, the authors conduct experiments to test whether differences in expression levels of four Acp genes with highly diverged promoter regions are maintained in hybrids of closely related species. They find that while two Acp genes maintain their expression level differences in hybrids, the other two exhibit dominance of one allele. The authors attribute these findings to transvection. Based on their data, they conclude that rapid evolution of Acp gene promoters, rather than changes in trans, drives changes in Acp gene expression that contribute to speciation.
Major comments:
Unfortunately, the presented data are not sufficient to fully support the conclusions. While many of the concerns can be addressed by revising the text to moderate the claims and acknowledge the methodological limitations, some key experiments require repetition with more controls, biological replicates, and statistical analyses to validate the findings.
Specifically, some of the main conclusions heavily rely on the RT-PCR experiments presented in Figure 4, which analyze the expression of four Acp genes in hybrid flies. The authors use PCR and RFLP to distinguish species-specific alleles but draw quantitative conclusions from what is essentially a qualitative experiment. There are several issues with this approach. First, the experiment includes only two biological replicates per sample, which is inadequate for robust statistical analysis. Second, the authors did not measure the intensity of the gel fragments, making it impossible to quantify allele-specific expression accurately. Third, no control genes were used as standards to ensure the comparability of samples.
The gold standard for quantifying allele-specific expression is using real-time PCR methods such as TaqMan assays, which allow precise SNP genotyping. To address this major limitation, the authors should ideally repeat the experiments using allele-specific real-time PCR assays. This would provide a reliable and quantitative measurement of allele-specific expression.
If the authors cannot implement real-time PCR, an alternative (though less rigorous) approach would be to continue using their current method with the following adjustments:
Minor comments
While the following comments are not necessarily minor, they can be addressed through revisions to the text without requiring additional experimental work. Some comments are more conceptual in nature, while others concern the interpretation and presentation of the experimental results. They are provided in no particular order. 1. A key limitation of this study is the use of RNA-seq datasets from whole adult flies for interspecies gene expression comparisons. Whole-body RNA-seq inherently averages gene expression across all tissues, potentially masking tissue-specific expression differences. While Acp genes are likely restricted to accessory glands, the non-Acp genes and the random gene sets used in the analysis may have broader expression profiles. As a result, their expression might be conserved in certain tissues while diverging in others- an aspect that whole-body RNA-seq cannot capture. The authors should acknowledge that tissue-specific RNA-seq analyses could provide a more precise understanding of expression divergence and potentially reveal reduced conservation when considering specific tissues independently. 2. The statement in line 128, "Consistent with this model," does not accurately reflect the findings presented in Figures 2A and B. Specifically, the data in Figure 2A show that Acp gene expression divergence is significantly different from the divergence of non-Acp genes or a random sample only in the comparison between D. melanogaster and D. simulans. However, when these species are compared to D. yakuba, Acp gene expression divergence aligns with the divergence patterns of non-Acp genes or random samples. In contrast, Figure 2B shows that the distribution of expression changes is skewed for Acp genes compared to random control samples when D. melanogaster or D. simulans are compared to D. yakuba. However, this skew is absent when the two D. melanogaster and D. simulans are compared. Therefore, the statement in line 128 should be revised to accurately reflect these nuanced results and the trends shown in Figure 2A and B. 3. The statement in lines 136-138, "Acps were enriched for significant expression changes in the faster evolving group across all species," while accurate, overlooks a key observation. This trend was also observed in other groups, including those with slower evolving promoters, in some of the species' comparisons. Therefore, the enrichment is not unique to Acps with rapidly evolving promoters, and this should be explicitly acknowledged in the text. 4. It would be helpful for the authors to explain the meaning of the d score at the beginning of the paragraph starting in line 131, to ensure clarity for readers unfamiliar with this metric. 5. In Figure 2C-E - the title of the Y-axis does not match the text. If it represents the percentage of genes with significant expression changes, as in Figure 2A, the discrepancies between the percentages in this figure and those in Figure 2A need to be addressed. 6. The experiment in Figure 3 needs a better explanation in the text. What is the analysis presented in Figure 3B-D. How many species were compared? 7. The concept of transvection should be omitted from this manuscript. First, the definition provided by the authors is inaccurate. Second, even if additional experiments were to convincingly show that one allele in hybrid animals is dominant over the other, there are alternative explanations for this phenomenon that do not involve transvection. The authors may propose transvection as a potential model in the discussion, but they should do so cautiously and explicitly acknowledge the possibility of other mechanisms. 8. The statement at the end of the introduction is overly strong and would benefit from more cautious phrasing. For instance, it could be reworded as: "These findings suggest that promoter changes, rather than genomic background, play a significant role in driving expression changes, indicating that promoter evolution may contribute to the rise of new species."
Text edits:
Throughout the manuscripts there are incomplete sentences and sentences that are not clear. Below is a list of corrections:
This study addresses an important question in evolutionary biology: how seminal fluid proteins achieve rapid evolution despite showing limited adaptive changes in their coding regions. By focusing on accessory gland proteins (Acps) and examining their promoter regions, the authors suggest promoter-driven evolution as a potential mechanism for rapid seminal fluid protein diversification. While this hypothesis is intriguing and can contribute to our understanding of speciation, more rigorous analysis and experimental validation would be needed to support the conclusions. The revised manuscript can be of interest to fly geneticists and to scientists in the fields of gene regulation and evolution.
Keywords for my expertise: Enhancers, transcriptional regulation, development, evolution, Drosophila.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary
This manuscript explores promoter evolution of genes encoding seminal fluid proteins expressed in the male accessory gland of Drosophila and finds cis-regulatory changes underlie expression differences between species. Although these genes evolve rapidly it appears that the coding regions rarely show signs of positive selection inferring that changes in their expression and hence promoter sequences can underlie the evolution of their roles within and among species.
Major comments
Figure 1 illustrates evidence that the promoter regions of these gene have accumulated more changes than other sampled genes from the Drosophila genome. While this convinces that the region upstream of the transcription start site has diverged considerably in sequence (grey line compared to black line), Figure 1A also suggests the "Genespan" region which includes the 5'UTR but presumably also part of the coding region is also highly diverged. It would be useful to see how the pattern extends into the coding region further to compare further to the promoter region (although Fig 1H does illustrate this more convincingly).
Figure 2 presents evidence for significant changes in (presumably levels of) expression of male accessory gland protein (AcP) genes and ribosomal proteins genes between pairs of species, which is reflected in the skew of expression compared to randomly selected genes.
Figure 3 shows detailed analysis for 3 selected AcP genes with significantly diverged expression. The authors claim this shows 'substitution' hotspots in the promoter regions of all 3 genes but this could be better illustrated by extending the plots in B-D further upstream and downstream to compare to these regions.
Figure 4 shows the results of expression analysis in parental lines of each pair of species and F1 hybrids. However the results are very difficult to follow in the figure and in the relevant text. While the schemes in A, C. E and G are helpful, the gel images are not the best quality and interpretations confusing. An additional scheme is needed to illustrate hypothetical outcomes of trans change, cis change and transvection to help interpret the gels. On line 169 (presumably referring to panels D and F although C and D are cited on the next line) the authors claim that Obp56f and CG11598 'were more expressed in D. melanogaster compared to D. simulans' but in the gel image the D. sim band is stronger for both genes (like D. sechellia) compared to the D. mel band. The authors also claim that the patterns of expression seen in the F1s are dominant for one allele and that this must be because of transvection. I agree this experiment is evidence for cis-regulatory change. However the interpretation that it is caused by transvection needs more explanation/justification and how do the authors rule out that it is not a cis X trans interaction between the species promoter differences and differences in the transcription factors of each species in the F1? Also my understanding is that transvection is relatively rare and yet the authors claim this is the explanation for 2/4 genes tested.
Minor comments
Line 112 states that the melanogaster subgroup contains 5 species - this is incorrect - while this study looked at 5 species there are more species in this subgroup such as mauritiana and santomea.
Lines 131-134 could explain better what the conservation scores and their groupings mean and the rationale for this approach.
Line 162 - the meaning of the sentence starting on this line is unclear - it sounds very circular.
Line 168 should cite Fig 4 H instead of F.
This paper is generally well written although some sections would benefit from more explanation. The paper demonstrates cis-regulatory changes between the promoters of orthologs of male accessory gland genes underlie expression differences but that the species differences are not always reflected in hybrids, which the authors interpret as being caused by transvection although there could be other explanations. Overall this provides new insights into the regulation and divergence of these interesting genes. The paper does not explore the consequences of these changes in gene expression although this is discussed to some extent in the Discussion section.
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
Summary:
In this manuscript, authors demonstrated the role of ECM-dependent MEK/ERK/MITF signaling pathway that influences the plasticity of MCs (melanocytes) through their interactions with the environment. The findings emphasize the essential role of the extracellular matrix (ECM) in controlling MC function and differentiation, highlighting a critical need for further research to understand the complex interactions between mechanical factors and ECM components in the cellular microenvironment. Overall, the manuscript is concise, written well and shed light on a complex relationship between ECM protein types and substrate stiffness that affects MC mechanosensation. However, understanding detailed molecular mechanisms involved, especially the roles of MITF and other key regulators, is crucial for comprehending MC function and related pathologies. Authors need to clarify some minor queries to be considered for publication.
We thank this reviewer for the time and caution taken to assess our work. To provide a better understanding of the molecular mechanisms involved in MITF modulation and MC function in response to ECM proteins, we substantially revised the manuscript and now included e.g. bulk RNA sequencing, pharmacological inhibition of FAK and ERK (in addition to MEK inhibition), and MITF depletion.
Major comments to the Authors:
- Authors have chosen ERK signaling pathways to test and draw their conclusion based on existing knowledge in the field, as several studies previously reported the role of ECM to modulate the ERK signaling pathway but it would be interesting to test other signaling pathways unbiasedly; e.g. ECM can also regulate Wnt signaling (PMID: 29454361) and connection of MITF and its target gene TYR expression is also regulated by Wnt in context of melanocyte. (PMID: 29454361, PMID: 34878101, PMID: 38020918).*
The new transcriptome analysis (line 258 ff., revised fig. 5, new fig. 6, new suppl. fig. S5) indeed showed that some components of the Wnt signaling pathway are differentially expressed in response to ECM proteins (new fig. 6B). In comparison, however, the expression of genes involved in MAPK/ERK signaling was more prominently affected by the specific ECM types (new fig. 6C, D), congruent with the biochemical results we presented in the original manuscript. We therefore focused our mechanistic analyses on this pathway, and we consolidated our initial findings with additional pharmacological inhibition experiments. Specifically, like MEK inhibition, ERK inhibition (new fig. 6J-L) increased both MITF nuclear localization and melanin production in MCs exposed to FN, reinforcing the relevance of this pathway in control of MC functions in the model used.
We agree that an additional contribution of Wnt signaling to ECM-dependent regulation of MC phenotypes is possible, including Mitf and Tyr expression. Next to the new Wnt-related transcriptome data (line 323 ff., new fig. 6B), we therefore now included a short discussion on that aspect (line 478 ff.). However, we feel that a comprehensive comparison of the individual contributions of Wnt vs. ERK signaling is beyond the scope of the current manuscript.
- Discussion line 340-344. Please provide the data as it is directly connected to the study, and it would be crucial to interpret data better. As FAK is upregulated and FAK inhibitor did not reduce pERK, is there any possibility that other kinases might involve. Please discuss. Again, authors should check Wnt activation as FAK can activate Wnt signaling in response to matrix stiffness as well. (PMID 29454361).*
We agree with the reviewer that the FAK data required further investigation. In the revised version, we re-examined the potential role of FAK as an upstream regulator of ERK activation using the FAK inhibitor Ifebemtinib, rather than Defactinib as used in our original experiments. Our previous conclusion-that ERK activation was independent of FAK-was likely influenced by limitations associated with Defactinib, which did not properly reduce p-FAK levels despite lowering focal adhesion numbers, accompanied with an increase of ERK phosphorylation alongside a decrease of nuclear MITF levels. In contrast, Ifebemtinib treatment led to a more effective inhibition of FAK, as evidenced by a marked reduction in both p-FAK levels and focal adhesion number (new suppl. fig. 6B,C). Importantly, this was accompanied by a significant decrease in p-ERK levels (new fig. 6M,N), suggesting that FAK contributes to ERK activation in response to ECM molecules in our model. Furthermore, FAK inhibition similar to MEK and ERK inhibition, led to increased melanin production in MCs cultured on FN (new fig. 6O). These new data are now included in the revised version of the manuscript (line 360 ff., new fig. 6M-O, new suppl. fig. 6).
Nonetheless, this does not exclude the possibility that additional kinases and pathways, including Wnt signaling, may also be involved. We acknowledge this possibility in the revised discussion (lines 478-488).
- Rationale for selecting MITF for the study is very weak. Please justify in the discussion why authors have chosen to study MITF/ERK axis with a more logistic approach.*
We have focused central aspects of our analyses on MITF because it is a central regulator of MC differentiation, pigmentation, and survival, and its activity has previously been reported to be modulated by ERK. Considering the observed changes in pigmentation, proliferation, and gene expression in response to distinct ECM molecules, we hypothesized that MITF acts as a key integrator of these ECM-dependent signals. Our data indeed support this rationale: we detected ECM-type-dependent MITF levels and localization, and manipulating the ERK pathway altered MITF activity and associated functional outputs. Moreover, siRNA-mediated downregulation of MITF in MCs cultured on COL I led to a marked reduction in melanin content (revised fig. 4D). Together, these data emphasize that the ERK/MITF axis serves as a pathway that responds to extracellular cues and links these to MC behavior. For clarity, we have included an additional explanation on our rationale in the revised manuscript (lines 146-152).
- It is suggested to check for the changes in the transcriptomic profile of melanocytes upon culturing on different matrix to get a more comprehensive view associated with the molecular mechanisms involved.*
We fully agree with the reviewer on the importance of assessing the ECM-dependent transcriptomes of MCs. Therefore, we have now performed RNA sequencing to compare the transcriptomic profiles of MCs cultured on COL IV-, COL I- and FN-coated stiff substrates (line 258 ff. and revised fig. 5, new fig. 6, new suppl. fig. S5). This analysis provided a broader view of the molecular responses of MCs to ECM molecules and complemented our previous molecular and phenotypes analyses. The obtained transcriptomes confirmed significant modulation of genes associated with MC differentiation and pigmentation, as well as genes involved in signaling pathways such as MAPK/ERK and Wnt (see also answers to points 1-3). These findings help contextualize the ECM-dependent phenotypic changes and strengthen the mechanistic insights presented in the study.
- Please provide the protein expression of genes involved in cell cycle progression and/or apoptosis to support the data in Fig. 3D-E.*
To support the observations presented in original fig. 3, we employed immunostaining to assess the protein expression of Ki67, which is both a well-established marker and a protein involved in cell cycle progression (PMID: 28630280). In revised figure 3E, a significant reduction in the proportion of Ki67-positive cells on FN compared to COL I was observed, reinforcing our initial findings derived from BrdU incorporation assays and direct microscopic monitoring of cell division (revised fig. 3D,F).
In addition, global gene expression analysis revealed differentially expressed genes related to cell cycle regulation and apoptosis (revised fig. 5C,D), in line with the reduced proliferation observed. Notably, FN also triggered the differential expression of genes associated with cellular senescence (revised fig. 5E). Together, these data suggest that adhesion to FN induces a transcriptional and phenotypic shift in MCs toward a less-proliferative state that is associated with differential cell cycle modulation and signs of senescence.
Minor comment to the Authors:
- Discussion line 358-359, using term synergy is an overstatement as the collective data do not support the claim. Very little role of matrix stiffness is demonstrated by experimental data.*
We thank the reviewer for this comment and agree that the term "synergy" may overstate the conclusions drawn from the current dataset. We have therefore removed this term from the revised version of the manuscript to more accurately reflect the data.
- Method section, BrdU assay and BrdU assay-cell proliferation can be combined in method section.*
We have combined the descriptions of the BrdU assay and BrdU-based cell proliferation assay into a single, unified section in the Methods.
- What trigger melanocytes to respond to different microenvironment. Please discuss.*
To address this question, we have added the following paragraph to the Discussion (lines 377-380): "Our study identifies ECM components as critical environmental triggers that instruct MC behavior. Through dynamic interactions with the ECM, MCs engage adhesion-dependent signaling pathways, such as FAK activation, enabling them to decode contextual ECM inputs and adapt their phenotype accordingly."
- Fig 3C and 5D Tyr mRNA expression is tested. Authors should also test for the protein expression in the similar set of studies.*
We thank the reviewer for this suggestion and agree that assessing TYR protein expression would be valuable. However, we have encountered difficulties with the currently available antibodies and detection methods, which in our hands appeared unreliable for consistently detecting endogenous TYR protein levels in MCs under these conditions. For this reason, we relied on Tyr mRNA expression as a robust and reproducible readout and complemented this with functional assays such as melanin content measurement as a read-out that indirectly reflects TYR enzymatic activity. Of note, our transcriptomic analysis also revealed Tyr and other melanogenesis genes as differentially expressed genes when comparing MCs grown on COLI vs FN (revised fig. 5A,B).
- Line 217-218, Authors claim stiffness mediated increase of MITF nuclear localization in Col I, however Fig. 4A-B does not represent that claim. Please justify.*
Fig. 4A shows representative images of MCs cultured on stiff substrates coated with different ECM types, while the original figure 4B included the comparison across substrate stiffnesses for each ECM condition. We have now generated additional datasets to assess global MITF levels as well as nuclear localization across stiffness conditions in the presence of the different ECM types, demonstrating that nuclear MITF is significantly higher in cells cultured on stiff vs. soft or intermediate stiffness (revised fig. 4B,C). Of note, we do not detect a significant difference between soft and intermediate substrate stiffness, which could hint to a threshold of MITF dynamics in stiffness sensitivity. We have updated the figure legend and corresponding text to ensure the data presentation accurately supports our conclusions.
Significance:
Overall, the study is well-planned, the experiments are well-designed and executed with appropriate use of statistical analysis. However, a more in-depth analysis of the molecular mechanisms is necessary to clarify how the extracellular matrix (ECM) regulates ERK or MITF nuclear translocation.
We agree and feel that the additional data in the revised manuscript that explored transcriptional changes and the FAK/MEK/ERK/MITF axis in response to ECM proteins provide improved insights into ECM-mediated regulation of ERK and MC pigmentation.
This study enhances our existing knowledge by linking the well-established role of the extracellular matrix (ECM) in regulating ERK signaling to ERK's involvement in controlling MITF, a key regulator of melanocyte differentiation. It further establishes the ECM's role in controlling melanocyte function and differentiation.
This study will interest readers working in the field of the tumor microenvironment, as it explores the role of the extracellular matrix and its complexity and stiffness in disease progression, not only in melanoma but also in other types of cancer.
Reviewer #2 Evidence, reproducibility and clarity:
Summary:
In their manuscript, Luthold et al describe the behaviour of immortalized mouse melanocytes cultured on various extracellular matrix (ECM) proteins and substrates of different stiffness. They found that fibronectin, collagen IV and collagen I have different effects on melanocyte morphology, migration, and proliferation. They further link these differential effects to MITF localization and MEK/ERK signalling. This work shows that fibronectin supports melanocyte migration, which was associated with a dendritic morphology and correlated with increased MEK/ERK signalling and decreased MITF nuclear localization. In contrast, collagen I promoted melanocyte proliferation with low MEK/ERK signalling, enhanced MITF nuclear localization and high melanin production.
While this study is well designed and the data adequately presented and interpreted, the impact of its conclusions is limited by the incomplete mechanistic characterization of the observed phenotypes and by the lack of parallels with physiological conditions. To strengthen their manuscript, the authors should consider the following comments:
We also wish to thank this reviewer for the efforts made to assess our work and help us improve the study. We substantially revised the manuscript and now included e.g. bulk RNA sequencing and various loss-of-function approaches to better delineate the signaling pathways involved in ECM-dependent control of MCs.
Major comments to the Authors:
- Characterization of observed phenotypes:*
*The link between matrix-sensing and intracellular signalling is missing. Which types of integrins are expressed by iMCs? *
This is indeed an interesting point. Our RNA sequencing analysis indicates that MCs express integrins known to mediate adhesion to COL I and FN, including Itga2, Itga3, Itga5, Itgav, Itgb1, and Itgb3 (revised fig. 5K). Importantly, the expression of these integrins remains relatively consistent across ECM conditions (COL I, COL IV, and FN), suggesting that the phenotypic differences observed may not be directly explained by variations in integrin expression.
Are any of these integrins required for the observed phenotypes?
To assess a functional involvement, we conducted a pilot experiment blocking β1-integrin in MCs seeded on COL I and observed a marked reduction in MC adhesion (see associated graph 1, provided to this reviewer). However, the compromised cell spreading and resulting widespread detachment introduced confounding effects, making it difficult to interpret downstream events such as MITF nuclear localization. Since such readouts can be indirectly influenced by the overall adhesion state and associated signaling pathways such as FAK, we chose not to pursue further mechanistic analysis using this approach. Targeted strategies (e.g., inducible knockdown, acute protein degradation) will be needed in the future to dissect the precise role of individual integrins in mediating ECM-specific signaling responses in MCs.
Graph 1: Effect of β1-integrin blocking on MC adhesion. iMCs were detached using PBS-EDTA (10 min, 37 {degree sign}C) and incubated for 15 min on ice with either 10 μg/mL β1-integrin-blocking antibody (CD29, clone TS2/16; Invitrogen, #AB_1210468) or 10 μg/mL IgG isotype control. Cells (5,000 per well) were then seeded on COL I-coated substrates. After 1 h, non-adherent cells were gently washed off with PBS, and adherent cells were fixed with 4% PFA. Cell adhesion was quantified by counting the number of attached cells per µm² under a microscope.
The phenotypic changes described here are interesting but only partially analysed. Transcriptomic studies would yield a more complete view of cell state transitions (optional). At a minimum, could the authors detect any changes in cadherin expression, or in other genes classically involved in phenotype switching, such as twist1, snail or zeb1?
We thank the reviewer for this important suggestion, which helped to improve this manuscript. We have now performed bulk RNA sequencing to analyze global gene expression changes in MCs cultured on different ECM substrates (revised fig. 5, new suppl. fig. 5). Among these, we explored gene expression programs associated with MC plasticity and differentiation (revised fig. 5F-H): MCs cultured on FN exhibited reduced expression of melanocytic differentiation markers and upregulation of genes linked to plasticity, dedifferentiation, and neural crest-like features, suggesting a shift toward a less differentiated state, reflecting aspects of a phenotypic switch.
Nonetheless, as part of this analysis (but not included in the manuscript), we found that Zeb2, Snai2, and Zeb1 were expressed at similar levels across ECM conditions. Similarly, among the cadherins, Cdh1 and Cdh2 were not differentially expressed, albeit the overall low expression of Cdh1 showed a trend towards a reduction on COL I. Finally, Snai1, Twist1, and Twist2 were detected at very low levels and not significantly regulated as well. These data suggest that, at the chosen experimental conditions, while a clear adaptive phenotypic cell plasticity is observed, classical EMT-like programs are not prominently activated. However, we cannot exclude the possibility that longer culture durations or additional cues could induce such transitions.
Lines 235-236, the authors write that ECM proteins regulate melanocyte behaviour "likely through modulation of MITF localization and activity". Could the authors support the role of MITF experimentally? Genetic experiments using different MITF mutants could address this question.
To experimentally support the role of MITF, we now performed melanin assays following siRNA-mediated knockdown of MITF in MCs grown on COL I or FN. On COL I-coated substrates, MITF depletion led to a marked reduction in melanin content, supporting the conclusion that ECM-dependent regulation of pigmentation in our culture model involves MITF activity. These findings are now included in the revised manuscript (lines 244-245, revised fig. 4D, new fig. S4B).
*Additionally, how does MEK/ERK signalling control MITF activity in these melanocytes? The trametinib experiment should be consolidated with other inhibitors (including ERK inhibitors) and/or genetic manipulation. *
To address this comment, we complemented our former Trametinib experiments with ERK inhibition using Ravoxertinib (new fig. 6J-L). ERK inhibition led to increased nuclear localization of MITF and elevated melanin production, supporting the involvement of MEK/ERK in restraining MITF activity in MCs in response to ECM molecules. These new data are now included in the revised manuscript (line 354 ff. and new fig. 6J-L).
Did the authors also measure the effect of trametinib on cell proliferation in Figure 5?
Overall, compared to the observed pronounced phenotypes like ECM-dependent cell morphology, melanin production and others, the differences in cell proliferation of MCs grown at different ECM conditions were statistically significant but not very large. We therefore refrained from additionally assessing the effect of trametinib on the observed ECM-dependent MC behaviour. Given the well-established role of ERK signaling in promoting cell proliferation, we indeed expect that MEK inhibition can reduce MC proliferation in our system, though it remains open whether there is an ECM-specific aspect to this.
- Parallels with physiological conditions:*
*Most experiments shown were performed with immortalized melanocytes even though authors mention the use of primary cells (pMCs, line 148). Were similar results obtained in primary melanocytes? Do human melanocytes in culture behave similarly? *
While we have not assessed human MCs, original fig. S2 (__revised fig. S3) __provides data using primary murine MCs (freshly isolated from newborn mice), confirming a similar behavior of primary cells compared to immortalized MCs in terms of cell area, p-FAK levels, number of FAs, melanin production, and MITF nuclear localization.
Are some of these observations also true in vivo, for example in mouse skin (optional)?
The current manuscript focuses on the behavior of MCs in culture, as it was important to use a reductionist model system that can uncouple the effect of distinct ECM types as well as substrate stiffness. However, as a perspective and beyond the scope of this manuscript, we indeed plan to translate our in vitro findings to mouse skin, taking different biophysical and biochemical cues into account. Data from the present in vitro study provides valuable insights into which parameters and which anatomical areas to study in vivo.
How do the authors reconciliate their findings that collagen IV induced melanocyte migration and decreased proliferation and melanin production with the fact that melanocytes in human skin are generally in contact with the collagen IV-rich basement membrane?
We indeed regarded the use of collagen IV (COL IV) as a physiological reference condition, and considered MC migration, proliferation, and melanin production on COL as baseline levels. Relative to COL IV, COL I reduced migration and increased melanin production, while FN led to increased migration, and a decrease of proliferation and melanin production. This suggests that ECM composition can selectively modulate distinct aspects of MC behavior compared to attachment to COL IV. The intermediate state observed on COL IV would be in line with a model in which this abundant basement membrane molecule enables MCs to maintain high flexibility in their phenotype, e.g. to further increase melanin production upon external stimuli other than ECM (UV, inflammation etc.). The perhaps unexpected, opposing response of MCs to FN and COL I, respectively, opens the possibility that under specific (patho)physiological conditions, the then abundant ECM can direct MC behaviour. Both plasma- and cellular-derived FN is deposited upon skin injury and instructs various cell types to promote skin repair. Taking our observations in vitro into account, it is tempting to speculate that this FN-enriched tissue enables MCs to quickly migrate into wound sites to re-establish protection to UV. Conversely, increased COL I levels-as observed in fibrotic conditions such as scleroderma-might favor a more differentiated, pigment-producing phenotype. Interestingly, cases of localized hyperpigmentation have been reported in scleroderma patients, possibly reflecting such matrix-driven MC reprogramming. Though requiring further investigation, these observations open new avenues to explore how dynamic changes in ECM composition contribute to MC behavior in tissue homeostasis and repair.
We now extended our original discussion to better emphasize the physiological relevance of our findings (lines 383-391) and hypothesize how ECM remodeling may contribute to the dynamic regulation of MC plasticity-not only during tissue homeostasis, but also in response to injury and in fibrotic conditions such as scleroderma (lines 393-406).
Minor comments to the Authors:
The evidence that FAK is not responsible for MEK/ERK activation could be presented in the main text rather than in the discussion.
We thank the reviewer for highlighting this important point. Our initial conclusion-that ERK activation was independent of FAK-likely stemmed from limitations of the previously used FAK inhibitor (Defactinib). In those earlier experiments, while FAK inhibition reduced focal adhesion numbers, p-FAK levels were not properly decreased, and paradoxically, ERK phosphorylation increased alongside decreased nuclear MITF levels. Based on this initial discrepancy and because of this reviewer's comment, we performed additional experiments using another selective FAK inhibitor, Ifebemtinib, which achieved an effective reduction in both p-FAK levels and focal adhesion number (new suppl. fig. S6B, C). In the revised version, we present new experiments using Ifebemtinib, demonstrating that FAK inhibition in fact does reduce p-ERK levels (new fig. 6M-N), thus supporting the notion that FAK contributes to ECM-dependent ERK pathway activation in our model. These findings are now shown in the results section (lines 357-364).
Significance:
General assessment: This study establishes the cellular impact of different types of extracellular matrix proteins and stiffness conditions relevant to skin biology on the behaviour of untransformed mouse melanocytes. In particular, it shows opposite effects of fibronectin and collagen I on cell proliferation and migration, which could prove relevant to certain skin conditions in human. However, the scope of these results is limited by the incomplete mechanistic characterization of the observed phenotypes and by the lack of parallels with physiological conditions.
Advance: The systematic comparison of different microenvironmental conditions on normal melanocyte behaviour is novel and opens perspectives to understand the role of melanocytes in some human skin pathologies.
Audience: The comparison of different environmental conditions on melanocyte behaviour is of interest to the melanocyte biology community and could have implications for basic and clinical understanding of some skin diseases.
My expertise is in melanoma biology, including the impact of the microenvironment on tumour cell behaviours.
Reviewer #3 Evidence, reproducibility and clarity:
In this manuscript Luthold et al. describe how extracelluar matrix proteins and mechanosensation affect melanocyte differentiation. In particular, they show that ECM proteins and surface stiffness lead to effects on the MEK/ERK pathway, thus affecting the MITF transcription factor. The manuscript is interesting, well written and the data presented in a clear and easy-to-follow manner. The data are nicely quantitated and largely convincing.
- However, the discussion of the nuclear location of MITF (Figure 4A) is not convincing. The images presented show that upon exposure to ColI, there is a lot of MITF in the nucleus, a lot less so upon ColIV and none upon FN exposure. However, we only see a snapshot of the cells and thus we do not know if we are witnessing effects on MITF protein synthesis, degradation or nuclear localization (the least likely scenario since M-MITF, the isoform present in melanocytes is predominantly nuclear anyway). Was there a cytoplasmic signal detected? Upon FN treatment, there is no MITF protein visible in the cells. Does this mean that the protein is not made, that it is degraded or present at such low levels that the antibody does not detect it? The claim of the authors that this affects nuclear localization of MITF needs more corroboration. *
We thank the reviewer for raising this important point regarding the interpretation of MITF localization. We agree that the data as represented in the original figure 4 cannot distinguish whether changes reflect differences in MITF expression, stability, or subcellular distribution.
To better address this, we now included a quantitative analysis of both nuclear and cytoplasmic MITF signals (revised fig. 4B). These data show that MITF is detectable in both compartments at all conditions tested. While total MITF levels were not reduced on FN, nuclear MITF was markedly decreased and cytoplasmic MITF was even increased compared to COL I. This indicates that the reduced nuclear signal on FN compared to COL I is not due to an overall loss of MITF protein but rather reflects a shift in its subcellular distribution. These findings support the idea that ECM composition influences MITF localization, consistent with functional changes in its activity and with the observed phenotypic changes.
- Also, the authors need to show immunocytochemical images for the effects on MITF nuclear localization for the images presented in Figure 5C. *
As requested, we now provide representative micrographs illustrating the effects on MITF nuclear localization corresponding to the conditions shown in Fig. 5C. These images have been included in the revised version of the manuscript (new fig. 6G), further supporting the quantitative data presented.
- It seems that the authors quantitated immune-reactivity for both MITF and YAP. What was the control and how was the data normalized? *
MITF and YAP immunodetection were performed in separate experiments and were not analyzed in the same cells. For both stainings, secondary antibody controls were included (secondary antibody alone without primary antibody), which showed no detectable signal. For MITF and YAP quantifications (revised fig. 4B,F), nuclear (for both) and cytoplasmic (for MITF) intensity values were normalized within each independent experiment by dividing each individual measurement by the mean nuclear intensity across all conditions. This approach allowed us to deal with total signal variability between experiments while preserving relative differences between ECM conditions. For the percentage of nuclear MITF no normalization was applied. We have added this description to the revised methods section.
- Similarly, the blots and data shown in Figure 5 are not consistent with the text as described in the results section. The differences observed are minor and the only set that is likely to be significant is the FN-set; the differences between soft, intermediate and stiff of the FN-set do not look significantly different. The description of this in the results section should be toned down accordingly.*
To strengthen the conclusions drawn from the original Fig. 5 (now fig. 6), we performed additional immunoblot experiments to increase the number of replicates. These extended results now show a statistically significant increase in pERK levels in MCs cultured on FN compared to COL I. However, consistent with the reviewer's observation, no significant differences were detected across the stiffness conditions within FN. We have revised the Results section accordingly to tone down the interpretation and to better reflect the revised data (revised fig. 6E, lines 339-355).
Significance:
Upon improvement, this paper will provide an early characterization of the effects of the ECM on melanocyte differentiation. If the link to MITF holds, this will be the first time that mechanosensation has been shown to mediate effects on this transcription factor.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
In this manuscript Luthold et al. describe how extracelluar matrix proteins and mechanosensation affect melanocyte differentiation. In particular, they show that ECM proteins and surface stiffness lead to effects on the MEK/ERK pathway, thus affecting the MITF transcription factor. The manuscript is interesting, well written and the data presented in a clear and easy-to-follow manner. The data are nicely quantitated and largely convincing. However, the discussion of the nuclear location of MITF (Figure 4A) is not convincing. The images presented show that upon exposure to ColI, there is a lot of MITF in the nucleus, a lot less so upon ColIV and none upon FN exposure. However, we only see a snapshot of the cells and thus we do not know if we are witnessing effects on MITF protein synthesis, degradation or nuclear localization (the least likely scenario since M-MITF, the isoform present in melanocytes is predominantly nuclear anyway). Was there a cytoplasmic signal detected? Upon FN treatment, there is no MITF protein visible in the cells. Does this mean that the protein is not made, that it is degraded or present at such low levels that the antibody does not detect it? The claim of the authors that this affects nuclear localization of MITF needs more corroboration. Also, the authors need to show immunocytochemical images for the effects on MITF nuclear localization for the images presented in Figure 5C. It seems that the authors quantitated immune-reactivity for both MITF and YAP. What was the control and how was the data normalized? Similarly, the blots and data shown in Figure 5 are not consistent with the text as described in the results section. The differences observed are minor and the only set that is likely to be significant is the FN-set; the differences between soft, intermediate and stiff of the FN-set do not look significantly different. The description of this in the results section should be toned down accordingly.
Upon improvement, this paper will provide an early characterization of the effects of the ECM on melanocyte differentiation. If the link to MITF holds, this will be the first time that mechanosensation has been shown to mediate effects on this transcription factor.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary:
In this manuscript, authors demonstrated the role of ECM-dependent MEK/ERK/MITF signaling pathway that influences the plasticity of MCs (melanocytes) through their interactions with the environment. The findings emphasize the essential role of the extracellular matrix (ECM) in controlling MC function and differentiation, highlighting a critical need for further research to understand the complex interactions between mechanical factors and ECM components in the cellular microenvironment. Overall, the manuscript is concise, written well and shed light on a complex relationship between ECM protein types and substrate stiffness that affects MC mechanosensation. However, understanding detailed molecular mechanisms involved, especially the roles of MITF and other key regulators, is crucial for comprehending MC function and related pathologies. Authors needs to clarify some minor queries to be considered for publication.
Major comment:
Minor comment:
Overall, the study is well-planned, the experiments are well-designed and executed with appropriate use of statistical analysis. However, a more in-depth analysis of the molecular mechanisms is necessary to clarify how the extracellular matrix (ECM) regulates ERK or MITF nuclear translocation.
This study enhances our existing knowledge by linking the well-established role of the extracellular matrix (ECM) in regulating ERK signaling to ERK's involvement in controlling MITF, a key regulator of melanocyte differentiation. It further establishes the ECM's role in controlling melanocyte function and differentiation.
This study will interest readers working in the field of the tumor microenvironment, as it explores the role of the extracellular matrix and its complexity and stiffness in disease progression, not only in melanoma but also in other types of cancer.
C’est une évolution, par rapport au modèle du salariat. Mais ce n’est pas une révolution. Le travail indépendant a déjà été important en proportion avant que le salariat s’impose
Certes le travail indépendant était important en terme de volume avant l'ère industrielle. Cela dit cette conclusion mériterait une comparaison des types de métiers concernés et de la place des "donneurs d'ordre" des deux systèmes qui n'ont plus gra,d chose à voir l'un avec l'autre.
Les nouveaux entrants dans le monde du travail doivent choisir une activité. Ils doivent être prêts à évoluer. Ils doivent apprendre à se vendre sur les réseaux sociaux, les places de marché
Là l'auteur répond à la question de départ, mais du point de vue des travailleurs qui doivent s'adapter aux mutations technologiques et leur impact sur le monde du travail. En revanche le système n'est pas remis en question et apparaît comme inéluctable...
Aujourd’hui, l’évolution technologique libère l’individu qui a la possibilité de maîtriser sa vie, dans la tradition humaniste
Point de vue global et qui ne tient pas compte des phénomènes d'aliénation liés aux nouvelles technologies, pas plus qu'il ne tient compte des problèmes de santé mentale (burn-out) liés à l'absence de temps de déconnexion (imbrication du professionnel dans la sphère privée).
Une tendance est l’autonomisation de l’individu
L'autonomisation n'est vue que sous la forme du statut du travailleur (salarié, entrepreneur, indépendant, travailleur ubérisé...). Oui il y a plus de travailleur.euses indépendant.es, mais est ce un choix ? Le recours à la sous-traitance et à l'externalisation des activités s'est accompagné, dans de nombreuses entreprises et administrations, par des suppressions de postes (ménage, courrier, livraison, fonctions transverses...).
Une tendance est l’autonomisation de l’individu. Ce qui correspond à ce que nous avons vu auparavant : le travail est une prestation de service. Une époque a vu une tendance au salariat
Ce point de vue est très restrictif. Il ne tient pas compte de l'analyse marxiste du travail et de la notion de classe (ceux qui possèdent les moyens de production / ceux et celles qui vendent leur force de travail).
Par exemple, Amazon, symbole de la nouvelle économie, emploie beaucoup de manutentionnaires, symboles de la vieille économie. On a tendance à raisonner en termes de destruction créatrice, selon la théorie de Schumpeter : une nouvelle activité remplace une ancienne
Référence à Schumpeter, économiste du XXème siècle qui a beaucoup théorisé et écrit sur l'évolution de l'économie (fonctionnement par vagues, marquées par des innovations).
Il faut souligner qu’il se diversifie et se transforme. Il n’y a pas forcément de disparition d’emplois, de remplacements. La situation est très diversifiée
Si la question de la diversification et la transformation est juste, en revanche on peut trouver beaucoup d'études sur les disparitions d'emplois liées à la numérisation et la dématérialisation. Cette affirmation va à l'encontre de son argument sus-cité. Il est étrange qu'il parle dans le même texte de compétences entièrement faisables par les machines (conduisant à la possibilité de devenir indépendant) et en même temps de l'absence d'impact sur le volume d'emplois. Si l'on suit son raisonnement, que sont donc devenues les secrétaires des cabinets d'avocats qui classaient les dossiers, archivaient les décisions de justice, géraient les agendas ?
On constate ainsi une autonomisation du travailleur. De plus en plus, l’individu dispose à bon prix des moyens de mettre en œuvre une activité en tant qu’indépendant. Il n’a plus forcément besoin de la structure d’un cabinet d’avocat, de celle d’une entreprise. Il n’a plus besoin d’énormément de moyens pour communiquer, trouver des clients, même à l’étranger. D’où la résurgence du travail indépendant, qui est par exemple très visible aux États-Unis d’Amérique
Là il ne parle que des indépendants. De nombreuses études sur les conditions de travail (ANACT, expertises de CSE) ont montré à quel point les outils numériques étaient synonymes de perte d'autonomie, d'automatisation et d'aliénation au travail. De même la question de l'isolement (lié au fait que beaucoup de métiers pourraient de faire de manière individuelle) n'est pas posée alors qu'elle a des impacts réels sur la santé mentale des travailleuses et travailleurs concerné.es.
Il y a là un gain de productivité. Là où il fallait disposer de la place pour stocker toute l’information juridique, il suffit d’un ordinateur. Par conséquent, un juriste, un comptable, n’a plus forcément besoin de la puissance d’un cabinet qui pouvait mutualiser ce genre d’informations.
Gain de place et donc de productivité, oui puisque tout es monétisé dans une entreprise (y compris les m² occupés). Cela dit les veilles juridiques continuent d'exister, et celles-ci doivent être compilées, classées et archivées pour en faire ressortir facilement les jurisprudences intéressantes. De ce point de vue le lien de cause à effet que fait l'auteur est un peu rapide. Il ne tient pas compte du savoir-faire et des nomenclatures propres à chaque individu/collectif.
Car seule la simple mise à disposition de l’information est concernée. Un juriste par exemple n’utilisera pas une application informatique pour rédiger un mémoire, mais il utilisera l’information dont il peut désormais disposer avec facilité.
Cette partie du texte montre à quel point il est obsolète. Aujourd’hui l'IA permet de rédiger à la "manière de", et en tout cas de proposer un premier jet structuré faisant gagner un temps considérable. Par ailleurs les avocat.es peuvent utiliser des mémoires précédemment rédigés (on ne réinvente pas l'eau chaude), en les adaptant à l'affaire qu'ils traitent.
Ainsi, le big data est le traitement de données. Ce traitement est ensuite mis à disposition d’un certain nombre de personnes. De même, les logiciels de conception industrielle traitent une multitude de données, et donnent un résultat qui retranscrit les relations entre elles. De même, un simple tableur traite des données, et met à disposition une synthèse sous forme de tableaux. Idem pour les logiciels comptables, qui traitent les données qu’on y entre, et peuvent ressortir toutes sortes de tableaux et synthèses. On y trouve même les factures dématérialisées. Enfin, on a aujourd’hui toutes sortes d’informations mises à disposition. Là où il fallait posséder une grande bibliothèque, renouvelée tous les ans pour être à jour des codes juridiques en tout genre, il suffit d’aller sur internet. Une information gratuite, ou moins onéreuse que l’information-papier d’autrefois, est disponible. Sans compter qu’elle est actualisée beaucoup plus souvent. Le livreur de pizza utilise aussi une application de traitement et de mise à disposition de l’information. Il entre une adresse, une application analyse une carte avec les sens de circulation, et ressort un itinéraire. Tout comme le chauffeur de VTC. Tout comme le restaurateur qui scrute les variations de météo sur son smartphone. Et une place de marché met en relation acheteurs et vendeurs. Les vendeurs de biens ou de services peuvent être notés. Ils indiquent leurs disponibilités. À nouveau un traitement et une mise à disposition d’informations
Dans ce chapitre l'auteur met en avant les facilités dans le travail permises par les nouvelles technologies, ainsi que le gain de temps grâce à celles-ci. En revanche s'il pose bien la question de partage et de transit de données, il ne se positionne pas sur la question de la confidentialité et de l'utilisation qui en sont faites (traitement des datas, reventes des données à des sociétés privées à des fins commerciales par exemple). Par ailleurs l'affirmation selon laquelle l'information est gratuite est erronée. En effet l'accès à certaines informations est soumis à abonnement (payant) ou au paiement d'un droit d'accès temporaire.
Elle tend au traitement et à la mise à disposition de l’information. D’où ma préférence pour la terminologie technologies de l’information et de la communication, ou TIC.
lien de causalité entre la réponse à la question argumentative et le parti pris d'utiliser l'expression TIC plutôt que big data
Enfin, on voit le développement de place de marché où chacun peut proposer ses compétences. Une foule de métiers est concernée. Des plus qualifiés, comme des aides ménagères ou pour le bricolage. Ou simplement partager son logement ou sa voiture. Les nouvelles technologies de l’information et de la communication irriguent toute l’économie
Il appuie son argumentation en donnant des exemples dans des secteurs très divers pour montrer qu'aucun secteur n'est épargné : argument épistémique
Cela fait longtemps que dans les grandes surfaces la gestion des stocks se fait en temps réel : le passage en caisse met à jour automatiquement le stock. L’information juridique est disponible sur le web. Là où il fallait soit une connaissance encyclopédique, soit une multitude de codes, il suffit aujourd’hui de rechercher l’information sur internet
L'auteur semble dire que le savoir et le savoir-faire sont remplacés par les outils numériques. Si certaines tâches sont effectivement remplacées par ces outils, cela n'empêche le besoin de connaissances a priori sur les process industriels ou de pensée.
Aujourd’hui, l’expression à la mode est big data. Ce qui traduit le fait que l’informatique actuelle est capable de traiter des montagnes de données (data en anglais). Ce qui permet de trouver des corrélations entre vos consultations internet et des propositions commerciales par exemple
L'auteur semble ne pas adhérer à cette expression, la qualifiant de "à la mode".
Un article de l’IREF souligne que l’emploi dans le numérique ne représente que 5 % du total de l’emploi privé aux États-Unis, selon les chiffres du Bureau of Labor Statistics. Ce qui montre que ce qui est appelé la révolution numérique n’est pas une révolution en termes de volume d’emplois, mais parce qu’elle se diffuse à travers toute l’économie. Tous les secteurs, des plus qualifiés aux moins qualifiés, sont touchés
Le volume d'emplois n'est vu que sous le prisme de métiers du numérique et non de l'impact réel sur les emplois dans tous les secteurs. Cela ne dit rien sur les suppressions d'emplois liées à la dématérialisation, l'automatisation et la numérisation.
L’impact de la révolution numérique sur le travail
question argumentative dans le titre. Titre assez neutre qui n'impose pas d'hypoyhèse.
Discuta o documento "Contrato de aprendizagem". Coloque questões, participe nos comentários dos colegas. Estude o documento e deixe um comentário final sobre as suas espetativas na disciplina
NA
There's a missing comma after this word.
Vista errorea del Data Frame para guardar el torneo Pokemon. Si que quiere ver la vista correcta hay que cliquera la soalapa “Tree”, en lugar de la solapa “Data”, en la parte superior.
Una vez hecho el ejercicio en GT y agregado el DataFrame, y cliquear en la solapa Tree, me salen los datos vacíos, no se si es que falta agregar una parte de código para ejecutar algo y organizarlos.
pokemonDataDictionary value: 'ditto'
Entiendo que los diccionarios traen los datos de de los valores desde la PokeApi. Pero en este caso, no me queda claro ¿cómo se da esta relación? Osea, cómo llego a que la Api me traiga los valores de un pokemon determinado?
:item
Doncente @offray, que la palabra ítem se encuentre antecedida por los dos puntos indica que es como el elemento que vamos a manipular a través del mensaje?
pokemonDataDictionary := [ :name | | dataLink pokemonRawData | dataLink := 'https://pokeapi.co/api/v2/pokemon/', name. pokemonRawData := dataLink asUrl retrieveContents. STONJSON fromString: pokemonRawData ]
Docente @Offray, sobre el siguiente bloque tengo las siguientes inquietudes: 1. técnicamente que significa := ? ya que esta siempre antecede un bloque. 2. con base en el documento Representando y procesando datos en Pharo no veo ejemplos como :name donde los dos puntos anteceden el valor, técnicamente que significa dicho mensaje dentro del boque, es lo referente al elemento que presentará los cambios dentro de un mensaje?
okemonDataDictionary value: 'pikachu'
Durante el ejercicio en Pharo con el documento importado en Glamorus, genera algunos errores al momento de aplicarlo (inspect), sin embargo, se ha corregido moviendo algunos espacios para la correcta ejecución, lo cual se debe validar como resultado de la importación del documento.
#('Eduar Daza' 'Nestor Cristancho' 'Maxi López-Gómez' 'Valentina Vanegas' 'Rubén Torres' 'Thomas Martínez' 'Lizeth Colorado' 'Dario Montenegro' 'Juan Pablo Arias Romero' 'Valentina Penagos Ariza')
En el presente caso podemos ver un ARREGLO, el cual está representado por #( ) y que son maneras de guardar colecciones de información diversa como en este caso se agrupan los repositorios de la clase Unisemánticas. Para mayor información ver el apartado 4.1. del documento [Representando y procesando datos en Pharo] (https://mutabit.com/repos.fossil/labci/doc/25A1/wiki/representando-y-procesando-datos-en-pharo--cek90.md.html)
pokemonDataDictionary := STONJSON fromString: pokemonRawData
A través de este código (STONJSON) podremos sustraer los datos de manera ligera a fin de contar con el diccionario de los datos., por otro lado, para tener una estructura de los datos se recomienda el navegador Mozilla Firefox, toda vez que tiene una interfaz nativa para ver los datos con más facilidad.
A continuación se ilustra un ejemplo de la visualización de JSON:
pokemonExampleLink asUrl retrieveContents
La siguiente operación implica un Receptor(pokemonExampleLink), un Selector (asUrl) y un Argumento (retrieveContents) donde básicamente se interpreta que la operación pokemonRawData indique que la cadena pokemonExampleLink como URL permita la sustracción de los contenidos de https://pokeapi.co/api/v2/pokemon/pikachu
Pokemon
Pokémon es una de las franquicias más populares del mundo del entretenimiento, originada en Japón. El nombre viene de la contracción de "Pocket Monsters" (Monstruos de Bolsillo) y fueron creados en 1996.

ormato ampliamente utilizado en el intercambio de datos en la web
Dicho formato (JSON) también es comúnmente utilizado para el desarrollo de interfaces de programación de aplicaciones (API REST), por ejemplo, en las Bibliotecas universitarias se utiliza usualmente para el vaciado de datos entre los sistemas de información académicos con los ILS para descargar/actualizar los datos de los estudiantes matriculados en una Universidad sin necesidad de hacer cargas por archivo txt, xls, etc.
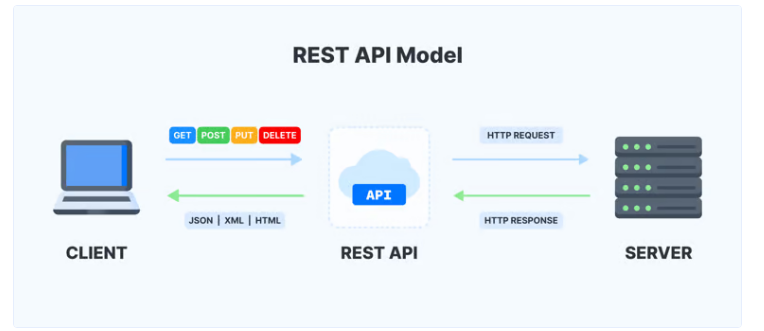
atRandom
En Pharo, atRandom es un método común que significa "elige un elemento aleatorio" que en este caso se utiliza para la selección aleatoria de los repositorios de los estudiantes, sin embargo, será utilizada para seleccionar aleatoriamente los datos de los Pokemon albergados en Pokeapi.
Author response:
The following is the authors’ response to the original reviews
Reviewer 1 (Public review):
Summary:
Gene transfer agent (GTA) from Bartonella is a fascinating chimeric GTA that evolved from the domestication of two phages. Not much is known about how the expression of the BaGTA is regulated. In this manuscript, Korotaev et al noted the structural similarity between BrrG (a protein encoded by the ror locus of BaGTA) to a well-known transcriptional anti-termination factor, 21Q, from phage P21. This sparked the investigation into the possibility that BaGTA cluster is also regulated by anti-termination. Using a suite of cell biology, genetics, and genome-wide techniques (ChIP-seq), Korotaev et al convincingly showed that this is most likely the case. The findings offer the first insight into the regulation of GTA cluster (and GTA-mediated gene transfer) particularly in this pathogen Bartonella. Note that anti-termination is a well-known/studied mechanism of transcriptional control. Anti-termination is a very common mechanism for gene expression control of prophages, phages, bacterial gene clusters, and other GTAs, so in this sense, the impact of the findings in this study here is limited to Bartonella.
Strengths:
Convincing results that overall support the main claim of the manuscript.
Weaknesses:
A few important controls are missing.
We sincerely appreciate reviewer #1's positive assessment of our manuscript. In response to the concern regarding control samples/experiments, we have addressed this issue in our revision, by providing data of the replicates of our experiments. We acknowledge that antitermination is a well-established mechanism of expression control in bacteria, including bacterial gene clusters, phages, prophages, and at least one other GTA. As reviewer #2 also noted, our study presents a unique example of phage co-domestication, where antitermination integrates both phage remnants at the regulatory level. We have emphasized this original aspect more clearly in the revised manuscript.
Reviewer 1 (Recommendations for the authors):
(1) Provide Rsmd and DALI scores to show how similar the AlphaFold-predicted structures of BrrG are to other anti-termination factors. This should be done for Fig1B and also for Suppl. Fig 1 to support the claim that BrrG, GafA, GafZ, Q21 share structural features.
In the revised manuscript we provide Rsmd and DALI scores in the supplementary Fig. 1A (Suppl. Fig. 1A). In Suppl. Fig. 1B we further include a heatmap of similiarity values.
(2) Throughout the manuscript, flow cytometry data of gfp expression was used and shown as single replicate. Korotaev et al wrote in the legends that error bars are shown (that is not true for e.g. Figs. 3, 4, and 5). It is difficult for reviewers/readers to gauge how reliable are their experiments.
In the revised manuscript we show all replicates for the flow cytometry histograms.
For Fig. 2C, all replicates are provided in Suppl. Fig. 3.
For Fig. 3B, all replicates are provided in Suppl. Fig. 4.
For Fig. 4B, all replicates are provided in Suppl. Fig. 5.
For Fig. 5B, all replicates are provided in Suppl. Fig. 6.
(3) I am unsure how ChIP-seq in Fig. 2A was performed (with anti-FLAG or anti-HA antibodies? I cannot tell from the Materials & Methods). More importantly, I did not see the control for this ChIP-seq experiment. If a FLAG-tagged BrrG was used for ChIP-seq, then a WT non-tagged version should be used as a negative control (not sequencing INPUT DNA), this is especially important for anti-terminator that can co-travel with RNA polymerase. Please also report the number of replicates for ChIP-seq experiments.
Fig. 2A presents the coverage plot from the ChIP-Seq of ∆brrG +pPtet:3xFLAG-brrG (N’ in green). As anticipated by the referee, we had used ∆brrG +pTet:brrG (untagged) as control (grey). Each strain was tested in a single replicate. The C-terminal tag produced results similar to the untagged version, suggesting it is non-functional. All tested tags are shown in Supplementary Figure 2.
(4) Korotaev et al mentioned that BrrG binds to DNA (as well as to RNA polymerase). With the availability of existing ChIP-seq data, the authors should be able to locate the DNA-binding element of BrrG, this additional information will be useful to the community.
We identified a putative binding site of BrrG using our ChIP-Seq data. The putative binding site is indicated in Fig. 2D of the revised manuscript.
(5) Mutational experiments to break the potential hairpin structure are required to strengthen the claim that this putative hairpin is the potential transcriptional terminator.
We did not claim the identified hairpin is a confirmed terminator, but proposed it as a candidate. We agree with the referee that the suggested experiment would be necessary to definitively establish its function. However, our main objective was to show that BrrG acts as a processive terminator, which we demonstrated by replacing the putative terminator with a well-characterized synthetic one that BrrG successfully bypassed. Therefore, we chose not to perform the proposed experiment and have accordingly softened our conclusions regarding the hairpin’s potential terminator function.
Reviewer 2 (Public review):
Summary:
In this study, the authors identified and characterized a regulatory mechanism based on transcriptional anti-termination that connects the two gene clusters, capsid and run-off replication (ROR) locus, of the bipartite Bartonella gene transfer agent (GTA). Among genes essential for GTA functionality identified in a previous transposon sequencing project, they found a potential antiterminatior of phage origin within the ROR locus. They employed fluorescence reporter and gene transfer assays of overexpression and knockout strains in combination with ChiPSeq and promoter-fusions to convincingly show that this protein indeed acts as an antiterminator counteracting attenuation of the capsid gene cluster expression.
Impact on the field:
The results provide valuable insights into the evolution of the chimeric BaGTA, a unique example of phage co-domestication by bacteria. A similar system found in the other broadly studied Rhodobacterales/Caulobacterales GTA family suggests that antitermination could be a general mechanism for GTA control.
Strengths:
Results of the selected and carefully designed experiments support the main conclusions.
Weaknesses:
It remains open why overexpression of the antiterminator does not increase the gene transfer frequency.
We are grateful for reviewer #2's thoughtful and encouraging feedback on our manuscript. The reviewer raises an important question about why overexpression of the antiterminator does not increase gene transfer frequency. While we acknowledge this point, we consider it beyond the scope of the current study. Our findings clearly demonstrate that the antiterminator induces capsid component expression in a large proportion of cells. However, the fact that this expression plateaus at high levels rather than exhibiting a transient peak, as seen in the wild type, suggests that antiterminators do not regulate GTA particle release via lysis. We are actively investigating this further through additional experiments, which we plan to publish separately from this study.
Reviewer 2 (Recommendations for the authors):
(1) The authors wrote "GTAs are not self-transmitting because the DNA packaging capacity of a GTA particle is too small to package the entire gene cluster encoding it" (page 3). I thought that at least the Bartonella capsid gene cluster should be self-transmissible within the 14 kb packaged DNA (https://doi.org/10.1371/journal.pgen.1003393, https://doi.org/10.1371/journal.pgen.1000546). This was also concluded by Lang et al (https://doi.org/10.1146/annurev-virology-101416-041624). In this case the presented results would have important implications. As the gene cluster and the anti-terminator required for its expression are separated on the chromosome, it would not be possible to transfer an active GTA gene cluster, although the DNA coding for the genes required for making the packaging agent itself, theoretically fits into a BaGTA particle. Could the authors comment on that? I think it would be helpful to add the sizes of the different gene clusters and the distance between them in Fig. 2A. The ROR amplified region spans 500kb, is the capsid gene cluster within this region?
We thank the reviewer for bringing up this interesting point. The ror gene cluster, which encodes the antiterminator BrrG, is approximately 9.2 kb in size and could feasibly be packaged in its entirety into a GTA particle. In contrast, the bgt cluster (capsid cluster) is approximately 20 kb in size —exceeding the packaging limit of GTA particles—and is separated from the bgt cluster by approximately 35 kb. Consequently, if the ror cluster is transferred via a GTA particle into a recipient host that does not encode the bgt gene cluster, the ror cluster would not be expressed.
We added the sizes of the gene clusters to Fig. 1A.
(2) Another side-note regarding the introduction: On page three the authors write: "GTAs encode bacteriophage-like particles and in contrast to phages transfer random pieces of host bacterial DNA". While packaging is not specific, certain biases in the packaging frequency are observed in both studied GTA families. For Bartonella this is ROR. In the two GTA-producing strains D. shibae and C. crescentus origin and terminus of replication are not packaged and certain regions are overrepresented (https://doi.org/10.1093/gbe/evy005, https://doi.org/10.1371/journal.pbio.3001790). Furthermore, D. shibae plasmids are not packaged but chromids are. I think the term "random" does not properly describe these observations. I would suggest using "not specific" instead.
We thank the reviewer for this suggestion and adjusted the wording on p. 3 accordingly.
(3) Page 5: Remove "To address this". It is not needed as you already state "To test this hypothesis" in the previous sentence.
We adjusted the working on p.5 accordingly.
(4) I think the manuscript would greatly benefit from a summary figure to visualize the Q-like antiterminator-dependent regulatory circuit for GTA control and its four components described on pages 15 and 16.
We thank the reviewer for this valuable suggestion. We included a summary figure (Fig. 6) in the discussion section of the revised manuscript.
(5) Page 17: It might be worth noting that GafA is highly conserved along GTAs in Rhodobacterales (https://doi.org/10.3389/fmicb.2021.662907) and so is probably regulatory integration into the ctrA network (https://doi.org/10.3389/fmicb.2019.00803). It's an old mechanism. It would be also interesting to know if it is a common feature of the two archetypical GTAs that the regulator is not part of the cluster itself.
We agree with the reviewer’s comments and have revised the wording to state that GafA is highly conserved.
Author response:
The following is the authors’ response to the previous reviews
General Response to Reviewers:
We thank the Reviewers for their comments, which continue to substantially improve the quality and clarity of the manuscript, and therefore help us to strengthen its message while acknowledging alternative explanations.
All three reviewers raised the concern that we have not proven that Rab3A is acting on a presynaptic mechanism to increase mEPSC amplitude after TTX treatment of mouse cortical cultures. The reviewers’ main point is that we have not shown a lack of upregulation of postsynaptic receptors in mouse cortical cultures. We want to stress that we agree that postsynaptic receptors are upregulated after activity block in neuronal cultures. However, the reviewers are not acknowledging that we have previously presented strong evidence at the mammalian NMJ that there is no increase in AChR after activity blockade, and therefore the requirement for Rab3A in the homeostatic increase in quantal amplitude points to a presynaptic contribution. We agree that we should restrict our firmest conclusions to the data in the current study, but in the Discussion we are proposing interpretations. We have added the following new text:
“The impetus for our current study was two previous studies in which we examined homeostatic regulation of quantal amplitude at the NMJ. An advantage of studying the NMJ is that synaptic ACh receptors are easily identified with fluorescently labeled alpha-bungarotoxin, which allows for very accurate quantification of postsynaptic receptor density. We were able to detect a known change due to mixing 2 colors of alpha-BTX to within 1% (Wang et al., 2005). Using this model synapse, we showed that there was no increase in synaptic AChRs after TTX treatment, whereas miniature endplate current increased 35% (Wang et al., 2005). We further showed that the presynaptic protein Rab3A was necessary for full upregulation of mEPC amplitude (Wang et al., 2011). These data strongly suggested Rab3A contributed to homeostatic upregulation of quantal amplitude via a presynaptic mechanism. With the current study showing that Rab3A is required for the homeostatic increase in mEPSC amplitude in cortical cultures, one interpretation is that in both situations, Rab3A is required for an increase in the presynaptic quantum.”
The point we are making is that the current manuscript is an extension of that work and interpretation of our findings regarding the variability of upregulation of postsynaptic receptors in our mouse cortical cultures further supports the idea that there is a Rab3Adependent presynaptic contribution to homeostatic increases in quantal amplitude.
Public Reviews:
Reviewer #1 (Public review):
Koesters and colleagues investigated the role of the small GTPase Rab3A in homeostatic scaling of miniature synaptic transmission in primary mouse cortical cultures using electrophysiology and immunohistochemistry. The major finding is that TTX incubation for 48 hours does not induce an increase in the amplitude of excitatory synaptic miniature events in neuronal cortical cultures derived from Rab3A KO and Rab3A Earlybird mutant mice. NASPM application had comparable effects on mEPSC amplitude in control and after TTX, implying that Ca2+-permeable glutamate receptors are unlikely modulated during synaptic scaling. Immunohistochemical analysis revealed no significant changes in GluA2 puncta size, intensity, and integral after TTX treatment in control and Rab3A KO cultures. Finally, they provide evidence that loss of Rab3A in neurons, but not astrocytes, blocks homeostatic scaling. Based on these data, the authors propose a model in which neuronal Rab3A is required for homeostatic scaling of synaptic transmission, potentially through GluA2-independent mechanisms.
The major finding - impaired homeostatic up-scaling after TTX treatment in Rab3A KO and Rab3 earlybird mutant neurons - is supported by data of high quality. However, the paper falls short of providing any evidence or direction regarding potential mechanisms. The data on GluA2 modulation after TTX incubation are likely statistically underpowered, and do not allow drawing solid conclusions, such as GluA2-independent mechanisms of up-scaling.
The study should be of interest to the field because it implicates a presynaptic molecule in homeostatic scaling, which is generally thought to involve postsynaptic neurotransmitter receptor modulation. However, it remains unclear how Rab3A participates in homeostatic plasticity.
Major (remaining) point:
(1) Direct quantitative comparison between electrophysiology and GluA2 imaging data is complicated by many factors, such as different signal-to-noise ratios. Hence, comparing the variability of the increase in mini amplitude vs. GluA2 fluorescence area is not valid. Thus, I recommend removing the sentence "We found that the increase in postsynaptic AMPAR levels was more variable than that of mEPSC amplitudes, suggesting other factors may contribute to the homeostatic increase in synaptic strength." from the abstract.
We have not removed the statement, but altered it to soften the conclusion. It now reads, “We found that the increase in postsynaptic AMPAR levels in wild type cultures was more variable than that of mEPSC amplitudes, which might be explained by a presynaptic contribution, but we cannot rule out variability in the measurement.”.
Similarly, the data do not directly support the conclusion of GluA2-independent mechanisms of homeostatic scaling. Statements like "We conclude that these data support the idea that there is another contributor to the TTX- induced increase in quantal size." should be thus revised or removed.
This particular statement is in the previous response to reviewers only, we deleted the sentence that starts, “The simplest explanation Rab3A regulates a presynaptic contributor….”. and “Imaging of immunofluorescence more variable…”. We deleted “ our data suggest….consistently leads to an increase in mEPSC amplitude and sometimes leads to….” We added “…the lack of a robust increase in receptor levels leaves open the possibility that there is a presynaptic contributor to quantal size in mouse cortical cultures. However, the variability could arise from technical factors associated with the immunofluorescence method, and the mechanism of Rab3A-dependent plasticity could be presynaptic for the NMJ and postsynaptic for cortical neurons.”
Reviewer #2 (Public review):
I thank the authors for their efforts in the revision. In general, I believe the main conclusion that Rab3A is required for TTX-induced homeostatic synaptic plasticity is wellsupported by the data presented, and this is an important addition to the repertoire of molecular players involved in homeostatic compensations. I also acknowledge that the authors are more cautious in making conclusions based on the current evidence, and the structure and logic have been much improved.
The only major concern I have still falls on the interpretation of the mismatch between GluA2 cluster size and mEPSC amplitude. The authors argue that they are only trying to say that changes in the cluster size are more variable than those in the mEPSC amplitude, and they provide multiple explanations for this mismatch. It seems incongruous to state that the simplest explanation is a presynaptic factor when you have all these alternative factors that very likely have contributed to the results. Further, the authors speculate in the discussion that Rab3A does not regulate postsynaptic GluA2 but instead regulates a presynaptic contributor. Do the authors mean that, in their model, the mEPSC amplitude increases can be attributed to two factors- postsynaptic GluA2 regulation and a presynaptic contribution (which is regulated by Rab3A)? If so, and Rab3A does not affect GluA2 whatsoever, shouldn't we see GluA2 increase even in the absence of Rab3A? The data in Table 1 seems to indicate otherwise.
The main body of this comment is addressed in the General Response to Reviewers. In addition, we deleted text “current data, coupled with our previous findings at the mouse neuromuscular junction, support the idea that there are additional sources contributing to the homeostatic increase in quantal size.” We added new text, so the sentence now reads: “Increased receptors likely contribute to increases in mESPC amplitudes in mouse cortical cultures, but because we do not have a significant increase in GluA2 receptors in our experiments, it is impossible to conclude that the increase is lacking in cultures from Rab3A<sup>-/-</sup> neurons.”
I also question the way the data are presented in Figure 5. The authors first compare 3 cultures and then 5 cultures altogether, if these experiments are all aimed to answer the same research question, then they should be pooled together. Interestingly, the additional two cultures both show increases in GluA2 clusters, which makes the decrease in culture #3 even more perplexing, for which the authors comment in line 261 that this is due to other factors. Shouldn't this be an indicator that something unusual has happened in this culture?
Data in this figure is sufficient to support that GluA2 increases are variable across cultures, which hardly adds anything new to the paper or to the field.
A major goal of performing the immunofluorescence measurements in the same cultures for which we had electrophysiological results was to address the common impression that the homeostatic effect itself is highly variable, as the reviewer notes in the comment “…GluA2 increases are variable across cultures…” Presumably, if GluA2 increases are the mechanism of the mEPSC amplitude increases, then variable GluA2 increases should correlate with variable mEPSC amplitude increases, but that is not what we observed. We are left with the explanation that the immunofluorescence method itself is very variable. We have added the point to the Discussion, which reads, “the variability could arise from technical factors associated with the immunofluorescence method, and the mechanism of Rab3A-dependent homeostatic plasticity could be presynaptic for the NMJ and postsynaptic for cortical neurons.”
Finally, the implication of “Shouldn’t this be an indicator that something unusual has happened in this culture?” if it is not due to culture to culture variability in the homeostatic response itself, is that there was a technical problem with accurately measuring receptor levels. We have no reason to suspect anything was amiss in this set of coverslips (the values for controls and for TTX-treated were not outside the range of values in other experiments). In any of the coverslips, there may be variability in the amount of primary anti-GluA2 antibody, as this was added directly to the culture rather than prepared as a diluted solution and added to all the coverslips. But to remove this one experiment because it did not give the expected result is to allow bias to direct our data selection.
The authors further cite a study with comparable sample sizes, which shows a similar mismatch based on p values (Xu and Pozzo-Miller 2007), yet the effect sizes in this study actually match quite well (both ~160%). P values cannot be used to show whether two effects match, but effect sizes can. Therefore, the statement in lines 411-413 "... consistently leads to an increase in mEPSC amplitudes, and sometimes leads to an increase in synaptic GluA2 receptor cluster size" is not very convincing, and can hardly be used to support "the idea that there are additional sources contributing to the homeostatic increase in quantal size.”
We have the same situation; our effect sizes match (19.7% increase for mEPSC amplitude; 18.1% increase for GluA2 receptor cluster size, see Table 1), but in our case, the p value for receptors does not reach statistical significance. Our point here is that there is published evidence that the variability in receptor measurements is greater than the variability in electrophysiological measurements. But we have softened this point, removing the sentences containing “…consistently leads and sometimes...” and “……additional sources contributing…”.
I would suggest simply showing mEPSC and immunostaining data from all cultures in this experiment as additional evidence for homeostatic synaptic plasticity in WT cultures, and leave out the argument for "mismatch". The presynaptic location of Rab3A is sufficient to speculate a presynaptic regulation of this form of homeostatic compensation.
We have removed all uses of the word “mismatch,” but feel the presentation of the 3 matched experiments, 23-24 cells (Figure 5A, D), and the additional 2 experiments for a total of 5 cultures, 48-49 cells (Figure 5C, F), is important in order to demonstrate that the lack of statistically significant receptor response is due neither to a variable homeostatic response in the mEPSC amplitudes, nor to a small number of cultures.
Minor concerns:
(1) Line 214, I see the authors cite literature to argue that GluA2 can form homomers and can conduct currents. While GluA2 subunits edited at the Q/R site (they are in nature) can form homomers with very low efficiency in exogenous systems such as HEK293 cells (as done in the cited studies), it's unlikely for this to happen in neurons (they can hardly traffic to synapses if possible at all).
We were unable to identify a key reference that characterized GluA2 homomers vs. heteromers in native cortical neurons, but we have rewritten the section in the manuscript to acknowledge the low conductance of homomers:
“…to assess whether GluA2 receptor expression, which will identify GluA2 homomers and GluA2 heteromers (the former unlikely to contribute to mEPSCs given their low conductance relative to heteromers (Swanson et al., 1997; Mansour et al., 2001)…”
(2) Lines 221-222, the authors may have misinterpreted the results in Turrigiano 1998. This study does not show that the increase in receptors is most dramatic in the apical dendrite, in fact, this is the only region they have tested. The results in Figures 3b-c show that the effect size is independent of the distance from soma.
Figure 3 in Turrigiano et al., shows that the increase in glutamate responsiveness is higher at the cell body than along the primary dendrite. We have revised our description to indicate that an increase in responsiveness on the primary dendrite has been demonstrated in Turrigiano et al. 1998.
“We focused on the primary dendrite of pyramidal neurons as a way to reduce variability that might arise from being at widely ranging distances from the cell body, or, from inadvertently sampling dendritic regions arising from inhibitory neurons. In addition, it has been shown that there is a clear increase in response to glutamate in this region (Turrigiano et al., 1998).”
“…synaptic receptors on the primary dendrite, where a clear increase in sensitivity to exogenously applied glutamate was demonstrated (see Figure 3 in (Turrigiano et al., 1998)).
(3) Lines 309-310 (and other places mentioning TNFa), the addition of TNFa to this experiment seems out of place. The authors have not performed any experiment to validate the presence/absence of TNFa in their system (citing only 1 study from another lab is insufficient). Although it's convincing that glia Rab3A is not required for homeostatic plasticity here, the data does not suggest Rab3A's role (or the lack of) for TNFa in this process.
We have modified the paragraph in the Discussion that addresses the glial results, to describe more clearly the data that supported an astrocytic TNF-alpha mechanism: “TNF-alpha accumulates after activity blockade, and directly applied to neuronal cultures, can cause an increase in GluA1 receptors, providing a potential mechanism by which activity blockade leads to the homeostatic upregulation of postsynaptic receptors (Beattie et al., 2002; Stellwagen et al., 2005; Stellwagen and Malenka, 2006).”
We have also acknowledged that we cannot rule out TNF-alpha coming from neurons in the cortical cultures: “…suggesting the possibility that neuronal Rab3A can act via a non-TNF-alpha mechanism to contribute to homeostatic regulation of quantal amplitude, although we have not ruled out a neuronal Rab3A-mediated TNF-alpha pathway in cortical cultures.”
Reviewer #3 (Public review):
This manuscript presents a number of interesting findings that have the potential to increase our understanding of the mechanism underlying homeostatic synaptic plasticity (HSP). The data broadly support that Rab3A plays a role in HSP, although the site and mechanism of action remain uncertain.
The authors clearly demonstrate that Rab3A plays a role in HSP at excitatory synapses, with substantially less plasticity occurring in the Rab3A KO neurons. There is also no apparent HSP in the Earlybird Rab3A mutation, although baseline synaptic strength is already elevated. In this context, it is unclear if the plasticity is absent, already induced by this mutation, or just occluded by a ceiling effect due to the synapses already being strengthened. Occlusion may also occur in the mixed cultures when Rab3A is missing from neurons but not astrocytes. The authors do appropriately discuss these options. The authors have solid data showing that Rab3A is unlikely to be active in astrocytes, Finally, they attempt to study the linkage between changes in synaptic strength and AMPA receptor trafficking during HSP, and conclude that trafficking may not be solely responsible for the changes in synaptic strength during HSP.
Strengths:
This work adds another player into the mechanisms underlying an important form of synaptic plasticity. The plasticity is likely only reduced, suggesting Rab3A is only partially required and perhaps multiple mechanisms contribute. The authors speculate about some possible novel mechanisms, including whether Rab3A is active pre-synaptically to regulate quantal amplitude.
As Rab3A is primarily known as a pre-synaptic molecule, this possibility is intriguing. However, it is based on the partial dissociation of AMPAR trafficking and synaptic response and lacks strong support. On average, they saw a similar magnitude of change in mEPSC amplitude and GluA2 cluster area and integral, but the GluA2 data was not significant due to higher variability. It is difficult to determine if this is due to biology or methodology - the imaging method involves assessing puncta pairs (GluA2/VGlut1) clearly associated with a MAP2 labeled dendrite. This is a small subset of synapses, with usually less than 20 synapses per neuron analyzed, which would be expected to be more variable than mEPSC recordings averaged across several hundred events. However, when they reduce the mEPSC number of events to similar numbers as the imaging, the mESPC amplitudes are still less variable than the imaging data. The reason for this remains unclear. The pool of sampled synapses is still different between the methods and recent data has shown that synapses have variable responses during HSP. Further, there could be variability in the subunit composition of newly inserted AMPARs, and only assessing GluA2 could mask this (see below). It is intriguing that pre-synaptic changes might contribute to HSP, especially given the likely localization of Rab3A. But it remains difficult to distinguish if the apparent difference in imaging and electrophysiology is a methodological issue rather than a biological one. Stronger data, especially positive data on changes in release, will be necessary to conclude that pre-synaptic factors are required for HSP, beyond the established changes in post-synaptic receptor trafficking.
Regarding the concern that the lack of increase in receptors is due to a technical issue, please see General Response to Reviewers, above. We have also softened our conclusions throughout, acknowledging we cannot rule out a technical issue.
Other questions arise from the NASPM experiments, used to justify looking at GluA2 (and not GluA1) in the immunostaining. First, there is a strong frequency effect that is unclear in origin. One would expect NASPM to merely block some fraction of the post-synaptic current, and not affect pre-synaptic release or block whole synapses. But the change in frequency seems to argue (as the authors do) that some synapses only have CP-AMPARs, while the rest of the synapses have few or none. Another possibility is that there are pre-synaptic NASPM-sensitive receptors that influence release probability. Further, the amplitude data show a strong trend towards smaller amplitude following NASPM treatment (Fig 3B). The p value for both control and TTX neurons was 0.08 - it is very difficult to argue that there is no effect. The decrease on average is larger in the TTX neurons, and some cells show a strong effect. It is possible there is some heterogeneity between neurons on whether GluA1/A2 heteromers or GluA1 homomers are added during HSP. This would impact the conclusions about the GluA2 imaging as compared to the mEPSC amplitude data.
The key finding in Figure 3 is that NASPM did not eliminate the statistically significant increase in mEPSC amplitude after TTX treatment (Fig 3A). Whether or not NASPM sensitive receptors contribute to mESPC amplitude is a separate question (Fig 3B). We are open to the possibility that NASPM reduces mEPSC amplitude in both control and TTX treated cells (p = 0.08 for both), but that does not change our conclusion that NASPM has no effect on the TTX-induced increase in mEPSC amplitude. The mechanism underlying the decrease in mEPSC frequency following NASPM is interesting, but does not alter our conclusions regarding the role of Rab3A in homeostatic synaptic plasticity of mEPSC amplitude. In addition, the Reviewer does not acknowledge the Supplemental Figure #1, which shows a similar lack of correspondence between homeostatic increases in mEPSC amplitude and GluA1 receptors in two cultures where matched data were obtained. Therefore, we do not think our lack of a robust increase in receptors can be explained by our failing to look at the relevant receptor.
To understand the role of Rab3A in HSP will require addressing two main issues:
(1) Is Rab3A acting pre-synaptically, post-synaptically or both? The authors provide good evidence that Rab3A is acting within neurons and not astrocytes. But where it is acting (pre or post) would aid substantially in understanding its role. The general view in the field has been that HSP is regulated post-synaptically via regulation of AMPAR trafficking, and considerable evidence supports this view. More concrete support for the authors' suggestion of a pre-synaptic site of control would be helpful.
We agree that definitive evidence for a presynaptic role of Rab3A in homeostatic plasticity of mEPSC amplitudes in mouse cortical cultures requires demonstrating that loss of Rab3A in postsynaptic neurons does not disrupt the plasticity, whereas loss in presynaptic neurons does. Without these data, we can only speculate that the Rab3A-dependence of homeostatic plasticity of quantal size in cortical neurons may be similar to that of the neuromuscular junction, where it cannot be receptors. We have added to the Discussion that the mechanism of Rab3A regulation of homeostatic plasticity of quantal amplitude could different between cortical neurons and the neuromuscular junction (lines 448-450 in markup,). Establishing a way to co-culture Rab3A-/- and Rab3A+/+ neurons in ratios that would allow us to record from a Rab3A-/- neuron that has mainly Rab3A+/+ inputs (or vice versa) is not impossible, but requires either transfection or transgenic expression with markers that identify the relevant genotype, and will be the subject of future experiments.
(2): Rab3A is also found at inhibitory synapses. It would be very informative to know if HSP at inhibitory synapses is similarly affected. This is particularly relevant as at inhibitory synapses, one expects a removal of GABARs or a decrease in GABA release (ie the opposite of whatever is happening at excitatory synapses). If both processes are regulated by Rab3A, this might suggest a role for this protein more upstream in the signaling; an effect only at excitatory synapses would argue for a more specific role just at those synapses.
We agree with the Reviewer, that it is important to determine the generality of Rab3A function in homeostatic plasticity. Establishing the homeostatic effect on mIPSCs and then examining them in Rab3A-/- cultures is a large undertaking and will be the subject of future experiments.
Recommendations for the authors:
Reviewer #1 (Recommendations for the authors):
Minor (remaining) points:
(1) The figure referenced in the first response to the reviewers (Figure 5G) does not exist.
We meant Figure 5F, which has been corrected in the current response.
(2) I recommend showing the data without binning (despite some overlap).
The box plot in Origin will not allow not binning, but we can make the bin size so small that for all intents and purposes, there is close to 1 sample in each bin. When we do this, the majority of data are overlapped in a straight vertical line. Previously described concerns were regarding the gaps in the data, but it should be noted that these are cell means and we are not depicting the distributions of mEPSC amplitudes within a recording or across multiple recordings.
(3) Please auto-scale all axes from 0 (e.g., Fig 1E, F).
We have rescaled all mEPSC amplitude axes in box plots to go from 0 (Figures 1, 2 and 6).
(4) Typo in Figure legend 3: "NASPM (20 um)" => uM
Fixed.
Reviewer #2 (Recommendations for the authors):
(1) Line 140, frequencies are reported in Hz while other places are in sec-1, while these are essentially the same, they should be kept consistent in writing.
All mEPSC frequencies have been changed to sec<sup>-1</sup>, except we have left “Hz” for repetitive stimulation and filtering.
(2) Paragraph starting from line 163 (as well as other places where multiple groups are compared, such as the occlusion discussion), the authors assessed whether there was a change in baseline between WT and mutant group by doing pairwise tests, this is not the right test. A two-way ANOVA, or at least a multivariant test would be more appropriate.
We have performed a two-way ANOVA, with genotype as one factor, and treatment as the other factor. The p values in Figures 1 and 2 have been revised to reflect p values from the post-hoc Tukey test on the specific interactions (for each particular genotype, TTX vs CON effects). The difference in the two WT strains, untreated, was not significant in the Post-Hoc Tukey test, and we have revised the text. The difference between the untreated WT from the Rab3A+/Ebd colony and the untreated Rab3AEbd/Ebd mutant was still significant in the Post-Hoc Tukey test, and this has replaced the Kruskal-Wallis test. The two-way ANOVA was also applied to the neuron-glia experiments and p values in Figure 6 adjusted accordingly.
(3) Relevant to the second point under minor concerns, I suggest this sentence be removed, as reducing variability and avoiding inhibitory projects are reasons good enough to restrict the analysis to the apical dendrites.
We have revised the description of the Turrigiano et al., 1998 finding from their Figure 3 and feel it still strengthens the justification for choosing to analyze only synapses on the apical dendrite.
Reviewer #3 (Recommendations for the authors):
Minor points:
The comments on lines 256-7 could seem misleading - the NASPM results wouldn't rule out contribution of those other subunits, only non-GluA2 containing combinations of those subunits. I would suggest revising this statement. Also, NASPM does likely have an effect, just not one that changes much with TTX treatment.
At new line 213 (markup) we have added the modifier “homomeric” to clarify our point that the lack of NASPM effect on the increase in mEPSC amplitude after TTX indicates that the increase is not due to more homomeric Ca<sup>2+</sup>-permeable receptors. We have always stated that NASPM reduces mEPSC amplitude, but it is in both control and treated cultures.
Strong conclusions based on a single culture (lines 314-5) seem unwarranted.
We have softened this statement with a “suggesting that” substituted for the previous “Therefore,” but stand by our point that the mEPSC amplitude data support a homeostatic effect of TTX in Culture #3, so the lack of increase in GluA2 cluster size needs an explanation other than variability in the homeostatic effect itself.
Saying (line 554) something is 'the only remaining possibility' also seems unwarranted.
We have softened this statement to read, “A remaining possibility…”.
Beattie EC, Stellwagen D, Morishita W, Bresnahan JC, Ha BK, Von Zastrow M, Beattie MS, Malenka RC (2002) Control of synaptic strength by glial TNFalpha. Science 295:2282-2285.
Mansour M, Nagarajan N, Nehring RB, Clements JD, Rosenmund C (2001) Heteromeric AMPA receptors assemble with a preferred subunit stoichiometry and spatial arrangement. Neuron 32:841-853. Stellwagen D, Malenka RC (2006) Synaptic scaling mediated by glial TNF-alpha. Nature 440:1054-1059.
Stellwagen D, Beattie EC, Seo JY, Malenka RC (2005) Differential regulation of AMPA receptor and GABA receptor trafficking by tumor necrosis factor-alpha. J Neurosci 25:3219-3228.
Swanson GT, Kamboj SK, Cull-Candy SG (1997) Single-channel properties of recombinant AMPA receptors depend on RNA editing, splice variation, and subunit composition. J Neurosci 17:5869.
Turrigiano GG, Leslie KR, Desai NS, Rutherford LC, Nelson SB (1998) Activity-dependent scaling of quantal amplitude in neocortical neurons. Nature 391:892-896.
Wang X, Wang Q, Yang S, Bucan M, Rich MM, Engisch KL (2011) Impaired activity-dependent plasticity of quantal amplitude at the neuromuscular junction of Rab3A deletion and Rab3A earlybird mutant mice. J Neurosci 31:3580-3588.
Wang X, Li Y, Engisch KL, Nakanishi ST, Dodson SE, Miller GW, Cope TC, Pinter MJ, Rich MM (2005) Activity-dependent presynaptic regulation of quantal size at the mammalian neuromuscular junction in vivo. J Neurosci 25:343-351.
🎯 **Objetivo da atividade ** Compreender os fundamentos e a lógica da Análise em Componentes Principais (ACP), reconhecendo o seu valor para a redução de dimensionalidade e para a análise de dados em contextos de gestão.
✅ Instruções
dim(V)=dim(W)+dim(W⊥).
Denne formel gælder også hvis \(\textup{dim}V=\infty\) eller \(\textup{dim} V=0\). (Har dog ikke fundet bevis for\((W^\perp)^\perp=W\) når \(\textup{dim}V=\infty\))
Bevis:
Hvis \(\textup{dim} V=0\), så må \(W=W^\perp=V={0}\) og derfor: \(\textup{dim}V=0=0+0=\textup{dim}W+\textup{dim}W^\perp\).
Hvis \(\textup{dim}V=\infty\), bemærk så at udsagnet holder trivielt hvis \(\textup{dim}W=\infty\) eller \(\textup{dim}W^\perp=\infty\). Så hvis \(\textup{dim}W=\infty\) er vi færdige. Antag derfor at \(\textup{dim}W<\infty\). Antag for modstrid at \(\textup{dim}W^\perp<\infty\). Så gælder per TØ-opgave 5 i ugen 10/3-16/3 at \(W+W^\perp={w+u\mid w\in W,u\in W^\perp}\) er et underrrum i \(V\), og per korollar 10.18 er \(V\subseteq W+W^\perp\). Så \(W+W^\perp =V\). Herudover per samme TØ-opgave gælder der:
\(\infty>\textup{dim}W+\textup{dim}W^\perp=\textup{dim}(W+W^\perp)+\textup{dim}(W\cap W^\perp)=\textup{dim}V+\textup{dim}{0}=\infty+0=\infty\)
Og dette er en modstrid. Dermed må \(\textup{dim}W^\perp=\infty\), og så holder udsagnet også.
Q.E.D.
ucleotide depletion, and telomere shortening. A second mechanism of p53 induction is activated by oncogenes such as Myc, which promote aberrant G1/S transition. This pathway is regulated by a second product of the Ink4a locus, p14ARF (p19 in mice), which is encoded b

gression are expressed. If the cell determines that it is unready to move ahead with DNA replication, a number of inhibitors are capable of blocking the action of the CDKs, including p21Cip2/Waf1, p16Ink4a, a

Reviewer #1 (Public review):
Summary:
The authors confirmed earlier findings that AVP influences α and β cells differently, depending on glucose concentrations. At substimulatory glucose levels, AVP combined with forskolin - an activator of cAMP -did not significantly stimulate β cells, although it did activate α cells. Once glucose was raised to stimulatory levels, β cells became active, and α cell activity declined, indicating glucose's suppressive effect on α cells and permissive effect on β cells. Under physiological glucose levels (8-9 mM), forskolin enhanced β-cell calcium oscillations, and AVP further modulated this activity. However, AVP's effect on β cells was variable across islets and did not significantly alter AUC measurements (a combined indicator of oscillation frequency and duration). In α cells, forskolin and AVP led to increased activity even at high glucose levels, suggesting that α cells remain responsive despite expected suppression by insulin and glucose.
Experiments with physiological concentrations of epinephrine suggest that AVP does not operate via Gs-coupled V2 receptors in β cells, as AVP could not counteract epinephrine's inhibitory effects. Instead, epinephrine reduced β cell activity while increasing α cell activity through different G-protein-coupled mechanisms. These results emphasize that AVP can potentiate α-cell activation and has a nuanced, context-dependent effect on β cells.
The most robust activation of both α and β cells by AVP occurred within its physiological osmo-regulatory range (~10-100 pM), confirming that AVP exerts bell-shaped concentration-dependent effects on β cells. At low concentrations, AVP increased β cell calcium oscillation frequency and reduced "halfwidths"; high concentrations eventually suppressed β cell activity, mimicking the muscarinic signaling. In α cells, higher AVP concentrations were required for peak activation, which was not blunted by receptor inactivation within physiological ranges.
Attempting to further dissect the role of specific AVP receptors, the authors designed and tested peptide ligands selective for V1b receptors. These included a selective V1b agonist; a V1b agonist with antagonist properties at V1a and oxytocin receptors; and a selective V1a antagonist. In pancreatic slices, these peptides seem to replicate AVP's effects on Ca²⁺ signaling, although responses were highly variable, with some islets showing increased activity and others no change or suppression. The variability was partly attributed to islet-specific baseline activity, and the authors conclude that AVP and V1b receptor agonists can modulate β cell activity in a state-dependent manner, stimulating insulin secretion in quiescent cells and inhibiting it in already active cells.
Strengths:
Overall, the study is technically advanced and provides useful pharmacological tools. However, the conclusions are limited by a lack of direct mechanistic and functional data. Addressing these gaps through a combination of signaling pathway interrogation, functional hormone output, genetic validation, and receptor localization would strengthen the conclusions and reduce the current (interpretive) ambiguity.
Weaknesses:
(1) The study is entirely based on pharmacological tools. Without genetic models, off-target effects or incomplete specificity of the peptides cannot be fully ruled out.
(2) Despite multiple claims about β cell activation or inhibition, the functional output - insulin secretion - is weakly assessed, and only in limited conditions. This aspect makes it very hard to correlate calcium dynamics with physiological outcomes.
(3) Insulin and glucagon secretion assays should be provided; the authors should measure hormone release in parallel with Ca2+ imaging, using perifusion assays, especially during AVP ramp and peptide ligand applications.
Additionally, there is no standardization of the metabolic state of islets. The authors should consider measuring islet NAD(P)H autofluorescence or mitochondrial potential (e.g., using TMRE) to control for metabolic variability that may affect responsiveness.
(4) There is a high degree of variability in response to AVP and V1b agonists across islets (activation, no effect, inhibition). Surprisingly, the authors do not fully explore the cause of this heterogeneity (whether it is due to receptor expression differences, metabolic state, experimental variability, or other conditions).
(5) There is no validation of V1b receptor expression at the protein or mRNA level in α or β cells using in situ hybridization, immunohistochemistry, or spatial transcriptomics.
(6) AVP effects are described in terms of permissive or antagonistic effects on cAMP (especially in relation to epinephrine), but direct measurements of cAMP in α and β cells are not shown, weakening these conclusions. The authors should use Epac-based cAMP FRET sensors in α and β cells to monitor the interaction between AVP, forskolin, and epinephrine more conclusively.
(7) Single-islet transcriptomics or proteomics (also to clarify variability) should be provided to analyze receptor expression variability across islets to correlate with response phenotypes (activation vs inhibition). Alternatively, the authors could perform calcium imaging with simultaneous insulin granule tracking or ATP levels to assess islet functional states.
(8) While the study implies AVP acts through V1b receptors on β cells, the signaling downstream (e.g., PLC activation, IP3R isoforms involved) is simply inferred but not directly shown.
(9) The interpretation that IP3R inactivation (mentioned in the title!) underlies the bell-shaped AVP effect is just hypothetical, without direct measurements. Assays in β (and/or α)-cell-specific V1b KO mice and IP3R KO mice must be provided to support these speculations.
Reviewer #2 (Public review):
Summary:
In this paper, Drs. Kercmar, Murko, and Bombek make a series of observations related to the role of AVP in pancreatic islets. They use the pancreatic slice preparation that their group is well known for. The observations on the slide physiology are technically impressive. However, I am not convinced by the conclusions of this manuscript for a number of reasons. At the core of my concern is perhaps that this manuscript appears to be motivated to resolve 'controversies' surrounding the actions of AVP on insulin and glucagon secretion. This manuscript adds more observations, but these do not move the field forward in improving or solidifying our mechanistic understanding of AVP actions on islets. A major claim in this manuscript is the beta cell expression of the V1b Receptor for AVP, but the evidence presented in this paper falls short of supporting this claim. Observations on the activation of calcium in alpha cells via V1b receptor align with prior observations of this effect.
I have focused my main concerns below. I hope the authors will consider these suggestions carefully - please be assured that they were made with the intent to support the authors and increase the impact of this work.
Strengths:
The main strength of this paper is the technical sophistication of the approach and the analysis and representation of the calcium traces from alpha and beta cells.
Weaknesses:
(1) The introduction is long and summarizes a substantive body of literature on AVP actions on insulin secretion in vivo. There are a number of possible explanations for these observations that do not directly target islet cells. If the goal is to resolve the mechanistic basis of AVP action on alpha and beta cells, the more limited number of papers that describe direct islet effects is more helpful. There are excellent data that indicate that the actions of AVP are mediated via V1bR on alpha cells and that V1bR is a) not expressed by beta cells and b) does not activate beta cell calcium at all at 10 nM - which is the same concentration used in this paper (Figure 4G) for peak alpha cell Ca2+ activation (see https://doi.org/10.1016/j.cmet.2017.03.017; cited as ref 30 in the current manuscript).
(2) We know from bulk RNAseq data on purified alpha, beta, and delta cells from both the Huising and Gribble groups that there is no expression of V2a. I will point you to the data from the Huising lab website published almost a decade ago (http://dx.doi.org/10.1016/j.molmet.2016.04.007) - which is publicly available and can be used to generate figures (https://huisinglab.com/data-ghrelin-ucsc/index.html). They indicate the absence of expression of not only AVP2 receptors anywhere in the islet, but also the lack of expression of V1bra, V1brb, and Oxtr in beta cells. Instead of the detailed list of expression of these 4 receptors elsewhere in the body, it would be more directly relevant to set up their pancreatic slice experiments to summarize the known expression in pancreatic islets that is publicly available. It would also have helped ground the efforts that involved the generation of the V1aR agonist and V2R antagonist, which confirm these known AVP/OXT receptor expression patterns.
(3) Importantly, the lack of V1br from beta cells does not invalidate observations that AVP affects calcium in beta cells, but it does indicate that these effects are mediated a) indirectly, downstream of alpha cell V1br or b) via an unknown off-target mechanism (less likely). The different peak efficacies in Figure 4G would also suggest that they are not mediated by the same receptor.
(4) The rationale for the use of forskolin across almost all traces is unclear. It is motivated by a desire to 'study the AVP dependence of both alpha and beta cells at the same time'. As best as I can determine, the design choice to conduct all studies under sustained forskolin stimulation is related to the permissive actions of AVP on hormone secretion in response to cAMP-generating stimuli. The permissive actions by AVP that are cited are on hormone secretion, which in many cell types requires activation of both calcium and cAMP signaling. Whether the activation of V1br and subsequent calcium response is permitted by cAMP is unclear. I believe the argument the authors are making here is that the activation of beta cell calcium by AVP is permitted by forskolin. i.e., the cAMP stimulated by it in beta cells. However, the design does not account for the elevation of cAMP in alpha cells and subsequent release of glucagon, particularly upon co-stimulation with AVP, which permits glucagon release by activating a calcium response in alpha cells. This glucagon could then activate beta cells. If resolving the mechanism of action is the goal, often less is more. The activation of Gaq-mediated calcium is not cAMP dependent (although the downstream hormone secretion clearly often is). As was shown, AVP does not activate calcium in beta cells in the absence of cAMP. The experiments in Figures 1, 2, and 4 should have been completed in the absence of cAMP first.
(5) It is unexpected that epinephrine in Figure 2 does not activate the alpha cell calcium? A recent paper from the same group (Sluga et al) shows robust calcium activation in alpha cells in a similar prep by 1 nM epinephrine, which is similar to the dose used here.
(6) Figure 8 suggests a pharmacological activation of beta cell V1bR in the low pM range. How do the authors reconcile this comparison with the apparent absence of an effect of AVP stimulation at low pM to low nM doses in beta cells (Figure 4A)? I note that there are changes over time with sustained beta cell stimulation with 8 mM glucose, but these changes are relatively subtle, gradual, and quite likely represent the progression of calcium behaviors that would have occurred under sustained glucose, irrespective of these very low AVP concentrations. I will note that the Kd of the V1bR for AVP is around 1 nM, with tracer displacement starting around 100 pM according to the data in figure 5B, which is hard to reconcile with changes in beta cell calcium by AVP doses that start 10-100-fold lower than this dose at 1 and 10 pM (Figure 8).
Reviewer #2 (Public review):
The manuscript entitled "Structure of an oxygen-induced tubular nanocompartment in Pyrococcus furiosus" by Wenfei Song et al. employs whole-cell mass spectrometry and cryo-EM (including tomography, helical reconstruction, and single-particle analysis) to investigate the structure and function of the oxidoreductase Rubrerythrin (Rbr) from Pyrococcus furiosus. The study reports that under oxidative stress, Rbr forms a tubular structure, in contrast to its behaviour under anaerobic conditions. Authors characterized oxidoreductase Rubrerythrin (Rbr) from Pyrococcus furiosus under anaerobic conditions and formed a tubular structure when induced with oxidative stress. This study is well-designed. However, I have several questions related to the experimental design and the results obtained from those experiments, which are listed below.
(1) The authors have mentioned that "Under aerobic conditions, Rbr levels are 3 to 13 times higher compared to anaerobic conditions (Figures 1a-d)." Also, they performed whole-cell mass spec to measure the overexpression of the Rbr enzyme under anaerobic conditions. Thus, from the above statement, I consider the authors' claim that P. furiosus cells were cultured under anaerobic conditions and then exposed to oxidative stress. While cell growth under anaerobic conditions appears perfectly fine, the authors conducted the rest of the experiment under aerobic conditions during mass spectrometry and cryo-EM sample preparation. As a baseline, the author first grew the cells in their preferred anaerobic environment and also imaged the same cells that were exposed to air (aerobic) after anaerobic growth. The cell growth in anaerobic conditions is perfectly fine. But how did authors make sure that during anaerobic conditions, the Rbr enzyme is not expressed or not formed? As a control experiment, authors should demonstrate that during mass spec and cryo-EM sample preparations, cells are not exposed to air or maintained in an anaerobic environment. From anaerobic conditions, whenever cells were selected for spec and cryo-EM, cells were exposed to O2, and definitely controlled cells were not in anaerobic conditions anymore.
The authors collected P. furiosus wild-type or Rbr knockout cells in an anaerobic hood, but after that, they centrifuged the cells and plunged them using a Vitrobot. Are the instrument, centrifuge, and Vitrobot kept in an anaerobic environment? Recently, a few studies (anaerobic plunge-freezing in cryo-electron microscopy, Cook et al. (2024), Hands-Portman and Bakker (2022) DOI: 10.1039/D2FD00060A ) have mentioned the anaerobic plunge freeze setup for protein sample or cell freezing. I guess the authors did not use that setup. In these circumstances, the cell is already exposed to O2 during centrifugation and Vitrobot freezing. How were the control experiments properly performed in anaerobic conditions? A similar argument is true for Lamella grid preparation, where the enzyme was already exposed to O2, and single-particle grid preparation, where the purified enzyme is already exposed to O2. How were the control experiments properly performed in anaerobic conditions?
(2) It is important to provide evidence that the overexpressed protein is actually in an anaerobic condition and is later induced with more O2. Also, authors should confirm biochemically that the overexpressed protein in their desired protein "oxidoreductase Rubrerythrin (Rbr)". No biochemical data were provided in this manuscript. During single-particle analysis, the authors had to purify the protein sample and confirm that these were their desired protein samples. No biochemical or biophysical experiments were performed to confirm that the overexpressed protein is the desired protein.
(3) Figure 3, the atomic model looks different in all four tetramers. However, I have fitted the atomic model into the cryo-EM map, which looks reasonable. However, it will be easier for the reader to evaluate the model if the authors show different orientations of the atomic model, as well as if the authors could show that the atomic model fits the cryo-EM map.
(4) How did the authors select initial particle sets like 24 lakhs when forming helices and not forming isolated particles?
(5) The authors proposed a model for electron transfer upon oxidative stress. However, the data is not convincing that VLP is surrounded by Rbr and forms a tube-like structure. Generally, VLP is a sphere-like structure, and Rbr can form a tube-like structure when it interacts with spherical VLP. Rbr will surround VLP, and it will form a Rbr-decorated sphere-like structure.
(6) It will also be important to comment on the diameter of Oxidative stress-induced tubules (OSITs) and 3D reconstruction and/or helical reconstruction of purified protein samples. The spherical cyan densities within the tube are not very clear. If VLP is surrounded by Rbr (Figure 4), extra Rbr densities will be observed on VLP in the tomogram (in Figure 1). However, in the tomogram, VLP is inside Oxidative stress-induced tubules (OSITs). Figure 1 is a contradicting Figure 4. The authors should explain it properly.
(7) The authors performed helical reconstruction. Where is the Layer line calculation in helical reconstruction, and how do authors identify helical parameters for reconstruction?
(8) The authors used an extremely confusing methodology, which was very difficult to follow. The authors performed tomography, helical reconstruction, and single-particle analysis. Why did the authors need 3 different image processing methods to resolve structures that are not clear to me? The authors should also show the proper fitting between the map and the model. In Supplemental Figure 6c, the overall fitting of the subdomain looks ok. However, many peptide chains and side chains are not fitted properly in the EM density map. It will be helpful to show proper side chain fitting. In Supplementary Fig. 6a, the authors binned the data (Bin 8 or Bin 2) but did not mention when they unbinned the data for data processing. Also, the authors implemented C2 symmetry during local refinement. Why do authors suddenly use C2 symmetry expansion?
Minor Comments:
(1) The authors should properly show a schematic diagram of the enzyme subdomains. It will help to understand interactions or tetrameric assembly.
(2) The introduction is poorly written. It will really be helpful for the reader if the authors provide a proper introduction.
(3) The atomic model did not fit into the cryo-EM, so it was hard to determine the overall fitting.
(4) 17.1A pixel size? It's surprising.
(5) It will be better to calculate local resolution and show the map's angular distribution. It is obvious that resolution at the peripheral region will be poorer than core region. Therefore, it will be better to calculate local resolution. Additionally, authors should show the map to model fitting.
Reviewer #3 (Public review):
Summary:
The manuscript authored by Song et al explores the oxidative stress response of Rubrerythrin in Pyrococcus furiosus and the formation of unique tubules that also encapsulate Encapsulin VLPs. This is an excellent study employing diverse methods to comprehensively study the formation of these assemblies under oxidative stress and lays the foundation of understanding oxidative stress through the formation of tubules among redox-sensing proteins like Rubrerythrin. The authors decipher the molecular structure of the tubules and also present a high-resolution reconstruction of the rubrerythin unit that forms the OSITs.
Strengths:
The study is done thoroughly by employing methods like cryoET, single particle cryoEM, mass spectrometry, and expression analyses of knockout strains to delve into an important mechanism to counter oxidative stress. The authors perform comprehensive analyses, and this study represents a vital contribution to understanding how anaerobic organisms can respond to oxidative stress.
Weaknesses:
Not all encapsulin particles seem to be inside the OSITs. Do the authors have any insights into how the tubules sequester these viral particles? Do the VLPs have a role in nucleating the OSIT assembly, and are there interactions between VLP and OSIT surfaces? These could be points that can be discussed in greater detail by the authors.
Can the authors get a subtomogram averaging done for the encapsulin VLPs? A higher resolution reconstruction may provide potential interaction details with the OSITs, if there are any.
The role of the dense granules observed in the rubrerythrin deletion strain is not very well discussed. Is there a way these granules counter oxidative stress? The EDX scanning seems to show a Phosphate increase similar to Ca and Mg. Are these aggregates therefore likely to be calcium and Mg phosphate aggregates? This section of the paper seems incompletely analysed.
The authors should provide density and coordination distances around the diiron ions and provide a comparison with available crystal structures and highlight differences, if any, in Figure 3. Local resolution for the high-res map may be provided for Supplementary Figure 6.
Overall, this is a well-performed study with clear conclusions. The discussion points need to be improved further.
Df(3L)BSC561
DOI: 10.1093/nsr/nwaf095
Resource: Bloomington Drosophila Stock Center (RRID:SCR_006457)
Curator: @maulamb
SciCrunch record: RRID:SCR_006457
RRID: AB_2336171
DOI: 10.1016/j.isci.2025.112767
Resource: (Vector Laboratories Cat# BA-9200, RRID:AB_2336171)
Curator: @dhovakimyan1
SciCrunch record: RRID:AB_2336171
RRID: AB_2313606
DOI: 10.1016/j.isci.2025.112767
Resource: (Vector Laboratories Cat# BA-1000, RRID:AB_2313606)
Curator: @dhovakimyan1
SciCrunch record: RRID:AB_2313606
#11150
DOI: 10.1007/s00418-025-02390-5
Resource: RRID:Addgene_11150
Curator: @dhovakimyan1
SciCrunch record: RRID:Addgene_11150
RRID:AB_2313581
DOI: 10.1093/cercor/bhaf090
Resource: (Vector Laboratories Cat# BA-2000, RRID:AB_2313581)
Curator: @scibot
SciCrunch record: RRID:AB_2313581
RRID:AB_314276
DOI: 10.1016/j.xcrm.2025.102145
Resource: (BioLegend Cat# 302606, RRID:AB_314276)
Curator: @scibot
SciCrunch record: RRID:AB_314276
RRID:SCR_002798
DOI: 10.1016/j.neo.2025.101179
Resource: GraphPad Prism (RRID:SCR_002798)
Curator: @scibot
SciCrunch record: RRID:SCR_002798
RRID:IMSR_JAX:002650
DOI: 10.1016/j.molmet.2025.102171
Resource: (IMSR Cat# JAX_002650,RRID:IMSR_JAX:002650)
Curator: @scibot
SciCrunch record: RRID:IMSR_JAX:002650
MGI:5003554
DOI: 10.1016/j.molcel.2025.05.005
Resource: None
Curator: @scibot
SciCrunch record: RRID:MGI:5003554
RRID:AB_10717953
DOI: 10.1016/j.isci.2025.112639
Resource: (Thermo Fisher Scientific Cat# 46-0391-82, RRID:AB_10717953)
Curator: @scibot
SciCrunch record: RRID:AB_10717953
RRID:SCR_003070
DOI: 10.1016/j.devcel.2025.04.026
Resource: ImageJ (RRID:SCR_003070)
Curator: @scibot
SciCrunch record: RRID:SCR_003070
RRID:AB_2313581
DOI: 10.1007/s12035-025-05048-4
Resource: (Vector Laboratories Cat# BA-2000, RRID:AB_2313581)
Curator: @scibot
SciCrunch record: RRID:AB_2313581
Ay, while you live, draw your neck out o' the collar.
Gregory mocks him, using draw to mean escaping hanging. Commentary: A dark joke implies Sampson’s bravado might get him killed.
Ay, the heads of the maids, or their maidenheads; take it in what sense thou wilt. 40
Sampson is saying that after fighting the Montague men, he will either cut off the heads of the women or take their virginity, he adds, this can be understood either way.
Screenwriters, Directors, Actors, and playwrights * Woody Allen Olympia SM3 * Julie Andrews IBM Selectric I * Paul Auster: Olympia SM9 * Ingrid Bergman: Smith-Corona Skyriter * Ray Bradbury: Underwood (No. 5?), Royal KMM * Marlon Brando: Royal Arrow or Aristocrat * Bertolt Brecht: Erika * Richard Brooks: Royal KMM, Royal Portable (30s or 40s) * Mikhail Bulgakov: Olympia 8 * George Burns: Royal HH * Stephen J. Cannell: IBM Selectric II or III * Johnny Carson: Royal KMM (or maybe a KMG), Olivetti Lettera 22 * Paddy Chayefsky: Underwood Standard Model 6, Royal HH, Olympia SG3 * Francis Ford Coppola: Olivetti Lettera 32 * Norman Corwin: Flattop Corona, Royal KMM * Noel Coward: Royal KH, Imperial Standard, Olivetti-Underwood Studio 44 * Michael Crichton: Olympia electric, IBM Selectric I * Bing Crosby: Corona 3, 1920s Royal portable * Bette Davis: Remington Noiseless portable * Joe Eszterhas: Olivetti Lettera 35 * Douglas Fairbanks: Underwood 5 * Federico Fellini: Olivetti Studio 44 * Jodie Foster: Olivetti Lettera 35 * Stephen Fry: Hermes 3000 * Greta Garbo: Olympia SM 7 * William Gibson: Hermes 2000 * William Goldman: Olympia SM9, Olympia SM9, Hermes Baby, Olympia Traveller<br /> Matt Groening: Hermes Rocket<br /> Oscar Hammerstein II: Royal portable * Tom Hanks: Smith-Corona Clipper, Hermes 2000, Hermes 3000<br /> Katherine Hepburn: Royal De Luxe * Alfred Hitchcock: '30s black Underwood Champion portable * John Hughes (director): Olympia SM3 * Eric Idle: Adler J3 * Elia Kazan: Royal KMG, Royal HH<br /> Buster Keaton: Blickensderfer no. 5 * Grace Kelly: Remington Super-Riter * Stanley Kubrick: IBM Model C, Adler Tippa S * Ring Lardner: L. C. Smith * Stan Lee: Remington noiseless portable, Olympia SG1 * Ernest Lehman: Royal Electress * David Letterman: Royal Empress * David Mamet: Smith-Corona portable, Olympia SM4, Olympia SM9, IBM Selectric * Terrence McNally: Olympia SG3 * Arthur Miller: Smith-Corona portable in the late '30s50s Smith; -Corona Silent Super; Royal KMG * Henry Miller: Underwood * F. W. Murnau: Remington portable no. 2 * David Niven: Royal Quiet DeLuxe, 1940s * Christopher Nolan: 1940s (?) Royal portable * Conan O'Brien: Royal 10 * Clifford Odets (1962): Royal Quiet DeLuxe, ca. 1957 * Louis Pollack (screenwriter): Royal desktop * Mario Puzo: Royal HH * Carl Reiner: Royal KMG, 1950s Smith-Corona Silent * Gene Roddenberry: IBM Selectric I, IBM Selectric II or III, Panasonic * Fred Rogers (Mr. Rogers): 1930s Royal portable (Model O?), Royal KMG * Rod Serling: Royal KMG * George Bernard Shaw: Bar-Lock; Remington portable no. 1; Smith Premier (Remington); Remington Noiseless Portable * Sam Shepard: '60s Hermes 3000, Olympia SM9 * Neil Simon: Olympia SM9 * Steven Spielberg: Smith-Corona Coronamatic 2200 * John Millington Synge: Blickensderfer #5 * Shirley Temple: white Student (Bing variant), white Underwood Champion portable, white Remington portable no. 5 or similar * David Thewlis: Olympia SM9 * James Thurber: Underwood no. 5 * Dalton Trumbo: Underwood, Royal KHM, IBM A or B * John Waters: ca. 1950 Underwood, IBM A or B * Orson Welles: 1926 woodgrain Underwood portable, ’30s Underwood Noiseless Portable, Smith-Corona (?) * Tennessee Williams: Remington portable no. 2, 1936 Corona Junior, mid-1940s Corona Sterling, Royal KMM, Hermes Baby, Olivetti Studio 44, Remington portable #5 flat top, Remington Standard M, Olympia SM8
Boost rigorous instruction, student feedback, and assessment in all content areas
Boost: Aumentar, mejorar o fortalecer. • Rigorous instruction: Se refiere a una enseñanza desafiante, profunda y de alta calidad que promueve el pensamiento crítico y la comprensión profunda, no solo la memorización. • Student feedback: Retroalimentación que reciben los estudiantes sobre su desempeño, útil para mejorar su aprendizaje. • Assessment: Evaluación del aprendizaje para medir el progreso y las necesidades. • All content areas: Todas las asignaturas o áreas del currículo (como matemáticas, lengua, ciencias, etc.).
3º Oferecer droga, eventualmente e sem objetivo de lucro, a pessoa de seu relacionamento, para juntos a consumirem:
Para a configuração do crime de oferecimento de droga para consumo conjunto, tipificado no art. 33, § 3º, da Lei n. 11.343/2006, é necessária a prática da conduta mediante o dolo “específico”. (CERTO)
Para a configuração do crime de oferecimento de droga para consumo conjunto, tipificado no art. 33, § 3º, da Lei n. 11.343/2006, é necessária a prática da conduta mediante o dolo “específico”. (CERTO)
declaração da falência
Vide que a declaração de falência sempre, de pleno direito, dissolve a sociedade. Observe que a declaração de falência do empresário sócio da sociedade gerará a dissolução parcial da sociedade, no que tange à quota do empresário.
divulgação
Ramo do Direito DIREITO CIVIL
Paz, Justiça e Instituições EficazesTema <br /> Vazamento de dados pessoais. Dados comuns e sensíveis. Dano moral presumido. Impossibilidade.
DESTAQUE - O vazamento de dados pessoais não gera dano moral presumido.
INFORMAÇÕES DO INTEIRO TEOR
Trata-se, na origem, de ação de indenização ajuizada por pessoa idosa contra concessionária de energia elétrica pleiteando indenização por danos morais decorrentes do vazamento e acesso, por terceiros, de dados pessoais.
O art. 5º, II, da Lei Geral de Proteção de Dados - LGPD dispõe de forma expressa quais dados podem ser considerados sensíveis e, devido a essa condição, exigir tratamento diferenciado, previsto em artigos específicos. Os dados de natureza comum, pessoais mas não íntimos, passíveis apenas de identificação da pessoa natural não podem ser classificados como sensíveis.
Os dados objeto da lide são aqueles que se fornece em qualquer cadastro, inclusive nos sites consultados no dia a dia, não sendo, portanto, acobertados por sigilo, e o conhecimento por terceiro em nada violaria o direito de personalidade da recorrida.
O vazamento de dados pessoais, a despeito de se tratar de falha indesejável no tratamento de dados de pessoa natural por pessoa jurídica, não tem o condão, por si só, de gerar dano moral indenizável. Ou seja, o dano moral não é presumido, sendo necessário que o titular dos dados comprove eventual dano decorrente da exposição dessas informações.
Diferente seria se, de fato, estivéssemos diante de vazamento de dados sensíveis, que dizem respeito à intimidade da pessoa natural.
Ramo do Direito DIREITO CIVIL
Paz, Justiça e Instituições EficazesTema <br /> Matéria jornalística. Críticas jornalísticas a magistrada. Autoridade pública. Direito de informação, expressão e liberdade de imprensa. Ausência de configuração de abuso no dever de informar. Interesse público. Dano moral. Afastamento. Prevalência da liberdade de informação e de crítica.
DESTAQUE - A divulgação de notícia ou crítica acerca de atos ou decisões do Poder Público, ou de comportamento de seus agentes, não configuram, a princípio, abuso no exercício da liberdade de imprensa, desde que não se refiram a núcleo essencial de intimidade e de vida privada da pessoa.
INFORMAÇÕES DO INTEIRO TEOR - Esta Corte Superior estabeleceu, para situações de conflito entre a liberdade de expressão e os direitos da personalidade, entre outros, os seguintes elementos de ponderação: "(I) o compromisso ético com a informação verossímil; (II) a preservação dos chamados direitos da personalidade, entre os quais incluem-se os direitos à honra, à imagem, à privacidade e à intimidade; e (III) a vedação de veiculação de crítica jornalística com intuito de difamar, injuriar ou caluniar a pessoa (animus injuriandi vel diffamandi)" (REsp 801.109/DF, Rel. Ministro Raul Araújo, Quarta Turma, DJe de 12/03/2013).
Em princípio, a publicação de matéria jornalística que narra fatos verídicos ou verossímeis não caracteriza hipótese de responsabilidade civil, ainda que apresentando opiniões severas, irônicas ou impiedosas, sobretudo quando se tratar de figura pública que exerça atividade tipicamente estatal, gerindo interesses da coletividade, e que se refira a fatos de interesse geral relacionados à atividade pública desenvolvida pela pessoa noticiada.
A liberdade de expressão, nessas hipóteses, é prevalente, atraindo verdadeira excludente anímica, a afastar o intuito doloso de ofender a honra da pessoa a que se refere a reportagem.
Contudo, a análise acerca da ocorrência de abuso no exercício da liberdade de expressão, a ensejar reparação por dano moral, deve ser feita em cada caso concreto, mormente quando a pessoa envolvida for investida de autoridade pública, pois, em tese, sopesados os valores em conflito, é recomendável que se dê primazia à liberdade de informação e de crítica, como decorrência da vida em um Estado Democrático.
Em observância à situação fática do processo em epígrafe, a reportagem baseou-se em relatos do superintendente da Polícia Civil do Estado, acerca da deflagração de operação que investigava pessoas envolvidas com o jogo do bicho em determinado Estado, citando a atuação da autora no exercício de seu cargo público (magistrada), tendo o Tribunal local consignado expressamente que "a intenção de narrar o ocorrido esteve presente durante toda a redação do texto".
Nesse prisma, tem-se que a matéria jornalística relacionou-se a fatos de interesse da coletividade, os quais dizem respeito diretamente com atos da magistrada enquanto autoridade pública.
Assim, verifica-se que, em que pese o tom ácido da referida reportagem, com o emprego de expressões como "aberração jurídica" e "descalabro", as críticas estão inseridas no âmbito da matéria jornalística de cunho informativo, baseada em levantamentos de fatos de interesse público, sem adentrar a intimidade e a vida privada da recorrida, o que significa que não extrapola claramente o direito de crítica, principalmente porque exercida em relação a caso que ostenta gravidade e ampla repercussão e interesse social.
Desse modo, quando não ficar caracterizado o abuso ofensivo na crítica exercida pela parte no exercício da liberdade de expressão jornalística, deve-se afastar o dever de indenização, por força da "imperiosa cláusula de modicidade" subjacente a que alude a eg. Suprema Corte no julgamento da ADPF 130/DF.
Ramo do Direito DIREITO CIVIL
Paz, Justiça e Instituições EficazesTema <br /> Direito ao esquecimento. Fatos verídicos. Exclusão de matéria jornalística. Impossibilidade.
DESTAQUE - O direito ao esquecimento não justifica a exclusão de matéria jornalística.
INFORMAÇÕES DO INTEIRO TEOR - O direito à liberdade de imprensa não é absoluto, devendo sempre ser alicerçado na ética e na boa-fé, sob pena de caracterizar-se abusivo. A esse respeito, a jurisprudência desta Corte Superior é consolidada no sentido de que a atividade da imprensa deve pautar-se em três pilares, a saber: (I) dever de veracidade, (II) dever de pertinência e (III) dever geral de cuidado. Ou seja, o exercício do direito à liberdade de imprensa será considerado legítimo se o conteúdo transmitido for verdadeiro, de interesse público e não violar os direitos da personalidade do indivíduo noticiado.
Se esses deveres não forem observados e disso resultar ofensa a direito da personalidade da pessoa objeto da comunicação, surgirá para o ofendido o direito de ser reparado.
No caso, consoante destacado pelo Tribunal de origem, não há dúvidas acerca da veracidade da informação divulgada. Ademais, tratando-se de fato relativo à esfera penal, revela-se presente o interesse público na notícia. Por sua vez, em que pese o recorrido tenha alegado que a notícia interferiu e interfere negativamente na sua vida profissional, não alegou que a sua divulgação pela imprensa teve o propósito de ofender a sua honra.
Desse modo, não houve abuso no exercício da liberdade de imprensa.
É preciso definir, então, se o tempo transcorrido desde a ocorrência do fato é capaz, por si só, de justificar a imposição do dever de proceder à exclusão da matéria jornalística.
Em algumas oportunidades, a Quarta e a Sexta Turmas desta Corte Superior se pronunciaram favoravelmente acerca da existência do direito ao esquecimento. Considerando os efeitos jurídicos da passagem do tempo, ponderou-se que o Direito estabiliza o passado e confere previsibilidade ao futuro por meio de diversos institutos (prescrição, decadência, perdão, anistia, irretroatividade da lei, respeito ao direito adquirido, ato jurídico perfeito e coisa julgada).
Ocorre que, em fevereiro deste ano, o Supremo Tribunal Federal definiu que o direito ao esquecimento é incompatível com a Constituição Federal (Tema 786). Assim, o direito ao esquecimento, porque incompatível com o ordenamento jurídico brasileiro, não é capaz de justificar a atribuição da obrigação de excluir a publicação relativa a fatos verídicos.
Recurso extraordinário com repercussão geral. Caso Aída Curi. Direito ao esquecimento. Incompatibilidade com a ordem constitucional. Recurso extraordinário não provido. 1. Recurso extraordinário interposto em face de acórdão por meio do qual a Décima Quinta Câmara Cível do Tribunal de Justiça do Estado do Rio de Janeiro negou provimento a apelação em ação indenizatória que objetivava a compensação pecuniária e a reparação material em razão do uso não autorizado da imagem da falecida irmã dos autores, Aída Curi, no programa Linha Direta: Justiça. 2. Os precedentes mais longínquos apontados no debate sobre o chamado direito ao esquecimento passaram ao largo do direito autônomo ao esmaecimento de fatos, dados ou notícias pela passagem do tempo, tendo os julgadores se valido essencialmente de institutos jurídicos hoje bastante consolidados. A utilização de expressões que remetem a alguma modalidade de direito a reclusão ou recolhimento, como droit a l’oubli ou right to be let alone, foi aplicada de forma discreta e muito pontual, com significativa menção, ademais, nas razões de decidir, a direitos da personalidade/privacidade. Já na contemporaneidade, campo mais fértil ao trato do tema pelo advento da sociedade digital, o nominado direito ao esquecimento adquiriu roupagem diversa, sobretudo após o julgamento do chamado Caso González pelo Tribunal de Justiça Europeia, associando-se o problema do esquecimento ao tratamento e à conservação de informações pessoais na internet. 3. Em que pese a existência de vertentes diversas que atribuem significados distintos à expressão direito ao esquecimento, é possível identificar elementos essenciais nas diversas invocações, a partir dos quais se torna possível nominar o direito ao esquecimento como a pretensão apta a impedir a divulgação, seja em plataformas tradicionais ou virtuais, de fatos ou dados verídicos e licitamente obtidos, mas que, em razão da passagem do tempo, teriam se tornado descontextualizados ou destituídos de interesse público relevante. 4. O ordenamento jurídico brasileiro possui expressas e pontuais previsões em que se admite, sob condições específicas, o decurso do tempo como razão para supressão de dados ou informações, em circunstâncias que não configuram, todavia, a pretensão ao direito ao esquecimento. Elas se relacionam com o efeito temporal, mas não consagram um direito a que os sujeitos não sejam confrontados quanto às informações do passado, de modo que eventuais notícias sobre esses sujeitos – publicadas ao tempo em que os dados e as informações estiveram acessíveis – não são alcançadas pelo efeito de ocultamento. Elas permanecem passíveis de circulação se os dados nelas contidos tiverem sido, a seu tempo, licitamente obtidos e tratados. Isso porque a passagem do tempo, por si só, não tem o condão de transmutar uma publicação ou um dado nela contido de lícito para ilícito. 5. A previsão ou aplicação do direito ao esquecimento afronta a liberdade de expressão. Um comando jurídico que eleja a passagem do tempo como restrição à divulgação de informação verdadeira, licitamente obtida e com adequado tratamento dos dados nela inseridos, precisa estar previsto em lei, de modo pontual, clarividente e sem anulação da liberdade de expressão. Ele não pode, ademais, ser fruto apenas de ponderação judicial. 6. O caso concreto se refere ao programa televisivo Linha Direta: Justiça, que, revisitando alguns crimes que abalaram o Brasil, apresentou, dentre alguns casos verídicos que envolviam vítimas de violência contra a mulher , objetos de farta documentação social e jornalística, o caso de Aida Curi, cujos irmãos são autores da ação que deu origem ao presente recurso. Não cabe a aplicação do direito ao esquecimento a esse caso, tendo em vista que a exibição do referido programa não incorreu em afronta ao nome, à imagem, à vida privada da vítima ou de seus familiares. Recurso extraordinário não provido. 7. Fixa-se a seguinte tese: “É incompatível com a Constituição a ideia de um direito ao esquecimento, assim entendido como o poder de obstar, em razão da passagem do tempo, a divulgação de fatos ou dados verídicos e licitamente obtidos e publicados em meios de comunicação social analógicos ou digitais. Eventuais excessos ou abusos no exercício da liberdade de expressão e de informação devem ser analisados caso a caso, a partir dos parâmetros constitucionais - especialmente os relativos à proteção da honra, da imagem, da privacidade e da personalidade em geral - e das expressas e específicas previsões legais nos âmbitos penal e cível”.
Tema - 786 - Aplicabilidade do direito ao esquecimento na esfera civil quando for invocado pela própria vítima ou pelos seus familiares.
Tese - É incompatível com a Constituição a ideia de um direito ao esquecimento, assim entendido como o poder de obstar, em razão da passagem do tempo, a divulgação de fatos ou dados verídicos e licitamente obtidos e publicados em meios de comunicação social analógicos ou digitais. Eventuais excessos ou abusos no exercício da liberdade de expressão e de informação devem ser analisados caso a caso, a partir dos parâmetros constitucionais - especialmente os relativos à proteção da honra, da imagem, da privacidade e da personalidade em geral - e as expressas e específicas previsões legais nos âmbitos penal e cível.
JURISPRUDÊNCIA EM TESES - EDIÇÃO 224
MARCO CIVIL DA INTERNET III - LEI N. 12.965/2014
Edição disponibilizada em: 27/10/2023
Edição atualizada em: 13/10/2023
4) O direito ao esquecimento, entendido como a possiblidade de obstar, em razão da passagem do tempo, a divulgação de fatos ou dados verídicos e licitamente obtidos e publicados em meios de comunicação social, analógicos ou digitais, não é aplicável ao ordenamento jurídico brasileiro.
5) A desindexação de conteúdos não se confunde com o direito ao esquecimento, pois não implica a exclusão de resultados, mas tão somente a desvinculação de determinados conteúdos obtidos por meio dos provedores de busca.
anfizem
dokularda hava ya da gaz birikmesi durumudur
Bifurkasyon
Bifurkasyon, bir şeyin iki dala ya da kola ayrıldığı yer anlamına gelir.
diverjan kökle
Diverjan kökler, bir dişin birden fazla kökünün birbirinden ayrılarak açılı şekilde konumlanması durumudur. Yani kökler birbirine paralel değil, "V" şeklinde ya da daha geniş açıyla ayrılmıştır.
trismusa
Trismus, çiğneme kaslarının spazmı veya kısıtlılığı nedeniyle ağız açıklığının azalması ya da tam açılamaması durumudur.
Tomber amoureux, c’est s’émerveiller et se laisser surprendre et sur les sites de rencontres, ça ne fonctionne pas.
Conclusion lapidaire qui néglige la diversité des expériences amoureuses issues de ces plateformes.
On finit par s’auto-dévaloriser, beaucoup plus qu’il ne faudrait et ça fait des petits ravages psychologiques.
Un des arguments phares : le décalage entre attentes et réalité. Toutefois, aucune donnée empirique ne vient soutenir cette affirmation.
L’exclu, c’est celui qui ne séduit pas dans la vraie vie et tombe dans le piège des sites de rencontres pensant qu’enfin, il pourra choper. Mais si on ne séduit pas dans la vraie vie, on ne séduit pas sur les sites de rencontre. Il y a 1000 et une façon de définir les critères de séduction : la beauté, l’humour, l’intelligence, un métier cool, du fric… Mais l’exclu n’a rien de tout ça. Et il se prend encore plus de râteaux que dans la vie.
Ici, l’auteur généralise un profil type en se basant sur une figure caricaturale. Cette typologie souffre d’un manque de nuance.
Les sites de rencontres, Stéphane Rose les a pratiqués durant des années. Pour lui, ils incarnent la misère sexuelle.
Cette entrée en matière donne le ton : une posture critique et désabusée qui imprègne tout l’article.
Reviewer #3 (Public review):
While the AAV capsid has long been the target of protein engineering, its Rep proteins have been comparatively less studied. Since Rep plays a variety of roles for genome replication and virion packaging, gaining a deeper mechanistic understanding of their function and/or engineering versions that enable higher packaging productivity would be of interest to the field. This study generates a library of non-synonymous mutations in AAV2 rep (intended to cover all 19 aa changes at all positions, and coming close), packaged an AAV with AAV9 capsid, and sequenced the results to assess which amino acid changes resulted in an enrichment/depletion of genomes containing that variant rep. They found that proline substitutions are disruptive, well known from protein mutagenesis studies. The most significant enrichment sfound, however, were a set of synonymous mutations (unintended members of the library, as the library was designed to contain non-synonymous mutations) that lie within the p19 promoter. However, attempts to package recombinant vector using these individual rep variants in the AAV helper construct did not increase viral titer.
A previous study conducted analogous mutagenesis on Rep: Jain et al., "Comprehensive mutagenesis maps the effect of all single-codon mutations in the AAV2 rep gene on AAV production" eLife 2024 (cited here as reference 19). It is not clear that this current study is a significant advance relative to the prior, quite comprehensive study. Both generated a library of non-synonymous mutations and observed fitness effects on Rep. Because this study sequenced the full rep, rather than barcodes associated with each rep variant, it found the enrichment in the synonymous mutations. However, these should ideally advance a basic understanding of Rep biology and/or result in better AAV production, but they did neither. It is speculated in the Discussion that the mutations generated additional GCTC motifs in p19, elements that mediate protein-DNA interactions. However, the role of GCTC motifs is speculative, and no transcriptional analysis is conducted. Furthermore, as discussed above, the mutations do not result in higher viral titers. Perhaps there's a transcriptional effect at the much lower copy numbers of vector genome present during library selection vs. rAAV packaging. They also found stop codons in Rep domains thought to be required for viral packaging, but functional studies confirming the screening findings are not conducted. As a result, the biological or technical relevance of the findings are extremely unclear, and thus the impact is relatively low.
The description of herring DNA co-transfection and cross-packaging (which is a well-known pitfall) are somewhat technical and arguably don't merit too much main manuscript attention.
Author response:
The following is the authors’ response to the current reviews.
Reviewer #1 (Public review):
Summary:
In this manuscript, Pakula et al. explore the impact of reactive oxygen species (ROS) on neonatal cerebellar regeneration, providing evidence that ROS activates regeneration through Nestin-expressing progenitors (NEPs). Using scRNA-seq analysis of FACS-isolated NEPs, the authors characterize injury-induced changes, including an enrichment in ROS metabolic processes within the cerebellar microenvironment. Biochemical analyses confirm a rapid increase in ROS levels following irradiation and forced catalase expression, which reduces ROS levels, and impairs external granule layer (EGL) replenishment post-injury.
Strengths:
Overall, the study robustly supports its main conclusion and provides valuable insights into ROS as a regenerative signal in the neonatal cerebellum.
Comments on revisions:
The authors have addressed most of the previous comments. However, they should clarify the following response:
*"For reasons we have not explored, the phenotype is most prominent in these lobules, that is why they were originally chosen. We edited the following sentence (lines 578-579):
First, we analyzed the replenishment of the EGL by BgL-NEPs in vermis lobules 3-5, since our previous work showed that these lobules have a prominent defect."*
It has been reported that the anterior part of the cerebellum may have a lower regenerative capacity compared to the posterior lobe. To avoid potential ambiguity, the authors should clarify that "the phenotype" and "prominent defect" refer to more severe EGL depletion at an earlier stage after IR rather than a poorer regenerative outcome. Additionally, they should provide a reference to support their statement or indicate if it is based on unpublished observations.
Our comment does not refer to a more severe EGL depletion at an earlier stage. There is instead poorer regeneration of the anterior region. The irradiation approach used provides consistent cell killing of GCPs across the cerebellum. This can be seen in Fig. 1c, e, g, i in our previous publication: Wojcinski, et al. (2017) Cerebellar granule cell replenishment post-injury by adaptive reprogramming of Nestin+ progenitors. Nature Neuroscience, 20:1361-1370). Also, Fig 2e, g, k, m in the paper shows that by P5 and P8, posterior lobule 8 recovers better than anterior lobules 1-5.
Reviewer #2 (Public review):
Summary:
The authors have previously shown that the mouse neonatal cerebellum can regenerate damage to granule cell progenitors in the external granular layer, through reprogramming of gliogenic nestin-expressing progenitors (NEPs). The mechanisms of this reprogramming remain largely unknown. Here the authors used scRNAseq and ATACseq of purified neonatal NEPs from P1-P5 and showed that ROS signatures were transiently upregulated in gliogenic NEPs ve neurogenic NEPs 24 hours post injury (P2). To assess the role of ROS, mice transgenic for global catalase activity were assessed to reduce ROS. Inhibition of ROS significantly decreased gliogenic NEP reprogramming and diminished cerebellar growth post-injury. Further, inhibition of microglia across this same time period prevented one of the first steps of repair - the migration of NEPs into the external granule layer. This work is the first demonstration that the tissue microenvironment of the damaged neonatal cerebellum is a major regulator of neonatal cerebellar regeneration. Increased ROS is seen in other CNS damage models, including adults, thus there may be some shared mechanisms across age and regions, although interestingly neonatal cerebellar astrocytes do not upregulate GFAP as seen in adult CNS damage models. Another intriguing finding is that global inhibition of ROS did not alter normal cerebellar development.
Strengths:
This paper presents a beautiful example of using single cell data to generate biologically relevant, testable hypotheses of mechanisms driving important biological processes. The scRNAseq and ATACseq analyses are rigorously conducted and conclusive. Data is very clearly presented and easily interpreted supporting the hypothesis next tested by reduce ROS in irradiated brains.
Analysis of whole tissue and FAC sorted NEPS in transgenic mice where human catalase was globally expressed in mitochondria were rigorously controlled and conclusively show that ROS upregulation was indeed decreased post injury and very clearly the regenerative response was inhibited. The authors are to be commended on the very careful analyses which are very well presented and again, easy to follow with all appropriate data shown to support their conclusions.
Weaknesses:
The authors also present data to show that microglia are required for an early step of mobilizing gliogenic NEPs into the damaged EGL. While the data that PLX5622 administration from P0-P5 or even P0-P8 clearly shows that there is an immediate reduction of NEPs mobilized to the damaged EGL, there is no subsequent reduction of cerebellar growth such that by P30, the treated and untreated irradiated cerebella are equivalent in size. There is speculation in the discussion about why this might be the case. Additional experiments and tools are required to assess mechanisms. Regardless, the data still implicate microglia in the neonatal regenerative response, and this finding remains an important advance.
As stated previously, the suggested follow up experiments while relevant are extensive and considered beyond the scope of the current paper.
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public review):
Summary:
In this manuscript, Pakula et al. explore the impact of reactive oxygen species (ROS) on neonatal cerebellar regeneration, providing evidence that ROS activates regeneration through Nestin-expressing progenitors (NEPs). Using scRNA-seq analysis of FACS-isolated NEPs, the authors characterize injury-induced changes, including an enrichment in ROS metabolic processes within the cerebellar microenvironment. Biochemical analyses confirm a rapid increase in ROS levels following irradiation, and forced catalase expression, which reduces ROS levels, and impairs external granule layer (EGL) replenishment post-injury.
Strengths:
Overall, the study robustly supports its main conclusion and provides valuable insights into ROS as a regenerative signal in the neonatal cerebellum.
Weaknesses:
(1) The diversity of cell types recovered from scRNA-seq libraries of sorted Nes-CFP cells is unexpected, especially the inclusion of minor types such as microglia, meninges, and ependymal cells. The authors should validate whether Nes and CFP mRNAs are enriched in the sorted cells; if not, they should discuss the potential pitfalls in sampling bias or artifacts that may have affected the dataset, impacting interpretation.
In our previous work, we thoroughly assessed the transgene using RNA in situ hybridization for Cfp, immunofluorescent analysis for CFP and scRNA-seq analysis for Cfp transcripts (Bayin et al., Science Adv. 2021, Fig. S1-2)(1), and characterized the diversity within the NEP populations of the cerebellum. Our present scRNA-seq data also confirms that Nes transcripts are expressed in all the NEP subtypes. A feature plot for Nes expression has been added to the revised manuscript (Fig 1E), as well as a sentence explaining the results. Of note, since this data was generated from FACS-isolated CFP+ cells, the perdurance of the protein allows for the detection of immediate progeny of Nes-expressing cells, even in cells where Nes is not expressed once cells are differentiated. Finally, oligodendrocyte progenitors, perivascular cells, some rare microglia and ependymal cells have been demonstrated to express Nes in the central nervous system; therefore, detecting small groups of these cells is expected (2-4). We have added the following sentence (lines 391-394):
“Detection of Nes mRNA confirmed that the transgene reflects endogenous Nes expression in progenitors of many lineages, and also that the perdurance of CFP protein in immediate progeny of Nes-expressing cells allowed the isolation of these cells by FACS (Figure 1E)”.
(2) The authors should de-emphasize that ROS signaling and related gene upregulation exclusively in gliogenic NEPs. Genes such as Cdkn1a, Phlda3, Ass1, and Bax are identified as differentially expressed in neurogenic NEPs and granule cell progenitors (GCPs), with Ass1 absent in GCPs. According to Table S4, gene ontology (GO) terms related to ROS metabolic processes are also enriched in gliogenic NEPs, neurogenic NEPs, and GCPs.
As the reviewer requested, we have de-emphasized that ROS signaling is preferentially upregulated in gliogenic NEPs, since we agree with the reviewer that there is some evidence for similar transcriptional signatures in neurogenic NEPs and GCPs. We added the following (lines 429-531):
“Some of the DNA damage and apoptosis related genes that were upregulated in IR gliogenic-NEPs (Cdkn1a, Phlda3, Bax) were also upregulated in the IR neurogenic-NEPs and GCPs at P2 (Supplementary Figure 2B-E).”
And we edited the last few sentences of the section to state (lines 453-459):
“Interestingly, we did not observe significant enrichment for GO terms associated with cellular stress response in the GCPs that survived the irradiation compared to controls, despite significant enrichment for ROS signaling related GO-terms (Table S4). Collectively, these results indicate that injury induces significant and overlapping transcriptional changes in NEPs and GCPs. The gliogenic- and neurogenic-NEP subtypes transiently upregulate stress response genes upon GCP death, and an overall increase in ROS signaling is observed in the injured cerebella.”
(3) The authors need to justify the selection of only the anterior lobe for EGL replenishment and microglia quantification.
We thank the reviewers for asking for this clarification. Our previous publications on regeneration of the EGL by NEPs have all involved quantification of these lobules, thus we think it is important to stay with the same lobules. For reasons we have not explored, the phenotype is most prominent in these lobules, that is why they were originally chosen. We edited the following sentence (lines 578-579):
“First, we analyzed the replenishment of the EGL by BgL-NEPs in vermis lobules 3-5, since our previous work showed that these lobules have a prominent defect.”
(4) Figure 1K: The figure presents linkages between genes and GO terms as a network but does not depict a gene network. The terminology should be corrected accordingly.
We have corrected the terminology and added the following (lines 487-489):
“Finally, linkages between the genes in differentially open regions identified by ATAC-seq and the associated GO-terms revealed an active transcriptional network involved in regulating cell death and apoptosis (Figure 1K).”
(5) Figure 1H and S2: The x-axis appears to display raw p-values rather than log10(p.value) as indicated. The x-axis should ideally show -log10(p.adjust), beginning at zero. The current format may misleadingly suggest that the ROS GO term has the lowest p-values.
Apologies for the mistake. The data represents raw p-values and the x-axis has been corrected.
(6) Genes such as Ppara, Egln3, Foxo3, Jun, and Nos1ap were identified by bulk ATAC-seq based on proximity to peaks, not by scRNA-seq. Without additional expression data, caution is needed when presenting these genes as direct evidence of ROS involvement in NEPs.
We modified the text to discuss the discrepancies between the analyses. While some of this could be due to the lower detection limits in the scRNA-seq, it also highlights that chromatin accessibility is not a direct readout for expression levels and further analysis is needed. Nevertheless, both scRNA-seq and ATAC-seq have identified similar mechanisms, and our mutant analysis confirmed our hypothesis that an increase in ROS levels underlies repair, further increasing the confidence in our analyses. Further investigation is needed to understand the downstream mechanisms. We added the following sentence (lines 478-481):
“However, not all genes in the accessible areas were differentially expressed in the scRNA-seq data. While some of this could be due to the detection limits of scRNA-seq, further analysis is required to assess the mechanisms of how the differentially accessible chromatin affects transcription.”
(7) The authors should annotate cell identities for the different clusters in Table S2.
All cell types have been annotated in Table S2.
(8) Reiterative clustering analysis reveals distinct subpopulations among gliogenic and neurogenic NEPs. Could the authors clarify the identities of these subclusters? Can we distinguish the gliogenic NEPs in the Bergmann glia layer from those in the white matter?
Thank you for this clarification. As shown in our previous studies, we can not distinguish between the gliogenic NEPs in the Bergmann glia layer and the white matter based on scRNA-seq, but expression of the Bergmann glia marker Gdf10 suggests that a large proportion of the cells in the Hopx+ clusters are in the Bergmann glia layer. The distinction within the major subpopulations that we characterized (Hopx-, Ascl1-expressing NEPs and GCPs) are driven by their proliferative/maturation status as we previously observed. We have included a detailed annotation of all the clusters in Table S2, as requested and a UMAP for mKi57 expression in Fig 1E. We have clarified this in the following sentence (lines 383-385):
“These groups of cells were further subdivided into molecularly distinct clusters based on marker genes and their cell cycle profiles or developmental stages (Figure 1D, Table S2).”
(9) In the Methods section, the authors mention filtering out genes with fewer than 10 counts. They should specify if these genes were used as background for enrichment analysis. Background gene selection is critical, as it influences the functional enrichment of gene sets in the list.
As requested, the approach used has been added to the Methods section of the revised paper. Briefly, the background genes used by the goseq function are the same genes used for the probability weight function (nullp). The mm8 genome annotation was used in the nullp function, and all annotated genes were used as background genes to compute GO term enrichment. The following was added (lines 307-308):
“The background genes used to compute the GO term enrichment includes all genes with gene symbol annotations within mm8.”
(10) Figure S1C: The authors could consider using bar plots to better illustrate cell composition differences across conditions and replicates.
As suggested, we have included bar plots in Fig. S1D-F.
(11) Figures 4-6: It remains unclear how the white matter microglia contribute to the recruitment of BgL-NEPs to the EGL, as the mCAT-mediated microglia loss data are all confined to the white matter.
We have thought about the question and had initially quantified the microglia in the white matter and the rest of the lobules (excluding the EGL) separately. However, there are very few microglia outside the white matter in each section, thus it is not possible to obtain reliable statistical data on such a small population. We therefore did not include the cells in the analysis. We have added this point in the main text (line 548).
“As a possible explanation for how white matter microglia could influence NEP behaviors, given the small size of the lobules and how the cytoarchitecture is disrupted after injury, we think it is possible that secreted factors from the white matter microglia could reach the BgL NEPs. Alternatively, there could be a relay system through an intermediate cell type closer to the microglia.” We have added these ideas to the Discussion of the revised paper (lines 735-738).
Reviewer #2 (Public review):
Summary:
The authors have previously shown that the mouse neonatal cerebellum can regenerate damage to granule cell progenitors in the external granular layer, through reprogramming of gliogenic nestin-expressing progenitors (NEPs). The mechanisms of this reprogramming remain largely unknown. Here the authors used scRNAseq and ATACseq of purified neonatal NEPs from P1-P5 and showed that ROS signatures were transiently upregulated in gliogenic NEPs ve neurogenic NEPs 24 hours post injury (P2). To assess the role of ROS, mice transgenic for global catalase activity were assessed to reduce ROS. Inhibition of ROS significantly decreased gliogenic NEP reprogramming and diminished cerebellar growth post-injury. Further, inhibition of microglia across this same time period prevented one of the first steps of repair - the migration of NEPs into the external granule layer. This work is the first demonstration that the tissue microenvironment of the damaged neonatal cerebellum is a major regulator of neonatal cerebellar regeneration. Increased ROS is seen in other CNS damage models including adults, thus there may be some shared mechanisms across age and regions, although interestingly neonatal cerebellar astrocytes do not upregulate GFAP as seen in adult CNS damage models. Another intriguing finding is that global inhibition of ROS did not alter normal cerebellar development.
Strengths:
This paper presents a beautiful example of using single cell data to generate biologically relevant, testable hypotheses of mechanisms driving important biological processes. The scRNAseq and ATACseq analyses are rigorously conducted and conclusive. Data is very clearly presented and easily interpreted supporting the hypothesis next tested by reduce ROS in irradiated brains.
Analysis of whole tissue and FAC sorted NEPS in transgenic mice where human catalase was globally expressed in mitochondria were rigorously controlled and conclusively show that ROS upregulation was indeed decreased post injury and very clearly the regenerative response was inhibited. The authors are to be commended on the very careful analyses which are very well presented and again, easy to follow with all appropriate data shown to support their conclusions.
Weaknesses:
The authors also present data to show that microglia are required for an early step of mobilizing gliogenic NEPs into the damaged EGL. While the data that PLX5622 administration from P0-P5 or even P0-P8 clearly shows that there is an immediate reduction of NEPs mobilized to the damaged EGL, there is no subsequent reduction of cerebellar growth such that by P30, the treated and untreated irradiated cerebella are equivalent in size. There is speculation in the discussion about why this might be the case, but there is no explanation for why further, longer treatment was not attempted nor was there any additional analyses of other regenerative steps in the treated animals. The data still implicate microglia in the neonatal regenerative response, but how remains uncertain.
Recommendations for the authors:
Reviewer #2 (Recommendations for the authors):
This is an exemplary manuscript.
The methods and data are very well described and presented.
I actually have very little to ask the authors except for an explanation of why PLX treatment was discontinued after P5 or P8 and what other steps of NEP reprogramming were assessed in these animals? Was NEP expansion still decreased at P8 even in the presence of PLX at this stage? Also - was there any analysis attempted combining mCAT and PLX?
We agree with the reviewer that a follow up study that goes into a deeper analysis of the role of microglia in GCP regeneration and any interaction with ROS signaling would interesting. However, it would require a set of tools that we do not currently have. We did not have enough PLX5622 to perform addition experiments or extend the length of treatment. Plexxikon informed us in 2021 that they were no longer manufacturing PLX5622 because they were focusing on new analogs for in vivo use, and thus we had to use what we had left over from a completed preclinical cancer study. We nevertheless think it is important to publish our preliminary results to spark further experiments by other groups.
References
(1) Bayin N. S. Mizrak D., Stephen N. D., Lao Z., Sims P. A., Joyner A. L. Injury induced ASCL1 expression orchestrates a transitory cell state required for repair of the neonatal cerebellum. Sci Adv. 2021;7(50):eabj1598.
(2) Cawsey T, Duflou J, Weickert CS, Gorrie CA. Nestin-Positive Ependymal Cells Are Increased in the Human Spinal Cord after Traumatic Central Nervous System Injury. J Neurotrauma. 2015;32(18):1393-402.
(3) Gallo V, Armstrong RC. Developmental and growth factor-induced regulation of nestin in oligodendrocyte lineage cells. The Journal of neuroscience : the official journal of the Society for Neuroscience. 1995;15(1 Pt 1):394-406.
(4) Huang Y, Xu Z, Xiong S, Sun F, Qin G, Hu G, et al. Repopulated microglia are solely derived from the proliferation of residual microglia after acute depletion. Nat Neurosci. 2018;21(4):530-40.
Author response:
The following is the authors’ response to the previous reviews
Reviewer #1 (Public review):
Jin et al. investigated how the bacterial DNA damage (SOS) response and its regulator protein RecA affects the development of drug resistance under short-term exposure to beta-lactam antibiotics. Canonically, the SOS response is triggered by DNA damage, which results in the induction of error-prone DNA repair mechanisms. These error-prone repair pathways can increase mutagenesis in the cell, leading to the evolution of drug resistance. Thus, inhibiting the SOS regulator RecA has been proposed as means to delay the rise of resistance.
In this paper, the authors deleted the RecA protein from E. coli and exposed this ∆recA strain to selective levels of the beta-lactam antibiotic, ampicillin. After an 8h treatment, they washed the antibiotic away and allowed the surviving cells to recover in regular media. They then measured the minimum inhibitory concentration (MIC) of ampicillin against these treated strains. They note that after just 8 h treatment with ampicillin, the ∆recA had developed higher MICs towards ampicillin, while by contrast, wild-type cells exhibited unchanged MICs. This MIC increase was also observed subsequent generations of bacteria, suggesting that the phenotype is driven by a genetic change.
The authors then used whole genome sequencing (WGS) to identify mutations that accounted for the resistance phenotype. Within resistant populations, they discovered key mutations in the promoter region of the beta-lactamase gene, ampC; in the penicillin-binding protein PBP3 which is the target of ampicillin; and in the AcrB subunit of the AcrAB-TolC efflux machinery. Importantly, mutations in the efflux machinery can impact the resistances towards other antibiotics, not just beta-lactams. To test this, they repeated the MIC experiments with other classes of antibiotics, including kanamycin, chloramphenicol, and rifampicin. Interestingly, they observed that the ∆recA strains pre-treated with ampicillin showed higher MICs towards all other antibiotic tested. This suggests that the mutations conferring resistance to ampicillin are also increasing resistance to other antibiotics.
The authors then performed an impressive series of genetic, microscopy, and transcriptomic experiments to show that this increase in resistance is not driven by the SOS response, but by independent DNA repair and stress response pathways. Specifically, they show that deletion of the recA reduces the bacterium's ability to process reactive oxygen species (ROS) and repair its DNA. These factors drive accumulation of mutations that can confer resistance towards different classes of antibiotics. The conclusions are reasonably well-supported by the data, but some aspects of the data and the model need to be clarified and extended.
Strengths:
A major strength of the paper is the detailed bacterial genetics and transcriptomics that the authors performed to elucidate the molecular pathways responsible for this increased resistance. They systemically deleted or inactivated genes involved in the SOS response in E. coli. They then subjected these mutants the same MIC assays as described previously. Surprisingly, none of the other SOS gene deletions resulted an increase in drug resistance, suggesting that the SOS response is not involved in this phenotype. This led the authors to focus on the localization of DNA PolI, which also participates in DNA damage repair. Using microscopy, they discovered that in the RecA deletion background, PolI co-localizes with the bacterial chromosome at much lower rates than wild-type. This led the authors to conclude that deletion of RecA hinders PolI and DNA repair. Although the authors do not provide a mechanism, this observation is nonetheless valuable for the field and can stimulate further investigations in the future.
In order to understand how RecA deletion affects cellular physiology, the authors performed RNA-seq on ampicillin-treated strains. Crucially, they discovered that in the RecA deletion strain, genes associated with antioxidative activity (cysJ, cysI, cysH, soda, sufD) and Base Excision Repair repair (mutH, mutY, mutM), which repairs oxidized forms of guanine, were all downregulated. The authors conclude that down-regulation of these genes might result in elevated levels of reactive oxygen species in the cells, which in turn, might drive the rise of resistance. Experimentally, they further demonstrated that treating the ∆recA strain with an antioxidant GSH prevents the rise of MICs. These observations will be useful for more detailed mechanistic follow-ups in the future.
Weaknesses:
Throughout the paper, the authors use language suggesting that ampicillin treatment of the ∆recA strain induces higher levels of mutagenesis inside the cells, leading to the rapid rise of resistance mutations. However, as the authors note, the mutants enriched by ampicillin selection can play a role in efflux and can thus change a bacterium's sensitivity to a wide range of antibiotics, in what is known as cross-resistance. The current data is not clear on whether the elevated "mutagenesis" is driven ampicillin selection or by a bona fide increase in mutation rate.
Furthermore, on a technical level, the authors employed WGS to identify resistance mutations in the treated ampicillin-treated wild-type and ∆recA strains. However, the WGS methodology described in the paper is inconsistent. Notably, wild-type WGS samples were picked from non-selective plates, while ΔrecA WGS isolates were picked from selective plates with 50 μg/mL ampicillin. Such an approach biases the frequency and identity of the mutations seen in the WGS and cannot be used to support the idea that ampicillin treatment induces higher levels of mutagenesis.
Finally, it is important to establish what the basal mutation rates of both the WT and ∆recA strains are. Currently, only the ampicillin-treated populations were reported. It is possible that the ∆recA strain has inherently higher mutagenesis than WT, with a larger subpopulation of resistant clones. Thus, ampicillin treatment might not in fact induce higher mutagenesis in ∆recA.
Comments on revisions:
Thank you for responding to the concerns raised previously. The manuscript overall has improved.
We sincerely thank the reviewer for raising this important point. In our initial submission, we acknowledge that our mutation analysis was based on a limited number of replicates (n=6), which may not have been sufficient to robustly distinguish between mutation induction and selection. In response to this concern, we have substantially expanded our experimental dataset. Specifically, we redesigned the mutation rate validation experiment by increasing the number of biological replicates in each condition to 96 independent parallel cultures. This enabled us to systematically assess mutation frequency distributions under four conditions (WT, WT+ampicillin, ΔrecA, ΔrecA+ampicillin), using both maximum likelihood estimation (MLE) and distribution-based fluctuation analysis (new Figure 1F, 1G, and Figure S5).
These expanded datasets revealed that:
(1) While the estimated mutation rate was significantly elevated in ΔrecA+ampicillin compared to ΔrecA alone (Fig. 1G),
(2) The distribution of mutation frequencies in ΔrecA+ampicillin was highly skewed with evident jackpot cultures (Fig. 1F), and
(3) The observed pattern significantly deviated from Poisson expectations, which is inconsistent with uniform mutagenesis and instead supports clonal selection from an early-arising mutational pool (Fig. S5).
Importantly, these new results do not contradict our original conclusions but rather extend and refine them. The previous evidence for ROS-mediated mutagenesis remains valid and is supported by our GSH experiments, transcriptomic analysis of oxidative stress genes, and DNA repair pathway repression. However, the additional data now indicate that ROS-induced variants are not uniformly induced after antibiotic exposure but are instead generated stochastically under the stress-prone ΔrecA background and then selectively enriched upon ampicillin treatment.
Taken together, we now propose a two-step model of resistance evolution in ΔrecA cells (new Figure 5):
Step i: RecA deficiency creates a hypermutable state through impaired repair and elevated ROS, increasing the probability of resistance-conferring mutations.
Step ii: β-lactam exposure acts as a selective bottleneck, enriching early-arising mutants that confer resistance.
We have revised both the Results and Discussion sections to clearly articulate this complementary relationship between mutational supply and selection, and we believe this integrated model better explains the observed phenotypes and mechanistic outcomes.
Reviewer #2 (Public review):
This study aims to demonstrate that E. coli can acquire rapid antibiotic resistance mutations in the absence of a DNA damage response. The authors employed a modified Adaptive Laboratory Evolution (ALE) workflow to investigate this, initiating the process by diluting an overnight culture 50-fold into an ampicillin selection medium. They present evidence that a recA- strain develops ampicillin resistance mutations more rapidly than the wild-type, as indicated by the Minimum Inhibitory Concentration (MIC) and mutation frequency. Whole-genome sequencing of recA- colonies resistant to ampicillin showed predominant inactivation of genes involved in the multi-drug efflux pump system, contrasting with wild-type mutations that seem to activate the chromosomal ampC cryptic promoter. Further analysis of mutants, including a lexA3 mutant incapable of inducing the SOS response, led the authors to conclude that the rapid evolution of antibiotic resistance occurs via an SOS-independent mechanism in the absence of recA. RNA sequencing suggests that antioxidative response genes drive the rapid evolution of antibiotic resistance in the recA- strain. They assert that rapid evolution is facilitated by compromised DNA repair, transcriptional repression of antioxidative stress genes, and excessive ROS accumulation.
Strengths:
The experiments are well-executed and the data appear reliable. It is evident that the inactivation of recA promotes faster evolutionary responses, although the exact mechanisms driving this acceleration remain elusive and deserve further investigation.
Weaknesses:
Some conclusions are overstated. For instance, the conclusion regarding the LexA3 allele, indicating that rapid evolution occurs in an SOS-independent manner (line 217), contradicts the introductory statement that attributes evolution to compromised DNA repair.
We thank the reviewer for this insightful observation, which highlights a central conceptual advance of our study. Our data indeed indicate that resistance evolution in ΔrecA occurs independently of canonical SOS induction (as shown by the lack of resistance in lexA3, dpiBA, and translesion polymerase mutants), yet is clearly associated with impaired DNA repair capacity (e.g., downregulation of polA, mutH, mutY).
This apparent “contradiction” reflects the dual role of RecA: it functions both as the master activator of the SOS response and as a key factor in SOS-independent repair processes. Thus, the rapid resistance evolution in ΔrecA is not due to loss of SOS, but rather due to the broader suppression of DNA repair pathways that RecA coordinates, which elevates mutational load under stress (This point is discussed in further detail in our response to Reviewer 1).
The claim made in the discussion of Figure 3 that the hindrance of DNA repair in recA- is crucial for rapid evolution is at best suggestive, not demonstrative. Additionally, the interpretation of the PolI data implies its role, yet it remains speculative.
We appreciate this comment and would like to respectfully clarify that our conclusion regarding the role of DNA repair impairment is supported by several independent lines of mechanistic evidence.
First, our RNA-seq analysis revealed transcriptional suppression of multiple DNA repair genes in ΔrecA cells following ampicillin treatment, including polA (DNA Pol I) and the base excision repair genes mutH, mutY, and mutM (Fig. 4K). This indicates that multiple repair pathways, including those responsible for correcting oxidative DNA lesions, are downregulated under these conditions.
Second, we observed a significant reduction in DNA Pol I protein expression as well as reduced colocalization with chromosomal DNA in ΔrecA cells, suggesting impaired engagement of repair machinery (Fig. 3C-E). These phenotypes are not limited to transcriptional signatures but extend to functional protein localization.
Third, and most importantly, resistance evolution was fully suppressed in ΔrecA cells upon co-treatment with glutathione (GSH), which reduces ROS levels. As GSH did not affect ampicillin killing (Fig. 4J), these findings suggest that mutagenesis and thus the emergence of resistance requires both ROS accumulation and the absence of efficient repair.
Therefore, we believe these data go beyond correlation and demonstrate a mechanistic role for DNA repair impairment in driving stress-associated resistance evolution in ΔrecA. We have revised the Discussion to emphasize the strength of this evidence while avoiding overstatement.
In Figure 2A table, mutations in amp promoters are leading to amino acid changes.
We thank the reviewer for spotting this inconsistency. Indeed, the ampC promoter mutations we identified reside in non-coding regulatory regions and do not result in amino acid substitutions. We have corrected the annotation in Fig. 2A and clarified in the main text that these mutations likely affect gene expression through transcriptional regulation, rather than protein sequence alteration.
The authors' assertion that ampicillin significantly influences persistence pathways in the wild-type strain, affecting quorum sensing, flagellar assembly, biofilm formation, and bacterial chemotaxis, lacks empirical validation.
We thank the reviewer for pointing this out. In the original version, we acknowledged transcriptional enrichment of genes related to quorum sensing, flagellar assembly, and chemotaxis in the wild-type strain upon ampicillin treatment. However, as we did not directly assess persistence phenotypes (e.g., biofilm formation or persister levels), we agree that such functional inferences were not fully supported. We have revised the relevant statements to focus solely on transcriptomic changes and have removed language suggesting direct effects on persistence pathways.
Figure 1G suggests that recA cells treated with ampicillin exhibit a strong mutator phenotype; however, it remains unclear if this can be linked to the mutations identified in Figure 2's sequencing analysis.
We appreciate the reviewer’s comment. This point is discussed in further detail in our response to Reviewer 1.
Reviewer #3 (Public review):
In the present work, Zhang et al investigate involvement of the bacterial DNA damage repair SOS response in the evolution of beta-lactam drug resistance evolution in Escherichia coli. Using a combination of microbiological, bacterial genetics, laboratory evolution, next-generation, and live-cell imaging approaches, the authors propose short-term (transient) drug resistance evolution can take place in RecA-deficient cells in an SOS response-independent manner. They propose the evolvability of drug resistance is alternatively driven by the oxidative stress imposed by accumulation of reactive oxygen species and compromised DNA repair. Overall, this is a nice study that addresses a growing and fundamental global health challenge (antimicrobial resistance).
Strengths:
The authors introduce new concepts to antimicrobial resistance evolution mechanisms. They show short-term exposure to beta-lactams can induce durably fixed antimicrobial resistance mutations. They propose this is due to comprised DNA repair and oxidative stress. Antibiotic resistance evolution under transient stress is poorly studied, so the authors' work is a nice mechanistic contribution to this field.
Weaknesses:
The authors do not show any direct evidence of altered mutation rate or accumulated DNA damage in their model.
We appreciate the reviewer’s comment. This point is discussed in further detail in our response to Reviewer 1.
Recommendations for the authors:
Reviewer #1 (Recommendations for the authors):
I would like to suggest two minor changes to the text.
(1) Re. WGS data.
The authors write in their response "We appreciate your concern regarding potential inconsistencies in the WGS methodology. However, we would like to clarify that the primary aim of the WGS experiment was to identify the types of mutations present in the wild type and ΔrecA strains after treatment of ampicillin, rather than to quantify or compare mutation frequencies. This purpose was explicitly stated in the manuscript.
I think the source of my confusion stemmed from this part in the text:
"In bacteria, resistance to most antibiotics requires the accumulation of drug resistance associated DNA mutations developed over time to provide high levels of resistance (29). To verify whether drug resistance associated DNA mutations have led to the rapid development of antibiotic resistance in recA mutant strain, we..."
I would change the phrase "verify whether drug resistance associated DNA mutations have led to the rapid development of antibiotic resistance in recA mutant strain" to "identify the types of mutations present in the wild type and ΔrecA strains after treatment of ampicillin." This would explicitly state what the sequencing was for (ie. ID-ing mutations). The current phrase can give the impression that WGS was used to validate rapid or high mutagenesis.
Thanks for this suggestion. We have revised this description to “In bacteria, resistance to most antibiotics requires the accumulation of drug resistance associated DNA mutations that can arise stochastically and, under stress conditions, become enriched through selection over time to confer high levels of resistance (33). Having observed a non-random and right-skewed distribution of mutation frequencies in ΔrecA isolates following ampicillin exposure, we next sought to determine whether specific resistance-conferring mutations were enriched in ΔrecA isolates following antibiotic exposure.”
(2) Re. whether the mutations are "induced" or "pre-existing."
The authors write:
"We appreciate your detailed feedback on the language used to describe our data. We understand the concern regarding the use of the term "induced" in relation to beta-lactam exposure. To clarify, we employed not only beta-lactam antibiotics but also other antibiotics, such as ciprofloxacin and chloramphenicol, in our experiments (data not shown). However, we observed that beta-lactam antibiotics specifically induced the emergence of resistance or altered the MIC in our bacterial populations. If resistance had pre-existed before antibiotic exposure, we would expect other antibiotics to exhibit a similar selective effect, particularly given the potential for cross-resistance to multiple antibiotics."
I think it is important to discuss the negative data for the other antibiotics (along with the other points made in your Reviewer response) in the main text.
This point is discussed in further detail in our response to Reviewer 1 (Public Review).
Reviewer #1 (Public review):
Summary:
This paper shows that maternal high-fat diet during lactation changes microglia morphology in the PVN, potentially to acquire a more active state. Further, the authors reveal that PVN microglia engulf AgRP terminals in the PVN during postnatal development, a previously unrecognized behavior. A notable finding of this paper is that pharmacological reduction of microglial cells can reverse weight gain and terminal loss in the offspring under maternal high fat diet conditions, even though an increase in microglial engulfment of AgRP+ terminals was not observed, suggesting an alternative mechanism. The data support these findings, although questions remain regarding the efficacy and timing of the pharmacological microglial knockdown.
Strengths
(1) The impact of microglia on hypothalamic synaptic pruning is poorly characterized, and thus, the findings herein are especially of interest.
Weaknesses
(1) Most minor concerns were addressed during revisions, including additional details in the methods and results sections that help interpret the data as presented.
(2) The AgRP staining is unclear. For example, in Figure 2, the figure legend says "labeled AgRP terminals (red)" (Fig 2A-D) but then concludes no difference in the number of "AgRP neurons" (Fig 2J). Is this quantification of AgRP+ neurons, terminals, or both?
(3) The PLX experiments are critical to their conclusion that during lactation, microglia in the PVN sculpt AgRP inputs; however, there is no demonstration that PLX treatment effectively eliminated microglia during this postnatal window. Microglia depletion was only assessed at P55, a month past the PLX treatment window making it unclear when and by what percentage the microglia were eliminated.
Author response:
The following is the authors’ response to the original reviews
Public Reviews:
Reviewer #1 (Public reviews):
(1) A cartoon paradigm of the HFD treatment window would be a helpful addition to Figure 1. Relatedly, the authors might consider qualifying MHFD as 'lactational MHFD.' Readers might miss the fact that the exposure window starts at birth.
This is a good suggestion. The MHFD-L model has been used previously (e.g. Vogt et al. 2014). We have included a cartoon of the MHFD-L model and the PLX treatments to Figure 4, which we feel helps the readers and thank the reviewer for the suggestion.
(2) More details on the modeling pipeline are needed either in Figure 1 or text. Of the ~50 microglia that were counted (based on Figure 1J), were all 50 quantified for the morphological assessments? Were equal numbers used for the control and MHFD groups? Were the 3D models adjusted manually for accuracy? How much background was detected by IMARIS that was discarded? Was the user blind to the treatment group while using the pipeline? Were the microglia clustered or equally spread across the PVN?
In response to this suggestion, we have expanded the description of the image analysis routine in the methods. The analysis focused on detailed changes in microglial morphology as opposed to overall changes in microglia throughout the PVH as a whole. Accordingly, we applied anatomically matched ROIs to the PVH for the measurements. As described in the methods, the Imaris Filaments tool was used to visualize microglia fully contained within a tissue section and a mask derived from the 3D model for these cells was used to isolate them for further analysis, thereby separating these cells from interstitial labeling corresponding to parts of cell processes or other labeling not associated with selected cells. There was no formal “background subtraction.” This was an error in the previous version of the manuscript and we have revised the methods to reflect the process actually used. The images were segmented (to enhance signal to noise for 3D rendering), and then a Gaussian filter was applied to improve edge detection, which facilitates the morphological measurements.
(3) Suggest toning back some of the language. For example: "...consistent with enhanced activity and surveillance of their immediate microenvironment" (Line 195) could be "...perhaps consistent with...". Likewise, "profound" (Lines 194, 377) might be an overstatement.
Revisions have been made to both the Introduction and Discussion to modulate our representation of this controversial issue.
(4) Representative images for AgRP+ cells (quantified in Figure 2J) are missing. Why not a co-label of Iba1+/AgRP+ as per Figure 1, 3? Also, what was quantified in Figure 2J - soma? Total immunoreactivity?
Because the density of AgRP labeling does not change in the ARH we omitted the red channel image (AgRP labeling) to highlight the similarity of the microglial morphology. To address the reviewer’s concerns, in the revised figure we have reconstituted the figure with both the green (microglial) and red (AgRP) channels depicted.
Figure 2J displays the numbers of AgRP neurons counted in the ARH in selected R01s through the ARH. The Methods section has been revised to include the visualization procedure used for the cell counts.
(5) For the PLX experiment:
a) "...we depleted microglia during the lactation period" (Line 234). This statement suggests microglia decreased from the first injection at P4 and throughout lactation, which is inaccurate. PLX5622 effects take time, upwards of a week. Thus, if PLX5622 injections started at P4, it could be P11 before the decrease in microglia numbers is stable. Moreover, by the time microglia are entirely knocked down, the pups might be supplementing some chow for milk, making it unclear how much PLX5622 they were receiving from the dam, which could also impact the rate at which microglia repopulation commences in the fetal brain. Quantifying microglia across the P4-P21 treatment window would be helpful, especially at P16, since the PVN AgRP microglia phenotypes were demonstrated and roughly when pups might start eating some chow. b) I am surprised that ~70% of the microglia are present at P21. Does this number reflect that microglia are returning as the pups no longer receive PLX5622 from milk from the dam? Does it reflect the poor elimination of microglia in the first place?
This is an important point and have revised the first sentence in section 2.3 to clarify the PLX treatment logic and added a cartoon to Fig. 4 to show the treatment timeline. The PLX5622 was not administered to the dams but daily to the pups. We also agree with the interpretation that PLX5622 depleted numbers of microglia, as supported by the microglial cell counts, rather than effected a complete elimination and have made revisions to clarify this distinction. Although mice were weighed at weaning, cellular measurements were only made in mice perfused at P55.
(6) Was microglia morphology examined for all microglia across the PVN? It is possible that a focus on PVNmpd microglia would reveal a stronger phenotype? In Figure 4H, J, AgRP+ terminals are counted in PVN subregions - PVNmpd and PVNpml, with PVNmpd showing a decrease of ~300 AgRP+ terminals in MHFD/Veh (rescued in MHFD/PLX5622). In Figure 1K, AgRP+ terminals across what appears to be the entire PVN decrease by ~300, suggesting that PVNmpd is driving this phenotype. If true, then do microglia within the PVNmpd display this morphology phenotype?
We have revised the description of the analysis procedures to clarify these points. All measurements were made in user defined, matched regions of interest according to morphological features of the PVH. No measurements were made that included the entire PVH and we revised the Methods section to improve clarity.
(7) What chow did the pups receive as they started to consume solid food? Is this only a MHFD challenge, or could the pups be consuming HFD chow that fell into the cage?
The pups were weaned onto the same normal chow diet that the dams received prior to MHFD-L treatment. The cages were inspected daily and minimal HFD spillage was observed, although we cannot rule out with certainty any contribution of the pups directly consuming the HFD. We have edited Methods section 5.2 for clarity.
(8) Figure 5: Does internalized AgRP+ co-localize with CD68+ lysosomes? How was 'internalized' determined?
This important point has been clarified by revisions to the Methods section.
(9) Different sample sizes are used across experiments (e.g., Figure 4 NCD n=5, MHFD n=4). Does this impact statistical significance?
Sample size does impact power of ANOVA with larger samples reducing the chance of errors. ANOVA is generally robust in the face of moderate departures from the assumption of equal sample sizes and equal variance such as we experienced in the PLX5622 experiment. Here we used t-tests to detect differences in a single variable between two groups and two-way ANOVA to compare treatment by diet and treatment changes in the PLX5622 studies. Additional detail has been added to the Methods section to clarify this point.
Reviewer #2 (Public reviews):
(1) Under chow-fed conditions, there is a decrease in the number of microglia in the PVH and ARH between P16 and P30, accompanied by an increase in complexity/volume. With the exception of PVH microglia at P16, this maturation process is not affected by MHFD. This "transient" increase in microglial complexity could also reflect premature maturation of the circuit.
This is an interesting possibility that requires future investigation (see response to Recommended Suggestions, above).
(2) The key experiment in this paper, the ablation of microglia, was presumably designed to prevent microglial expansion/activation in the PVH of MHFD pups. However, it also likely accelerates and exaggerates the decrease in cell number during normal development regardless of maternal diet. Efforts to interpret these findings are further complicated because microglial and AgRP neuronal phenotypes were not assessed at earlier time points when the circuit is most sensitive to maternal influences.
We agree that evaluations of microglia and hypothalamic circuits at many more time points would indeed be informative (see comments above).
(3) Microglial loss was induced broadly in the forebrain. Enhanced AgRP outgrowth to the PVH could be caused by actions elsewhere, such as direct effects on AgRP neurons in the ARH or secondary effects of changes in growth rates.
A local effect of microglia in the ARH that affects growth of AgRP axons remains a distinct possibility that deserves a targeted examination (see response to Recommended Suggestions, above).
(4) Prior publications from the authors and other groups support the idea that the density of AgRP projections to the PVH is primarily driven by factors regulating outgrowth and not pruning. The failure to observe increased engulfment of AgRP fibers by PVH microglia is therefore not surprising. The possibility that synaptic connectivity is modulated by microglia was not explored.
Synaptic pruning and regulation of axon targeting are not mutually exclusive processes and microglia may participate in both. Here we evaluated innervation of the PVH, which is sensitive to MHFD-L exposure, and engulfment of AgRP terminals by microglia, which does appear to be altered by MHFD-L. Given previous observations of terminal engulfment by microglia in other brain regions in response to environmental changes (e.g. prolonged stress) it is not unreasonable to expect this outcome in the offspring of MHFD-L dams. In future work it will be important to profile multiple cell types in the PVH for microglial dependent and MHFDL-sensitive changes in targeting of AgRP axons. Equally important is a full characterization of postsynaptic changes in PVH neurons.
Reviewer #3 (Public reviews):
There was no attempt to interrogate microglia in different parts of the hypothalamus functionally. Morphology alone does not reflect a potential for significant signaling alterations that may occur within and between these and other cell types.
The authors should discuss the limitations of their approach and findings and propose future directions to address them.
We agree that evaluations of microglia and hypothalamic circuits at many more time points that include analyses of multiple regions would indeed be informative. We have added statements to the manuscript that address the limitations of our experimental approach and suggest future studies that will extend understanding of underlying mechanisms beyond those investigated here.
Recommendations for the authors:
Reviewing Editors Comments:
(1) The Abstract is 405 words and should be shortened to less than 200 words.
The abstract has been edited to 200 words.
(2) The authors might consider raising the question in the Introduction of whether reduced AgRP innervation of the PVN in MHFD-treated mice is due to decreased axonal growth, enhanced microglial-mediated pruning, or a combination of both. The potential effects on axonal growth should be given more consideration.
This is an important point that we agree deserves additional consideration in the manuscript. Our past work has focused on leptin’s ability to influence axonal targeting of PVH neurons by AgRP and PPG neurons through a cell-autonomous mechanism and our conclusion is that leptin primarily induces axon growth. Because in this study our design did not focus on changes in axon growth over time but on regional changes in microglia and their interactions with AgRP terminals we did not want to divert attention from our logic in the introduction by highlighting multiple mechanisms. However, we have added a brief mention in the Introduction and have expanded consideration of axonal growth effects to the Discussion. Distinguishing between microglia’s role in synaptic density or axon targeting in this pathway is an important goal of future work.
(3) Line 37, a high-fat diet should be defined here as HFD and used consistently thereafter. Note that "high-fat-diet exposure" requires two hyphens.
The suggested revisions have been made throughout the manuscript.
(4) Line 38 and elsewhere, MHFD does not adequately describe the treatment being limited to the lactation period, perhaps MLHFD would be better or just LHFD (because the pups can't lactate).
The suggested revisions have been made throughout the manuscript, and we have used MHFD-L to describe maternal consumption of a high-fat diet that is restricted to the lactation period.
(5) Line 110, leptin-deficient mice (add hyphen).
(6) Line 183, NCD should be defined.
The suggested revisions have been made throughout the manuscript.
(7) Lines 237- 238, it is not clear what is widespread in the rostral forebrain. Is it the loss of microglia? What is the dividing point between the rostral and caudal forebrain? Were microglia depleted in the caudal forebrain too?
We have revised this section of the manuscript to focus the description on the hypothalamus alone and specify that the reduction in microglial density is not restricted to the PVH.
(8) Line 245, microglial-mediated effects (add hyphen).
(9) Line 247, vehicle-treated mice (add hyphen).
The suggested revisions have been made throughout the manuscript.
(10) Line 457, when referring to genes, the approved gene name should be used in italics, AgRP should be Agrp (italics).
The suggested revision has been made throughout the manuscript.
(11) Line 459, the name of the Syn-Tom mice in the Key Resource table, Methods, and Text should be consistent. It would be best to use the formal name of the Ai34 line of mice on the JAX website.
The suggested revisions have been made throughout the manuscript.
(12) Figure 1G H, and I um should have Greek micro; Fig. 1J and K, Replace # with Number. The same suggestions apply to all the other figures.
Both the manuscript and figures have been revised in accordance with this recommendation.
(13) Figures 4 G, H, I and J. and Figures 5 M and O. The font size is too small to see well.
Fonts have been changed in the figures to improve visibility.
Reviewer #1 (Recommendations for the authors):
(1) Figures are out of order in the text. For example, Figure 1A is followed next by the results for Figure 1J instead of Figure 1B.
We regret that the organization of figure panels makes for awkward matching for the reader as they proceed through the text. We designed the figures to facilitate comparisons between cellular responses and differences in labeling. After evaluating a reorganization, we decided to maintain the original panel configurations, but have revised the text to more closely follow the presentation of cellular features in the figures.
(2) Figure 1B.: All images lack scale bars.
(3) Line 433 - 'underlie' is spelled wrong.
(4) Rosin et al should be 2019 and not 2018.
These corrections have been implemented in the revised text and figures.
(5) The statement that "the effects of MHFD on microglial morphology in the PVH of offspring display both temporal and regional specificity, which correspond to a decrease in the density of AgRP inputs to the PVH" (Line 196) needs clarification, as the phrase "regional specificity" has not been substantiated in this section even though it is discussed later.
We agree with this comment and have revised section 2.1 to more closely match the data presented to this point in the manuscript.
Reviewer #2 (Recommendations for the authors):
(1) The claim of "spatial specificity" in the effects of MHFD on microglia is based on an increase in the complexity/volume of microglia at P16 in the PVH that was not seen in the ARH or BNST. The transient nature of the effect raises several questions: Does the effect on the PVH represent premature maturation?
This is an interesting suggestion. However, given how little is known about microglial maturation in the hypothalamus it is difficult to address. It is indeed possible that microglia mature at different rates in each AgRP target, and that MHFD-L exposure alters the rate of maturation in some regions but not others. This will require a great deal more analysis of both microglia and ARH projections to understand fully (see below).
(2) To support their central claim that microglia in the PVH "sculpt the density of AgRP inputs to the PVH" the authors report effects on Iba1+ cells in the PVH of chow-fed dams at P55, body weight at P21, and AgRP projections in the PVH at an unspecified age. It is hard to understand what is happening across "normal" development in chow-fed dams since the number of Iba1+ cells decreases from ~50 to ~25 between P16 and P30 (Figure 1), but then increases to >60 cells at P55 (Figure 4). Given the large fluctuations in microglial population across time, analyzing the same parameters (i.e. microglial number/morphology in the ARH and PVH, AgRP neuronal number in the ARH, and fiber density in the PVH, and body weight) across time points before, during and after the critical period in chow and MHFD conditions would be very helpful.
The time points we evaluated were chosen to be during and after the previously determined critical period for development of AgRP projections to the PVH, which were then compared with adults (which were all P55) to assess longevity of the effects. We have incorporated revisions to improve the clarity of when measurements were assessed, and treatments implemented. Defining the cellular dynamics of microglia across time remains a major challenge for the field and will certainly be informed by future studies with additional time points, as well as by in vivo imaging studies focused on regions identified here. Although such studies are beyond the scope of the present work, their completion would advance our current understanding of how microglia respond to nutritional changes during development of feeding circuits.
(3) As microglia are also ablated in the ARH, direct effects on AgRP neurons or indirect effects via changes in growth rates could also contribute to increased AgRP fiber density in the PVH. In support of the first possibility, postnatal microglial depletion increases the number of AgRP neurons (Sun, et al. 2023).
We agree with the suggestion, also raised by the Reviewing Editor, which has been addressed briefly in the Introduction, and in more detail by revisions to the Discussion section.
(4) The failure to assess alpha-MSH fibers in the same animals was a missed opportunity. They are also affected by MHFD but likely involve a distinct mechanism (Vogt, et al 2014).
Given the paired interest in POMC neurons and AgRP neurons I understand the reviewer’s comment. We chose to focus solely on AgRP neurons because we do not currently have a way to genetically target axonal labeling exclusively to POMC neurons due to the shared precursor origin of POMC neurons and a percentage of NPY neurons in the ARH, as shown by Lori Zeltser’s laboratory. Moreover, the elegant work by Vogt et al. focused on responses of POMC neurons in the MHFD-L model. However, it certainly remains possible that microglia in the PVH interact with terminals derived from POMC neurons, as well as with terminals derived from other afferent populations of neurons.
(5) All statistical analyses involved unpaired t-tests. Two-way ANOVAs should be used to assess the effects of age and HFD and interactions between these factors.
We used t-tests to detect differences in a single variable between two groups and two-way ANOVA to compare treatment by diet and treatment changes in the PLX5622 studies. Additional detail has been added to the Methods section and information added to the figure legend for Fig. 4 to clarify this point.
Reviewer #3 (Recommendations for the authors):
I suggest exploring the deeper characterization of the microglia in various parts of the hypothalamus in different conditions. This could include cytokine assessment or spatial transcriptomic.
We agree that a great deal more work is needed to improve our understanding of how microglia impact hypothalamic development more broadly and to identify underlying molecular mechanisms. We are hopeful that the data presented here will motivate additional study of microglial dynamics in multiple hypothalamic regions, as well as detailed studies of cellular signaling events for factors derived from MHFD-L dams that impact neural development in the hypothalamus.
Author response:
Public Reviews:
Reviewer #1 (Public review):
Summary:
The manuscript finds a negative relationship between tuberculin skin test-induced type I interferon activity with chest X-ray tuberculosis severity in humans. This evidence is between incomplete and solid. It needs a bioinfomatics/transcriptomics reviewer to make a more insightful judgement. The manuscript demonstrates a convincing role for Stat2 in controlling Mycobacterium marinum infection in zebrafish embryos, incomplete data are presented linking reduced leukocyte recruitment to the infection susceptibility phenotype.
Strengths:
(1) An interesting analysis of TST response correlated with chest X-ray pathology.
(2) Novel data on a protective role for Stat2 in a natural host-mycobacterial species infection pairing.
We appreciate the reviewer’s positive comments.
Weaknesses:
(1) The transcriptional modules are very large sets of genes that do not present a clear picture of what is actually being measured relative to other biological pathways.
The transcriptional module analysis is a major strength of our approach. These gene signatures are derived from independent experiments, most of which have been previously published/validated [1,2]. To clarify, they represent co-regulated gene sets downstream of signalling pathways. Increased number of genes in these modules increases their combinatorial specificity for a given biological pathway. In the human data, they serve as orthogonal validation for the bioinformatic analysis showing enrichment of the type I IFN pathway among TST transcriptome genes that are negatively correlated with radiographic disease severity in pulmonary TB (see Figure 2). Importantly, our modules confirm the relationship with type I IFN signalling (see Figure 2E) by discriminating from type II IFN signalling, which is not statistically significantly correlated with radiographic TB severity (see Figure S6C-E).
(2) The link between infection-Stat2-leukocyte recruitment and containment of infection is plausible, but lacks a specific link to the first part of the manuscript.
For clarification, the first part of the study seeks to identify immune response pathways that relate to severity of human disease, leading to the identification of type I IFN signalling. Since the human data are limited to an observational analysis in which we cannot test causality, the second part of our study uses a genetically tractable experimental model to test the hypothesis that type I IFN signalling is host-protective and explore possible mechanisms for a beneficial effect. This leads to the observation that type I IFN responses contribute to early myeloid cell recruitment to the site of infection, that has previously been shown to be crucial for containment of mycobacterial infection in zebrafish larvae. We will further evaluate the introduction and results sections to ensure a clear link between the human and zebrafish work.
Major concerns
(1) Line 158: The two transcriptional modules should be placed in the context of other DEG patterns. The macrophage type I interferon module, in particular, is quite large (361 genes). Can this be made more granular in terms of type I IFN ligands and STAT2-dependent genes?
We respectfully disagree with this comment. For clarification, the 360 gene module reflects the zebrafish larval response to IFNphi1 protein [3]. Type I IFNs are known to induce hundreds of interferon stimulated genes [4]. As explained above, the size of the modules increases specificity for a given signalling pathway. In this case, we are most interested in discriminating type I and type II IFN signalling pathways that represent very different upstream biological processes. The discrimination we achieve with our modular approach is a major advance over previous reports of gene signatures in TB that do not discriminate between the two pathways. In this study, we did not discriminate between signalling downstream of type I IFN ligands and STAT2, consistent with existing literature showing that type I IFN signalling is STAT2 dependent [5,6].
(2) The ifnphi1 injection into mxa:mCherry stat2 crispants is a nice experiment to demonstrate loss of type I IFN responsiveness. Further data is required to demonstrate if important mycobacterial control pathways (IFNy, TNF, il6?, etc) are intact in stat2 crispants before being able to conclude that these phenotypes are specific to type I IFN.
Thank you for the positive comment. We acknowledge this point and will attempt to evaluate whether pro-inflammatory cytokine responses are intact in stat2 CRISPants by qPCR or bulk RNAseq. However, these experiments may prove inconclusive because of the limited sensitivity in this approach.
Reviewer #2 (Public review):
Summary:
This study shows that type I interferon (IFN-I) signaling helps protect against mycobacterial infection. Using human gene expression data and a zebrafish model, the authors find that reduced IFN-I activity is linked to more severe disease. They also show that zebrafish lacking the IFN-I signaling gene stat2 are more vulnerable to infection due to poor macrophage migration. These results suggest a protective role for IFN-I in mycobacterial disease, challenging previous findings from other animal models.
Strengths:
Strengths of the manuscript include the use of human clinical samples to support relevance to disease, along with a genetically tractable zebrafish model that enables mechanistic insight.
We welcome the reviewer’s positive summary of our study.
Weaknesses:
(1) The manuscript presents intriguing human data showing an inverse correlation between IFN-I gene signatures and TB disease, but the findings remain correlative and may be cohort-specific. Given that the skin is not a primary site of TB and is relatively immunotolerant, the biological relevance of downregulated IFN-I-related genes in this tissue to systemic or pulmonary TB is unclear.
We agree with the reviewer that the observational human data are correlative. That is precisely why we extend the study to undertake mechanistic studies in a genetically tractable animal model, using M. marinum infection of zebrafish larvae. In the introduction, we already provide a detailed rationale for the strengths of the TST model to study human immune responses to a standardised mycobacterial challenge. This approach mitigates against the confounding of heterogeneity in bacterial burden and sampling different stages of the natural history of infection in conventional observational human studies. Therefore, the application of the TST is a major strength of this study. We do not understand the context in which the reviewer suggests the skin is immunotolerant. In the present study and previous work we provide molecular level analysis of the TST as a robust cell mediated immune response that reflects molecular perturbation in granuloma from the site of pulmonary TB disease 1.
(2) The reliance on stat2 CRISPants in zebrafish offers a limited view of IFN-I signaling. Including additional crispant lines targeting other key regulators (e.g., ifnar1, tyk2, irf3, irf7) would strengthen the interpretation and clarify whether the observed effects reflect broader IFN-I pathway disruption.
We respectfully disagree with this comment. Our objective was to test the role of type I IFN signalling in M. marinum infection of zebrafish. We show that stat2 deletion effectively disrupts type I IFN signalling (Figure S8). Therefore, we do not see a compelling rationale to evaluate other molecules in the signalling pathway.
(3) The conclusion that IFN-I is protective contrasts with established findings from murine and non-human primate models, where IFN-I is often detrimental. While the authors highlight species differences, the lack of functional human data and reliance on M. marinum in zebrafish limit the translational relevance. A more balanced discussion addressing these discrepancies would improve the manuscript.
We acknowledge that our findings contrast with the prevailing view in published literature to date. We will further review the discussion to see how we can elaborate on the potential strengths and weaknesses of different experimental approaches, which may underpin these discrepancies.
(4) Quantification of bacterial burden using fluorescence intensity alone may not accurately reflect bacterial viability. Complementary methods, such as qPCR for bacterial DNA, would provide a more robust assessment of antimicrobial activity.
We and others have previously validated the use of the quantitative measures of fluorescence, used here as a measure of bacterial load [7,8]. Importantly, our measurements do not rely purely on the total fluorescence signal, but also measures of dissemination of infection, for which we see consistent findings. It is also widely recognised that DNA measurements do not necessarily correlate well with bacterial viability. Therefore, we respectfully disagree that a PCR-based approach will add substantial value to our existing analysis.
(5) Finally, the authors should clarify whether impaired macrophage recruitment in stat2 crispants results from defects in chemotaxis, differentiation, or survival, and address discrepancies between their human blood findings and prior studies.
We acknowledge that these are important questions. Our data show that stat2 disruption does not impact total macrophage numbers at baseline (Figure 4A,B) and therefore do not support any effect of Stat2 signalling on steady state macrophage survival or differentiation. The downregulation of macrophage mpeg1 expression in M. marinum infection precludes long-term follow-up of these cells in the context of infection [9]. Therefore, we cannot currently test the hypothesis that Stat2 signalling may influence death of macrophages recruited to the site of infection or make them more susceptible to the cytopathic effects of direct mycobacterial infection. We will attempt to confirm using short-term time-lapse imaging that cellular migration to the site of hindbrain M. marinum infection is reduced in stat2 deficient zebrafish. On the strength of what is possible to test and the established role of type I IFNs in induction of several chemokines [10,11], the most likely effect is that Stat2 signalling increases recruitment through chemokine production. We are exploring the possibility of testing changes to the chemokine profile in stat2 CRISPants by qPCR or bulk RNAseq, but these experiments may prove inconclusive because of the limitations of sensitivity in this approach.
We recognize that our finding of no relationship between peripheral blood type I IFN activity and severity of human TB contrasts with that of previous studies. As stated in the discussion, the most likely explanation for this is our use of transcriptional modules which reflect exclusive type I IFN responses. The signatures used in other studies include both type I and type II IFN inducible genes and therefore also reflect IFN gamma driven responses.
Reviewer #3 (Public review):
Summary:
In this manuscript, the authors presented an interesting study providing an insight into the role of Type-I interferon responses in tuberculosis (TB) pathogenesis by combining transcriptome analysis of PBMCs and TST from tuberculosis patients. The zebrafish model was used to identify the changes in the innate immune cell population of macrophages and neutrophils. The findings suggested that Type-I interferon signatures inversely correlated with disease severity in the TST transcriptome data. The authors validated the observations by CRISPR-mediated disruption of stat2 (a critical transcription factor for type I interferon signaling) in zebrafish larvae, showing increased susceptibility to M. marinum infection. Traditionally, type-I interferon responses have been viewed as detrimental in mycobacterial infections, with studies suggesting enhanced susceptibility in certain mouse models. The study tried to identify and further characterize the understanding of the role of type-I interferons in TB.
Strengths:
Traditionally, type-I interferon responses have been viewed as detrimental in mycobacterial infections, with studies suggesting enhanced susceptibility in certain mouse models. The study tried to further understand the role of type-I interferons in TB pathogenesis.
We thank the reviewer for their summary.
Weaknesses:
Though the study showed an inverse correlation of Type-I interferon with radiological features of TB, the molecular mechanism is largely unexplored in the study, which is making it difficult to understand the basis of the results shown in the manuscript by the authors.
We respectfully disagree with this comment. The observations in the human data lead to the hypothesis that type I IFN responses may be host-protective, which we then test specifically in the zebrafish model, and explore candidate mechanisms, focussing on myeloid cell recruitment to the site of infection.
References
(1) Bell, L.C.K., Pollara, G., Pascoe, M., Tomlinson, G.S., Lehloenya, R.J., Roe, J., Meldau, R., Miller, R.F., Ramsay, A., Chain, B.M., et al. (2016). In Vivo Molecular Dissection of the Effects of HIV-1 in Active Tuberculosis. PLoS Pathog. 12, e1005469. https://doi.org/10.1371/journal.ppat.1005469.
(2) Pollara, G., Turner, C.T., Rosenheim, J., Chandran, A., Bell, L.C.K., Khan, A., Patel, A., Peralta, L.F., Folino, A., Akarca, A., et al. (2021). Exaggerated IL-17A activity in human in vivo recall responses discriminates active tuberculosis from latent infection and cured disease. Sci. Transl. Med. 13, eabg7673. https://doi.org/10.1126/scitranslmed.abg7673.
(3) Levraud, J.-P., Jouneau, L., Briolat, V., Laghi, V., and Boudinot, P. (2019). IFN-Stimulated Genes in Zebrafish and Humans Define an Ancient Arsenal of Antiviral Immunity. J. Immunol. Baltim. Md 1950 203, 3361–3373. https://doi.org/10.4049/jimmunol.1900804.
(4) Schoggins, J.W. (2019). Interferon-Stimulated Genes: What Do They All Do? Annu. Rev. Virol. 6, 567–584. https://doi.org/10.1146/annurev-virology-092818-015756.
(5) Blaszczyk, K., Nowicka, H., Kostyrko, K., Antonczyk, A., Wesoly, J., and Bluyssen, H.A.R. (2016). The unique role of STAT2 in constitutive and IFN-induced transcription and antiviral responses. Cytokine Growth Factor Rev. 29, 71–81. https://doi.org/10.1016/j.cytogfr.2016.02.010.
(6) Begitt, A., Droescher, M., Meyer, T., Schmid, C.D., Baker, M., Antunes, F., Knobeloch, K.-P., Owen, M.R., Naumann, R., Decker, T., et al. (2014). STAT1-cooperative DNA binding distinguishes type 1 from type 2 interferon signaling. Nat. Immunol. 15, 168–176. https://doi.org/10.1038/ni.2794.
(7) Stirling, D.R., Suleyman, O., Gil, E., Elks, P.M., Torraca, V., Noursadeghi, M., and Tomlinson, G.S. (2020). Analysis tools to quantify dissemination of pathology in zebrafish larvae. Sci. Rep. 10, 3149. https://doi.org/10.1038/s41598-020-59932-1.
(8) Takaki, K., Davis, J.M., Winglee, K., and Ramakrishnan, L. (2013). Evaluation of the pathogenesis and treatment of Mycobacterium marinum infection in zebrafish. Nat. Protoc. 8, 1114–1124. https://doi.org/10.1038/nprot.2013.068.
(9) Benard, E.L., Racz, P.I., Rougeot, J., Nezhinsky, A.E., Verbeek, F.J., Spaink, H.P., and Meijer, A.H. (2015). Macrophage-expressed perforins mpeg1 and mpeg1.2 have an anti-bacterial function in zebrafish. J. Innate Immun. 7, 136–152. https://doi.org/10.1159/000366103.
(10) Lehmann, M.H., Torres-Domínguez, L.E., Price, P.J.R., Brandmüller, C., Kirschning, C.J., and Sutter, G. (2016). CCL2 expression is mediated by type I IFN receptor and recruits NK and T cells to the lung during MVA infection. J. Leukoc. Biol. 99, 1057–1064. https://doi.org/10.1189/jlb.4MA0815-376RR.
(11) Buttmann, M., Merzyn, C., and Rieckmann, P. (2004). Interferon-beta induces transient systemic IP-10/CXCL10 chemokine release in patients with multiple sclerosis. J. Neuroimmunol. 156, 195–203. https://doi.org/10.1016/j.jneuroim.2004.07.016.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary
Enhancer RNAs (eRNAs) are early indicators of transcription factor (TF) activity and can identify distinct molecular subtypes and pathological outcomes in breast cancer. In this study, Patel et al. analysed 302,951 polyadenylated eRNA loci from 1,095 breast cancer patients using RNA-seq data, applying machine learning (ML) to classify eRNAs associated with specific molecular subtypes and survival. They discovered subtype-specific eRNAs that implicate both established and novel regulatory pathways and TFs, as well as prognostic eRNAs -specifically, LumA and HER2-survival- that distinguish favorable from poor survival outcomes. Overall, this ML-based approach illustrates how eRNAs reveal the molecular grammar and pathological implications underlying breast cancer heterogeneity.
Major comments
Minor comments
Referee cross-commenting
I share Reviewer #1's opinion that the manuscript should assess whether mRNA or eRNA is the stronger predictor of breast cancer subtypes and clinical outcomes. It will greatly improve the novelty if eRNA is shown to be a better indicator for cancer characterization.
Also, I strongly concur with Reviewer #3 that the current informatics approach is superficial and that several conclusions are contentious. The authors need to resolve the inconsistencies in their ML statistics and the potentially misleading interpretations of the ChIP‑seq and motif‑enrichment results.
It is further recommended that, building Reviewer #3's comment, the study integrate eRNA signatures with their proximal genes to address 1) whether genes located near these enhancers are differentially expressed-and correlated with enhancer activity-across cancer subtypes, and 2) whether it provides insights into understanding the enhancer-gene regulatory architecture in a subtype-specific context.
General Assessment
This study provides insights into the potential use of eRNA to classify breast cancer subtypes and refine prognostic markers. A strength is the integration of large-scale RNA-seq data with machine learning to identify eRNA signatures in biologically-meaningful patient samples, revealing both established and novel TF networks. The study also discovered eRNA clusters that correlate with the survival of patients, thus providing strong clinical implications. However, the ML approach yields several inconsistencies-for instance, unsatisfactory classification results for the Her2 subtype as well as the confused statistical metrics in the results. Furthermore, the ML model struggles to differentiate more nuanced molecular classes (e.g., LumA vs. LumB) and higher-level histological subtypes (e.g., lobular vs. ductal), thus limiting its power to dissect more delicate pathological and molecular mechanisms. Another limitation worth noting of this ML approach is the exclusive use of only polyadenylated eRNAs via RNA-seq, which excludes perhaps the more prominent 2D eRNA expressed in regulatory enhancers. Moreover, certain datasets appear to be of suboptimal quality, leading to assertions that would benefit from additional supporting evidence. Altogether, while the study offers a promising angle on eRNA-based tumor stratification, more robust experimental validations are needed to resolve inconsistencies and clarify the mechanistic underpinnings.
Advance
Conceptually, the study highlights the potential for eRNA-based signatures to capture regulatory variation beyond classical markers. However, the utility of these signatures is constrained by the focus on polyadenylated transcripts alone, likely underrepresenting key enhancer regions, and certain evidence presented in this study is not substantial enough to support some statements. While the work adds an important dimension to the understanding of enhancer biology in breast cancer, the resulting insights are partly hampered by limitations in data coverage and quality.
Audience
The primary audience includes cancer epigenetics, functional genomics, and bioinformatics researchers who are interested in leveraging eRNAs as biomarkers and dissecting complex regulatory networks in breast cancer. Clinically oriented scientists focusing on molecular diagnostics may also find relevance in the authors' approach to stratify subtypes and outcomes. The research is most relevant to a specialized audience within basic and translational cancer genomics, as well as computational biology groups interested in eRNA analysis.
Field of Expertise
I evaluate this manuscript as a researcher specializing in cancer epigenetics, functional genomics, and NGS-based data analysis. Parts of the manuscript touching on clinical outcome measures may require additional review from practicing oncologists.
Briefing : Une cure de détox numérique
Ce document explore les thèmes de l'addiction aux smartphones et aux écrans, des tentatives de déconnexion, et des effets de ces expériences sur les jeunes et les adultes.
Il met en contraste un programme de détox numérique d'une semaine pour des lycéens en Allemagne et un camp de survie plus radical en Pologne axé sur le retour à la nature.
Thèmes Principaux :
L'Addiction Numérique et les "Digital Natives" : Le document souligne que la génération Z est la première à avoir grandi avec des smartphones dès leur plus jeune âge. L'utilisation constante des appareils, même pendant les activités quotidiennes comme cuisiner, manger, réviser et avant de dormir, est généralisée. Cela est décrit comme une "drogue" par un enseignant.
Expériences de Déconnexion : Le document présente deux approches de détox numérique :
Une semaine sans smartphone et ordinateur pour des lycéens allemands.
Un camp de survie en Pologne mené par Camille Fialkovski, axé sur le renoncement aux commodités modernes et un retour à la vie dans la nature.
Les Défis et les Effets de la Déconnexion : Les expériences de déconnexion révèlent les difficultés rencontrées par les participants, notamment l'ennui, le stress lié à l'absence de musique ou de moyens de communication, et le sentiment d'être "pas au courant des choses". Cependant, des effets positifs sont également observés, comme une plus grande interaction sociale en personne, une augmentation de l'activité physique (marcher davantage), la reprise de la lecture et une meilleure concentration sur le travail scolaire.
La Nécessité d'Apprendre à Se Débrouiller dans le Monde Réel : Le camp de survie de Camille insiste sur l'importance d'acquérir des compétences pratiques et de savoir se débrouiller sans l'aide de la technologie. Il considère qu'il est plus dangereux pour les enfants de grandir sans ces compétences que de passer du temps dans la nature.
Le Sentiment Ambivalent envers les Smartphones : À la fin de l'expérience de détox numérique des lycéens, il est clair que leur relation avec leurs smartphones est complexe. Ils le perçoivent à la fois comme une "malédiction et une bénédiction", conscients de leur dépendance mais aussi de l'utilité et de la fascination de l'appareil.
Idées ou Faits les Plus Importants :
La "génération z est la première à avoir eu un smartphone entre les mains dès le plus jeune âge".
Certains jeunes utilisent leur smartphone jusqu'à "11 12h" par jour.
La déconnexion est une démarche à la mode car "l'addiction au smartphones et aux écrans en général s'est généralisée".
Sébastien Metner, professeur d'histoire, propose de "priver ses élèves de leur drogue pendant une semaine".
Les lycéens commencent l'expérience avec des "sentiments mitigés", mais beaucoup reconnaissent avoir développé une "addiction".
"la plupart d'entre eux n'ont jamais été déconnecté pendant aussi longtemps".
La déconnexion "touche les jeunes de façon existentielle".
Les participants à la détox numérique s'attendent à s'ennuyer, à être stressés par l'absence de musique et à manquer de moyens de communication avec leurs amis.
Ils cherchent des substituts analogiques, comme des "appareils photos jetables", pour "prendre des photos en général on a besoin de souvenirs".
Les lycéens en Allemagne passeraient en moyenne "9h par jour en ligne d'après une étude".
Camille Fialkovski, l'instructeur de survie en Pologne, a une "mission personnelle dans la vie rassembler davantage de personnes autour d'un feu de camp et les éloigner des écrans".
Camille choisit de vivre de manière autonome et de ne pas être "joignable" constamment pour pouvoir se concentrer sur ses tâches.
Les participants à ses stages sont généralement des "citadins stressés qui le sollicitent pour se déconnecter du monde numérique et se ressourcer au grand że przyjeżdansune douza d'anné po iler dans cette tente par mo 10 degrzy".
Les enfants dans le camp de survie apprennent à "faire du feu à grimper aux arbres à pêcher des poissons".
"Ce qui est dangereux pour les enfants c'est pas d'être ici c'est de grandir sans savoir se débrouiller dans la vie".
Le professeur Sébastien Metner est conscient que vivre hors ligne est "devenu quasiment impossible aujourd'hui".
À la fin de la semaine de détox, un élève a reçu "394 messages" non lus.
La "flamme" sur Snapchat est un exemple de la façon dont les jeunes s'investissent émotionnellement dans les interactions en ligne.
Les jeunes "n'arrivent pas à s'en éloigner par eux-mêmes et ils en ont conscience".
Le smartphone est à la fois "une malédiction et une bénédiction".
Points Spécifiques :
Les difficultés pratiques sans smartphone incluent ne pas pouvoir joindre des amis, ne pas être au courant des informations de transport en commun (travaux, trains de substitution), et manquer les annulations de cours via l'application scolaire.
Les effets positifs incluent passer plus de temps à l'extérieur, faire plus d'exercice, lire davantage, et avoir des discussions plus approfondies en personne.
L'expérience de déconnexion a montré à certains participants qu'ils n'étaient "pas aussi dépendants que je le pensais".
La déconnexion a permis à certains de "faire plus de choses pour moi" et de "s'informer sur des sujets qui m'intéressent" au lieu de passer du temps non productif sur leur téléphone.
Le camp de survie de Camille est intentionnellement difficile pour enseigner la résilience, avec un dicton : "ton état d'esprit doit ressembler à ton feu de camp ça veut dire que si tu n'es pas bien préparé tu auras froid".
Ce document offre un aperçu des défis et des opportunités associés à notre dépendance croissante aux technologies numériques, en présentant des exemples concrets d'efforts de déconnexion et leurs conséquences.
Note de Synthèse : Où commence la folie ?
Ce document de synthèse examine les principales thématiques et idées importantes présentées dans les extraits de "Où commence la folie ? | 42 - La réponse à presque tout | ARTE".
Thèmes Principaux :
La Normalité comme Construction Sociale : Le concept de "norme" est présenté non pas comme une réalité intrinsèque mais comme une construction sociale nécessaire à l'interaction. En conséquence, la folie, définie par opposition à la norme, devient également relative.
La Folie au Sens Psychiatrique : La psychiatrie tend à éviter le terme "folie", préférant parler de troubles variés, principalement la psychose. La psychose est caractérisée par une altération du lien avec la réalité et affecte le raisonnement et la perception.
Mécanismes de la Psychose (notamment la Schizophrénie) : Les extraits décrivent comment la psychose, souvent associée à la schizophrénie, peut impliquer l'interprétation de coïncidences comme des signes ou des messages codés. Un sentiment d'insécurité identitaire et environnementale peut conduire à des interprétations délirantes.
La "Bouffée Délirante" : Un épisode intense d'angoisse et d'interprétation délirante, qui paradoxalement peut apporter un soulagement en trouvant une explication (même absurde) à l'insécurité ressentie.
La Folie comme Hyper-sensibilité ou Perception Décalée : L'idée est explorée que les troubles psychiatriques, y compris la psychose, pourraient être des formes amplifiées de caractéristiques présentes chez les personnes dites "normales". Notre perception de la réalité est toujours une interprétation influencée par notre histoire personnelle.
Absence de Frontière Claire entre Folie et Normalité : Il n'existe pas de critères strictement définis pour séparer clairement la folie de la normalité. Le critère principal d'une conviction erronée et inébranlable peut aussi s'appliquer à d'autres formes de croyances non pathologiques.
Le Diagnostic et la Souffrance : Le but du diagnostic en psychiatrie est de soigner les personnes en souffrance. Cependant, la souffrance n'est pas toujours perçue par la personne concernée (ex: épisode maniaque), mais peut être ressentie par l'entourage ou apparaître après l'épisode (souffrance des conséquences). La pertinence d'un diagnostic se mesure plutôt par la souffrance qu'il vise à soulager.
Symptômes Psychotiques chez les Personnes "en Bonne Santé" : Un pourcentage significatif de personnes sans diagnostic formel peuvent présenter des symptômes psychotiques (perceptions différentes, voix, idées bizarres) sans être considérées comme malades.
L'Approche centrée sur le Patient : L'importance de déterminer les objectifs d'un traitement en fonction du patient, et non uniquement du médecin. Un symptôme (comme entendre une voix) peut ne pas être source de souffrance pour le patient et ne pas nécessiter d'être supprimé.
La Neurodiversité : Ce concept remet en question l'idée que certaines particularités, notamment neurodéveloppementales (comme l'autisme), soient des maladies. Elles sont vues comme des expressions naturelles de la diversité humaine. La distinction est faite entre troubles neurodéveloppementaux (avec lesquels on naît et qui ne guérissent pas) et maladies psychiatriques (acquises et potentiellement curables).
L'Évolution du Diagnostic et la Stigmatisation Historique : Les critères diagnostiques ont évolué, menant à une augmentation des diagnostics pour certaines conditions (comme l'autisme). L'histoire montre une persécution et une stigmatisation des personnes jugées différentes, culminant avec les atrocités du régime nazi. La stigmatisation et la discrimination persistent aujourd'hui, notamment dans la sphère professionnelle.
La Folie et la Créativité : Le lien potentiel entre une prédisposition aux troubles psychotiques et une tendance accrue à la créativité est évoqué, citant des études génétiques. Cependant, l'idéalisation de ce lien est tempérée par le fait que les phases aiguës d'une psychose entravent généralement la création.
Le Diagnostic et les Particularités : Certaines particularités (comme le haut potentiel intellectuel) peuvent être confondues avec des troubles (comme le TDAH), soulignant la nécessité d'une évaluation précise pour distinguer une aptitude d'un handicap. Le diagnostic est évolutif et peut enfermer l'individu.
L'Acceptation de la Différence et de sa Propre "Folie" : L'importance pour la société d'accepter les écarts à la norme et pour les individus d'accepter leurs propres singularités et vulnérabilités.
Idées ou Faits Importants :
La norme est une invention sociale et non une réalité absolue.
La "folie" dans le langage courant correspond souvent à ce que la psychiatrie appelle "psychose", caractérisée par une perte du lien avec la réalité.
Un critère clé de la folie est une conviction inébranlable jugée erronée par les autres, mais ce critère peut s'appliquer au-delà des pathologies.
Il n'y a pas de frontière nette entre folie et normalité.
La souffrance est un critère important pour le diagnostic psychiatrique, même si elle n'est pas toujours perçue par le patient lui-même durant l'épisode.
Des symptômes psychotiques peuvent exister chez des personnes sans diagnostic formel.
L'objectif d'un traitement devrait être défini par le patient.
Le concept de neurodiversité voit les particularités (notamment neurodéveloppementales) comme des expressions de la diversité humaine, et non nécessairement comme des maladies.
Les troubles neurodéveloppementaux sont innés, tandis que les maladies psychiatriques sont acquises.
L'histoire a montré une stigmatisation et une persécution des personnes jugées différentes, et cette stigmatisation persiste aujourd'hui.
Il pourrait exister un lien entre la prédisposition aux troubles psychotiques et la créativité.
Le monde peut parfois nous rendre "fous" face aux expériences traumatisantes, soulignant la vulnérabilité humaine.
Accepter la diversité humaine et même sa propre "folie" est crucial.
Citations Clés :
"je pense que la norme en tant que telle elle n'existe pas c'est un concept qu'on a qu'on a inventé qu'on a construit parce qu'on en a besoin pour interagir socialement tous ensemble"
"ce que l'on désigne par folie correspond en général à ce que la psychiatrie appelle une psychose à savoir un état dans lequel tout lien avec la réalité ou du moins ce qu'on perçoit comme tel se brouille"
"lorsque vous vous dites "Je suis en train de donner une interview" vous savez pertinemment qui est le jeu en question pour une personne schizophrène en revanche c'est loin d'être aussi évident"
"il n'existe pas de critères rigoureusement définis qui permettent de tracer une frontière claire entre la folie et la normalité"
"ce qui compte dans le diagnostic des maladies psychiatriques c'est plutôt de savoir si une personne souffre et non pas si elle est atteinte de folie ou si elle a un comportement normal"
"le but d'un traitement doit être déterminé par le patient pas par le médecin"
"en définitive la folie n'est qu'une question de point de vue tout ce qui semble décalé n'a pas forcément besoin d'être soigné"
"Le concept de neurodiversité remet en question l'idée même que certaines particularités soient considérées comme des maladies ou des troubles il les entrevoi plutôt comme une expression naturelle de la diversité humaine"
"la différence entre un trouble neurodéveloppemental et une maladie psychiatrique c'est que la maladie psychiatrique on on l'acquiert comme on se casse une jambe finalement... alors qu'un trouble neuréveloppemental on vient au monde avec et on le garde toute sa vie"
"l'idée reste profondément ancrée dans notre société [la stigmatisation] et que la seule manière d'y faire face ce qui est fondamental notamment pour les personnes atteintes de troubles psychiatriques c'est de sensibiliser le grand public"
"il pourrait exister un lien entre une prédisposition au trouble psychotique et une tendance accrue à la créativité"
"le monde peut parfois nous rendre fou des expériences traumatisantes peuvent conduire certaines personnes à sombrer dans la folie au fond je crois que cela fait partie de la condition humaine"
"peut-être devrions-nous aussi apprendre à mieux accepter notre propre folie"
En résumé, les extraits remettent en question les notions rigides de folie et de normalité, les présentant comme des constructions sociales ou des spectres de perception et de fonctionnement humain.
Ils soulignent la complexité des troubles psychiatriques, l'importance de l'approche centrée sur le patient, et appellent à une plus grande acceptation de la diversité humaine et à la déstigmatisation.
Compte rendu détaillé des thèmes et idées clés du podcast "Les enfants rendent-ils heureux ? | Unhappy | ARTE"
Ce podcast explore la question complexe de l'impact des enfants sur le bonheur parental, en interrogeant différentes perspectives et expériences.
Il s'appuie sur des témoignages personnels, des opinions d'experts et des études socio-économiques.
Thèmes principaux et idées clés :
La parentalité : Un défi enrichissant mais exigeant ("Une vie d'extrémiste")
Le podcast commence par souligner que, bien que les parents puissent parfois préférer des activités solitaires comme regarder la télévision, le temps partagé avec les enfants est souvent source d'enrichissement et de sens.
Le bonheur parental ne réside pas toujours dans des moments de joie continue, mais aussi dans la confrontation et le dépassement des défis inhérents à l'éducation.
Un parallèle est établi entre la parentalité et des défis extrêmes comme l'escalade du Kilimanjaro, mais avec la différence fondamentale qu'on ne peut pas "revenir en arrière". La philosophe d'iter Tomy est citée, affirmant que la parentalité est "réservée aux extrémistes".
Louise, une mère célibataire, témoigne du sentiment d'être constamment dépassée et de la peur de "couler" au début, un mélange de surmenage, d'insomnie, d'angoisse existentielle et de crainte face à la vulnérabilité d'un si petit enfant.
La parentalité est décrite comme une "activité extrêmement intense" qui demande énormément de temps et d'énergie, conduisant à reléguer les propres besoins au second plan.
L'instinct versus la raison dans la décision d'avoir des enfants :
Établir une liste de "pour" et de "contre" pour avoir un enfant est jugé "incongru", notamment en considérant l'impact potentiel sur l'enfant qui grandit et pourrait découvrir qu'il ne "coche pas tous les éléments de la liste".
La parentalité est présentée comme un "mythe redoutable très fortement chargé émotionnellement par la société". Il est conseillé de se demander sincèrement si le désir d'avoir un enfant vient de soi ou est influencé par les "désirs des autres".
Louise, ayant découvert sa grossesse de manière inattendue, témoigne de s'être fiée à son instinct et de n'avoir jamais douté de vouloir garder son enfant malgré les difficultés.
Les aspects du bonheur parental :
Au-delà des défis, les enfants sont sources de bonheur. Louise explique que le bonheur vient en partie de l'acceptation que son fils peut aussi lui apprendre beaucoup de choses et lui a permis de découvrir de nouvelles facettes d'elle-même. Cela aide aussi à "être plus tolérant envers soi-même".
L'attirance innée pour les caractéristiques physiques des bébés (grands yeux, front haut, joues rebondies) est mentionnée comme un facteur initial de sentiment positif.
Tilman, père de quatre filles, affirme que pour lui, le bonheur est indissociable : celui qu'il partage en couple, en famille avec les enfants, et celui qu'il cultive de son côté. C'est un bonheur "avec les enfants", et non "malgré" ou "à cause" d'eux.
Les réalités socio-économiques de la parentalité :
Une étude longitudinale allemande (panel socio-économique) révèle que la satisfaction de vie augmente avant la naissance, atteint un pic la première année, puis diminue et revient au niveau initial. La satisfaction de vie ne diffère pas significativement entre les personnes avec ou sans enfants sur le long terme.
Le coût financier de l'éducation d'un enfant est souligné (environ 150 000 € pour le premier enfant en Allemagne) ainsi que l'impact sur la carrière professionnelle de nombreux parents.
Une augmentation du niveau de colère quotidienne chez les parents 5 ans après la naissance est notée, probablement liée à l'augmentation du stress.
La dynamique du couple et le partage des tâches :
La parentalité transforme la relation de couple en une "sorte de relation de travail", nécessitant une communication constante et une répartition des tâches.
Il est observé que les hommes ont tendance à surestimer leur part de travail ménager par rapport à la réalité.
Le manque de congé parental pris par les pères est souligné (seulement 18% en Allemagne, prenant en moyenne 2 mois contre 13 pour les femmes), indiquant un déséquilibre persistant dans la charge parentale.
Tilman et Ilana, avec quatre enfants, insistent sur l'importance de l'improvisation et de la communication permanente pour concilier famille nombreuse et vies professionnelles.
Les regrets et le tabou de la non-maternité :
La publication de l'étude "Regretting Motherhood" par Orna Donat, interrogeant des femmes qui aiment leurs enfants mais regretteraient de devenir mères si elles pouvaient revenir en arrière, est mentionnée comme ayant brisé un tabou important.
La recherche suggère qu'environ 10% des parents, s'ils pouvaient revenir en arrière, choisiraient de ne pas avoir d'enfants.
Il est souligné que les femmes devraient avoir la liberté de dire que les enfants ne sont pas toujours une bénédiction, une liberté que les hommes possèdent depuis longtemps.
Certains parents peuvent regretter de ne pas avoir consacré autant d'énergie à leur développement personnel après être devenus parents.
Le concept de "village" et la parentalité partagée :
Louise, en tant que mère célibataire, a construit un "village" autour de son fils, un réseau d'amis, de baby-sitters et de connaissances qui la soutiennent. Ce réseau est crucial mais doit être entretenu.
Le podcast suggère d'élargir la définition de la parentalité et de revenir à l'idée qu'il faut "tout un village pour élever un enfant". Il n'est pas nécessaire d'avoir enfanté pour être un "parent" dans ce sens.
L'expérience en Namibie, où les enfants sont confiés à la garde plus tôt, est citée comme une source d'inspiration pour Louise, lui montrant qu'il est possible de concilier parentalité et autres engagements.
La transformation de soi et la responsabilité parentale :
Devenir parent est décrit comme une "seconde naissance" et une "expérience transformatrice", au point de devenir "quelqu'un d'autre". Il est difficile de savoir à l'avance si l'on appréciera ce changement.
Être parent implique d'être "sous surveillance" et de "servir d'exemple", ce qui peut être difficile. Louise confie échouer souvent dans ce rôle, notamment par manque de patience.
La capacité à rire de soi et des autres est présentée comme un outil essentiel pour faire face aux imperfections de la vie de famille.
L'être humain est vu comme un être en devenir, avec un potentiel de transformation, y compris sur le plan émotionnel.
Le détachement et la projection dans l'avenir :
La relation parent-enfant est réciproque en termes d'attachement, et les deux parties doivent éventuellement parvenir à se détacher l'une de l'autre.
Malgré les défis, les enfants permettent aux parents de "voir le monde à travers leurs yeux", de les regarder grandir et devenir autonomes, offrant une "projection dans la société à venir".
Le rôle de l'instinct et du plaisir des parents :
Les parents ne doivent pas se forcer à faire des choses avec leurs enfants s'ils n'en ont pas envie. Le plaisir des parents dans certaines activités est un "acquis précieux pour la vie" des enfants.
Les enfants ont besoin de parents "vraiment là", présents et impliqués, plutôt que de directives données de loin.
Tilman et Ilana soulignent que la vie avec des enfants est "plus riche", "plus imprévisible", les entraîne sur des chemins inattendus, et surtout, qu'ils rient beaucoup en famille, ce qui serait moins le cas sans leurs enfants.
Citations importantes :
"...le bonheur vient notamment du fait de se confronter à des défis que ce soit en élevant un enfant en courant un marathon ou en escaladant une montagne Surmonter ces difficultés enrichit nos vies"
"Je suis d'accord avec le philosophe d'iter Tomy qui a dit un jour que la parentalité était réservée aux extrémistes car contrairement à l'ascension du Kilimanjaro par exemple on ne peut pas revenir en arrière On ne peut pas faire demi-tour On est parent Vitam et Ternam On reste toute sa vie sur le Kilimanjaro quoi qu'il arrive Une vie d'extrémiste"
"...la parentalité est un mythe redoutable très fortement chargé émotionnellement par la société Je conseillerai donc de se poser la question est-ce que je le veux vraiment ou est-ce plutôt le désir des autres que j'essaie de satisfaire"
"J'étais perdu Je n'étais pas sûr de savoir m'y prendre Mais il a 6 ans maintenant Tu as surmonté pas mal de choses déjà Étonnamment Oui Qu'est-ce qui rend heureux dans le fait d'avoir un enfant pour moi c'est d'admettre qu'il puisse lui aussi m'apprendre beaucoup de choses Grâce à lui j'ai découvert de nombreuses facettes de moi que je n'aurais peut-être pas remarqué sans lui On apprend à être plus tolérant envers soi-même Aujourd'hui je me rends compte que cela me rend vraiment heureuse et que c'est un cadeau"
"Une vie d'extrémiste Ce n'est pas vraiment comme ça que j'imaginais la parentalité mais à la réflexion les jeunes parents ont parfois un petit côté fanatique"
"...nos résultats montrent que la satisfaction de vie et le sentiment de bonheur augmente au cours des années précédents la naissance Atteignent un pic au cours de la première année après la naissance puis diminuent et reviennent à leur niveau initial La satisfaction de vie ne diffère pas énormément entre les personnes avec ou sans enfant Celles avec enfant ne sont pas plus heureuse ou plus contente de leur vie que les autres"
"...lorsque en tant qu'être humain nous sommes face à un être avec de grands yeux un front haut et des joues rebondis nous ressentons un sentiment positif Nous nous sentons bien et nous voulons nous en occuper Et ce sont bien sûr exactement les caractéristiques des bébés"
"Quand on devient parent on s'engage dans un travail d'équipe au sein du couple On est dans une sorte de relation de travail et on se critique mutuellement"
"Cette étude a brisé un tabou Les femmes devraient être autorisées à dire que les enfants ne sont pas toujours une bénédiction Cela fait longtemps qu'on accorde aux hommes cette liberté de parole"
"Peut-être devrions-nous élargir la définition de la parentalité et revenir à l'idée qu'il faut tout un village pour élever un enfant Il n'est pas nécessaire d'avoir enfanté pour être parent"
"En devenant parents on devient quelqu'un d'autre et il est difficile de savoir à l'avance si on va apprécier ce changement"
"Pour moi le bonheur ne fait qu'un Celui que nous partageons tous les deux celui que nous vivons en famille avec les enfants et celui que chacun cultive de son côté Je trouve que c'est difficile de les séparer C'est un bonheur qui n'est pas malgré à cause d'eux mais avec les enfants"
"Ce dont les enfants ont besoin ce sont des parents qui sont vraiment là qui ne crient pas de loin 'Rangent ta chambre' mais qui sont présents mettent la main à la pâte et disent 'Vien on va faire ça ensemble.'"
"À une époque où l'on cherche à se prémunir contre tous les risques où l'on cherche à tout sécuriser je trouve tout à fait formidable d'oser se lancer Se jeter ou pas dans le grand bain de la parentalité c'est peut-être au bout du compte aussi une question de confiance"
"la vie devient plus riche elle devient aussi plus imprévisible Cela nous entraîne sur des chemins que l'on aurait pas emprunté sinon Et puis c'est très amusant On rit beaucoup en famille On ne rirait pas autant je pense si on avait pas tous ces enfants autour de nous"
En résumé, le podcast offre une perspective nuancée sur la parentalité, reconnaissant à la fois ses difficultés et ses récompenses.
Il remet en question les mythes sociaux, souligne l'importance du soutien et de l'adaptation, et explore les multiples facettes du bonheur que les enfants peuvent apporter à la vie des parents, malgré les défis inévitables.
Document de Synthèse : Le Nerf Vague
Source: Extraits de "Le nerf vague : dialogue entre le cœur et le cerveau | ARTE"
Introduction :
Ce document explore le rôle multifacette du nerf vague, le présentant comme un "super nerf" essentiel à la communication entre le cerveau et le corps, et à l'adaptation aux changements.
La recherche scientifique actuelle s'intéresse de près à ses capacités, laissant entrevoir un potentiel révolutionnaire en médecine.
Thèmes Principaux et Idées Clés :
Le Rôle Central du Nerf Vague dans l'Adaptation et l'Équilibre du Système Nerveux Autonome:
Après l'épisode stressant, "le système parasympathique prend la relève pour que l'organisme récupère" et "le nerf vague va alors jouer un rôle central" en "apais[ant] le cœur et les poumons et relanc[ant] la digestion".
Le Stress Chronique et l'Inhibition du Nerf Vague:
La Variabilité de la Fréquence Cardiaque comme Indicateur de l'Activité du Nerf Vague:
Potentialités Thérapeutiques de la Stimulation Vagale:
Elle devient une option "lorsque les médicaments ne suffisent pas" ou quand la zone causale ne peut être localisée.
La stimulation vagale peut réduire l'intensité, la fréquence et l'extension des manifestations épileptiformes, améliorant "clairement la qualité de vie" des patients.
La stimulation vagale est également explorée pour traiter d'autres pathologies, notamment la fatigue liée au Covid long, en agissant sur les processus inflammatoires via le nerf vague qui incite la rate à "diminuer sa production de protéines proinflammatoires".
Des études fondamentales explorent l'influence de la stimulation vagale sur la motivation et l'humeur, notamment dans le traitement de la dépression, en influençant "les signaux émis par le corps et la façon dont ils sont traités par le cerveau".
Méthodes de Stimulation Vagale (Médicales et de Bien-être):
La stimulation vagale invasive implique l'implantation chirurgicale d'un générateur à pile connecté au nerf vague au niveau des cervicales.
La stimulation vagale transcutanée, moins invasive, peut être réalisée au niveau du cou et de l'oreille (rameau auriculaire), utilisant des électrodes pour envoyer de faibles impulsions électriques.
L'article met en garde contre la prolifération de stimulateurs vendus dans le domaine du bien-être, soulignant que la plupart n'ont "pas été testés dans des études cliniques" et qu'il est "très problématique de commercialiser un appareil sous le nom de stimulateur vagal s'il n'a pas été prouvé qu'il l'active effectivement".
Entraînement du Nerf Vague par le Mode de Vie:
Le concept de "récipient" qui se vide pendant l'effort et se remplit plus qu'avant pendant la récupération illustre l'adaptabilité accrue. "Pour entraîner son nerf vague ça va aussi vouloir dire aller faire du sport bouger".
La respiration abdominale lente et profonde est une technique efficace pour influencer l'activité du SNA et "venir solliciter le nerf vague en tant qu'allié contre le stress". Il est recommandé que l'expiration dure plus longtemps que l'inspiration.
L'exposition au froid et le réflexe d'immersion sont également mentionnés comme des moyens de provoquer un stress thermique et d'entraîner le nerf vague.
Le chant, en particulier le chant choral, est suggéré comme une activité stimulant le nerf vague par la respiration coordonnée, la vibration du larynx et la libération d'ocytocine.
Le Nerf Vague et la Guérison/Récupération:
Points Importants à Retenir :
Le nerf vague est crucial pour l'équilibre interne et la capacité d'adaptation du corps.
Le stress chronique a un impact négatif majeur sur l'activité du nerf vague.
L'entraînement du nerf vague par l'activité physique, la respiration et d'autres techniques peut améliorer la résilience au stress et la récupération.
La stimulation vagale offre des perspectives thérapeutiques prometteuses pour diverses maladies, mais la recherche est encore en cours pour optimiser son utilisation.
Il est important de distinguer les approches médicales validées des produits de bien-être dont l'efficacité n'est pas toujours prouvée.
Conclusion :
Les mécanismes précis de son influence et ses applications thérapeutiques optimales pour différentes pathologies restent à explorer.
Cependant, les résultats actuels soulignent son importance fondamentale pour la santé physique et mentale et suggèrent des approches concrètes pour améliorer son activité par le mode de vie.
The mammary glands in the inguinalregion are generally larger (for example, the #4 and 5 mam-mary gland pairs in mice) (Fig. 2a). The precocial type specieshave nipples developed in the inguinal region (Fig. 2b). Thelimited number of offspring produced by these animals meansthere is not a need for a great number of nipples whichessentially serve as points of milk outlet. The location wherenipples are developed in the precocial type occurs in theregion where the mammary gland appears to have the capacityto develop to a maximum size. This extensive glandular tissuein ungulates provides the rather large and active offspring ofthese animals with abundant milk, and this capacity has beenfurther expanded in domesticated species.
Inguinal nipples allow for larger mammary glands and this is why they have migrated down.
Seulement voilà, tout en démocratisant la prise de parole, ces circuits conversationnels de l'information peuvent être exploités par certains producteurs de fausses nouvelles. Enrobées de blagues ou de commentaires affectifs, certaines informations graves ou frauduleuses peuvent en effet s'immiscer, aussi insidieusement qu'un cheval de Troie, dans ces niches conversationnelles.
Ici, l'auteure en parlant de "cheval de Troie" alerte sur les producteurs de fake news pour galvaniser de la fausse information dans des conversations à hauts potentiels de visibilités...et rendre exponentielle la visibilité de leurs fausses informations, pour manipuler le public, ou tout du moins faire parler du sujet souhaité.
Or cette nouvelle structure de la visibilité a ainsi favorisé l'apparition de nouveaux registres d'énonciation plus familiers et désinhibés.
Ici l'auteure conclue que l'information est désinhibée, familière...et ne respecte plus les codes entre place publique et institutionnelle que l'on pouvait observer autrefois, étant donné que les réseaux sociaux font le pont entre ces deux mondes. Nous l'avons notamment vu à travers les résultats de cette étude, l'auteure pourrait alerter sur les dérives liées à de tels comportements : s'ils se multiplient et disposent de plus en plus de visibilité...cela pourrait normaliser des discours qui ne respectent pas l'intégrité de l'autre et où la désinformation pourrait être monnaie courante.
Elles ne sont pas plus murmurées, dans des espaces privés, au sein de contextes de communication interpersonnelle, mais désormais exposées au sein d'espaces de visibilité en « clair-obscur » . Dominique Cardon utilise cette expression pour souligner que, pour autant qu'ils soient visibles, ces bavardages sont remplis de sous-entendus et d'indices complices destinés à n'être compréhensibles que pour un cercle restreint : les proches du réseau relationnel des internautes.au sein d'espaces de visibilité frDominique Cardon utilise cette expression pour souligner que, pour autant qu'ils soient visibles,ces bavardages sont remplis de sous-entendus et d'indices complices destinés à n'être compréhensibles que pour un cercle restreint : les proches du réseau relationnel des internautes.
Ici l'auteure évoque une communication sur la sphère publique que le sociologue Dominique Cardon qualifie de "clair-obscur", à savoir des propos assumés, clairement affichés mais peu lisibles par les lecteurs "hors cercle" puisqu'on y trouve seulement des sous-entendus. Pour autant, même sans être clairs pour les autres lecteurs, de tels propos, sous couverts de faire réagir pourraient avoir un impact ou une interprétation négative et participer à cette spirale du "bad buzz", de la polémique, à échelle du public évoquée plus haut.
Si le phénomène des « fake news » n’est pas si nouveau et ravive d’une certaine manière les ragots et commérages qui se transmettaient par bouche à oreille dans les coulisses d’un café, sa grande nouveauté aujourd’hui est qu’il peut projeter certains racontages douteux sur le devant de la scène, au sein d’espace à haute visibilité du web comme les groupes Facebook ou les fils de discussions Twitter.
Ici, l'autrice assimile les fausses informations que l'on observe aux discussions d'autrefois, mais en nuançant leur impact, qui est exponentiel aujourd'hui par l'essor des réseaux sociaux. On pourrait faire le parallèle avec le "badbuzz" phénomène qui privilégie les nouvelles catastrophes, à sensation, plus vendeuses que les nouvelles plus positives, se demander si tout cela n'est pas un engrenage par les outils dont nous disposons mais finalement un comportement humain caractéristique.
« C’est très drôle. À mon avis c’est une fake news mais c’est drôle, ça me fait rire. Que ça soit vrai ou faux je m’en fous, j’en parle. » (Homme, 67 ans, retraité)
Ici l'expérience et ses conclusions sont appuyées par les propos des participants, où l'on voit que l'importance de la véracité de l'information n'est pas un élément priorisé. On aurait pu pousser le questionnement au "pourquoi" ce besoin de faire rire, dans le rapport à l'autre.
Probablement parce dans ces contextes, l’on ne risque pas grand-chose à diffuser quelque chose de faux. Et puis surtout parce que l’on ne se préoccupe pas vraiment de la valeur de vérité d’une information car nos conversations sont animées par d’autres motivations et s’apparentent alors davantage à des bavardages cacophoniques mobilisant des registres d’énonciation divers et variés oscillant par exemple de la plaisanterie à la provocation :
à nouveau ici, face à la naïveté que peuvent avoir certaines personnes à diffuser de fausses information pour divertir, dans un contexte sécurisant pour eux, il pourrait être opportun d'alerter sur la responsabilité de chacun à transmettre des informations, quelqu'en soit le contexte.
En fait, les informations fausses et sans intérêt public ont surtout été transmises au sein d’espace de communication aux contraintes de prise de parole très relâchées, par exemple à un·e ami·e très proche qui a la même opinion que nous sur de nombreux sujets ou dans un groupe de conversation dans lequel notre identité peut rester masquée. Pourquoi ?
l'autrice lie les fake news & la typologie d'espaces de communication. Elle pose la question de la plus grande diffusion de fausses informations dans un cadre moins institutionnel et plus relâché ou avec une identité masquée. Il aurait pu être intéressant de redéfinir la différence entre ces espaces privés et public. Egalement, appuyer les propos sur m'effet levier auquel le public participe en relatant de genre de désinformation, et l'impact que cela peut avoir, pour chaque contextes.
Pour explorer cette question, j’ai réalisé une enquête expérimentale dans le cadre d’un travail exploratoire auprès de 15 personnes, pour mon mémoire de fin d’études à Sciences Po, encadré par le sociologue Dominique Cardon. Ce questionnement est aujourd’hui approfondi dans mes recherches doctorales.
après avoir démontré ce que la désinformation n'est pas seulement le fruit d'une diffusion massive de fake news, mais que le contexte de discussion est un facteur important à prendre en compte, l'auteure continue sa démonstration par une expérience. cependant, il est à noter que le nombre de participant de 15 personnes est faible et il n'y a pas d'indication concernant la sélection de cet échantillon.
Ha indicado que el análisis interno realizado en estas semanas apunta a la necesidad de “profun
Usar siempre
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
Reviewer #1 (Evidence, reproducibility and clarity (Required)):
In this manuscript the authors have done cryo-electron tomography of the manchette, a microtubule-based structure important for proper sperm head formation during spermatogenesis. They also did mass-spectrometry of the isolated structures. Vesicles, actin and their linkers to microtubules within the structure are shown.
__We thank the reviewer for the critical reading of our manuscript; we have implemented the suggestions as detailed below, which we believe indeed improved the manuscript. __
Major:
The data the conclusions are based on seem very limited and sometimes overinterpreted. For example, only one connection between actin and microtubules was observed, and this is thought to be MACF1 simply based on its presence in the MS.
__We regret giving the impression that the data is limited. We in fact collected >100 tilt series from 3 biological replicas for the isolated manchette. __
__In the revised version, we added data from in-situ studies showing vesicles interacting with the manchette (as requested below, new Fig. 1). __
Specifically, for the interaction of actin with microtubule we added more examples (Revised Fig. 6) and we toned down the discussion related to the relevance of this interaction (lines 193-194, 253-255). MACF1 is mentioned only as a possible candidate in the discussion (line 254).
Another, and larger concern, is that the authors do a structural study on something that has been purified out of the cell, a process which is extremely disruptive. Vesicles, actin and other cellular components could easily be trapped in this cytoskeletal sieve during the purification process and as such, not be bona fide manchette components. This could create both misleading proteomics and imaging. Therefore, an approach not requiring extraction such as high-pressure freezing, sectioning and room-temperature electron tomography and/or immunoEM on sections to set aside this concern is strongly recommended. As an additional bonus, it would show if the vesicles containing ATP synthase are deformed mitochondria.
__We recognise the concern raised by the reviewer. __
__To alleviate this concern, we added imaging data of manchettes in-situ that show vesicles, mitochondria and filaments interacting with the manchette (new Fig. 1), essentially confirming the observations that were made on the isolated manchette. __
__The benefits of imaging the isolated manchette were better throughput (being able to collect more data) and reaching higher resolution allowing to resolve unequivocally the dynein/dynactin and actin filaments. __
Minor: Line 99: "to study IMT with cryo-ET, manchettes were isolated ...(insert from which organism)..."
__Added in line 102 in the revised version. __
Line 102 "...demonstrating that they can be used to study IMT".. can the authors please clarify?
This paragraph was revised (lines 131-137), we hope it is now more clear.
Line 111 "densities face towards the MT plus-end" How can a density "face" anywhere? For this, it needs to have a defined front and back.
Microtubule motor proteins (kinesin and dynein) are often attached to the microtubules with an angle and dynactin and cargo on one side (plus end). We rephrased this part and removed the word “face” in the revised version to make it more clear (lines 161-162).
Line 137: is the "perinuclear ring" the same as the manchette?
The perinuclear ring is the apical part of the manchette that connects it to the nucleus. We added to the revised version imaging of the perinuclear ring with observations on how it changes when the manchette elongates (new Fig. 2).
Figure 2B: How did the authors decide not to model the electron density found between the vesicle and the MT at 3 O'clock? Is there no other proteins with a similar lollipop structure as ATP synthase, so that this can be said to be this protein with such certainty?
__The densities connecting the vesicles to the microtubules shown in (now) Fig. 4D are not consistent enough to be averaged. __
__The densities resembling ATP synthase are inside the vesicles. Nevertheless, we have decided to remove the averaging of the ATP synthases from the revised manuscipt as they are not of great importance for this manuscript. Instead, the new in-situ data clearly show mitochondria (with their characteristic double membrane and cristae) interacting with manchette microtubule (new Fig 1C). __
Line 189: "F-actin formed organized bundles running parallel to mMTs" - this observation needs confirming in a less disrupted sample.
__Phalloidin (actin marker) was shown before to stain the manchette (PMID: 36734600). As actin filaments are very thin (7 nm) they are very hard to observe in plastic embedded EM. __
In the in-situ data we added to the revised manuscipt (new Fig 1D), we observe filaments with a diameter corresponding to actin. In addition, we added more examples of microtubules interacting with actin in isolated manchette (new Fig. 6 E-K).
Line 242 remove first comma sign.
Removed.
Line 363 "a total of 2 datasets" - is this manuscript based on only two tilt-series? Or two datasets from each of the 4 grids? In any case, this is very limited data.
We apologise for not clearly providing the information about the data size in the original manuscipt. The data is based on three biological replicas (3 animals). We collected more than 100 tomograms of different regions of the manchettes. As such, we would argue that the data is not limited per se.
Reviewer #1 (Significance (Required)):
The article is very interesting, and if presented together with the suggested controls, would be informative to both microtubule/motorprotein researchers as well as those trying studying spermatogenesis.
Reviewer #2 (Evidence, reproducibility and clarity (Required)):
The manchette appears as a shield-like structure surrounding the flagellar basal body upon spermiogenesis. It consists of a number of microtubules like a comb, but actin (Mochida et al. 1998 Dev. Biol. 200, 46) and myosin (Hayasaka et al. 2008 Asian J. Androl. 10, 561) were found, suggesting transportation inside the manchette. Detailed structural information and functional insight into the manchette was still awaited. There is a hypothesis called IMT (intra-machette transport) based on the fact that machette and IFT (intraflagellar transport) share common components (or homologues) and on their transition along the stages of spermiogenesis. While IMT is considered as a potential hypothesis to explain delivery of centrosomal and flagellar components, no one has witnessed IMT at the same level as IFT. IMT has never been purified, visualized in motion or at high resolution. This study for the first time visualized manchette using high-end cryo-electron tomography of isolated manchettes, addressing structural characterization of IMT. The authors successfully microtubular bundles, vesicles located between microtubules and a linker-like structure connecting the vesicle and the microtubule. On multilamellar membranes in the vesicles they found particles and assigned them to ATPase complexes, based on intermediate (~60A) resolution structure. They further identified interesting structures, such as (1) particles on microtubules, which resemble dynein and (2) filaments which shows symmetry of F-actin. All the molecular assignments are consistent with their proteomics of manchettes.
__We thank the reviewer for highlighting the novelty of our study.____ __
Their assignment of ATPase will be strengthened by MS data, if it proves absence of other possible proteins forming such a membrane protein complex.
All the ATPase components were indeed found in our proteomics data. Nevertheless, we have decided to remove the averaging of the ATPase as it does not directly relate to IMT, the focus of this manuscript.
They discussed possible role of various motor proteins based on their abundance (Line 134-151, Line 200). This makes sense only with a control. Absolute abundance of proteins would not necessarily present their local importance or roles. This reviewer would suggest quantitative proteomics of other organelles, or whole cells, or other fractions obtained during manchette isolation, to demonstrate unique abundance of KIF27 and other proteins of their interest.
We agree with the reviewer that absolute abundance does not necessarily indicate importance or a role. As such, we removed this part of the discussion from the revised manuscript.
A single image from a tomogram, Fig.6B, is not enough to prove actin-MT interaction. A gallery and a number (how many such junctions were found from how many MTs) will be necessary.
We agree that one example is not enough. In the new Fig. 6E-K, we provide a gallery of more examples. We have revised the text to reflect the point that these observations are still rare and more data will be needed to quantify this interaction (Lines 253-254).
Minor points: Their manchette purification is based on Mochida et al., which showed (their Fig.2) similarity to the in vivo structure (for example, Fig.1 of Kierszenbaum 2001 Mol. Reproduc. Dev. 59, 347). Nevertheless, since this is not a very common prep, it is helpful to show the isolated manchette’s wide view (low mag cryo-EM or ET) to prove its intactness.
We thank the reviewer for this suggestion, in the revised version, new Fig. 2 provides a cryo-EM overview of purified manchette from different developmental stages.
Line 81: Myosin -> myosin (to be consistent with other protein names)
Corrected.
This work is a significant step toward the understanding of manchettes. While the molecular assignment of dynein and ATPase is not fully decisive, due to limitation of resolution (this reviewer thinks the assignment of actin filament is convincing, based on its helical symmetry), their speculative model still deserves publication.
Reviewer #2 (Significance (Required)):
This work is a significant step toward the understanding of manchettes. While the molecular assignment of dynein and ATPase is not fully decisive, due to limitation of resolution (this reviewer thinks the assignment of actin filament is convincing, based on its helical symmetry), their speculative model still deserves publication.
Reviewer #3 (Evidence, reproducibility and clarity (Required)):
->Summary:
The manchette is a temporary microtubule (MT)-based structure essential for the development of the highly polarised sperm cell. In this study, the authors employed cryo-electron tomography (cryo-ET) and proteomics to investigate the intra-manchette transport system. Cryo-EM analysis of purified rat manchette revealed a high density of MTs interspersed with actin filaments, which appeared either bundled or as single filaments. Vesicles were observed among the MTs, connected by stick-like densities that, based on their orientation relative to MT polarity, were inferred to be kinesins. Subtomogram averaging (STA) confirmed the presence of dynein motor proteins. Proteomic analysis further validated the presence of dynein and kinesins and showed the presence of actin crosslinkers that could bundle actin filaments. Proteomics data also indicated the involvement of actin-based transport mediated by myosin. Importantly, the data indicated that the intraflagellar transport (IFT) system is not part of the intra-manchette transport mechanism. The visualisation of motor proteins directly from a biological sample represents a notable technical advancement, providing new insights into the organisation of the intra-manchette transport system in developing sperm.
We thank the reviewer for summarising the novelty of our observations.
-> Are the key conclusions convincing? Below we comment on three main conclusions. MT and F-actin bundles are both constituents of the manchette While the data convincingly shows that MT and F-actin are part of the manchette, one cannot conclude from it that F-actin is an integral part of the manchette. The authors would need to rephrase so that it is clear that they are speculating.
We have rephrased our statements and replaced “integral” with ‘actin filaments are associated’. Of note previous studies suggested actin are part of the manchette including staining with phalloidin (PMID: 36734600, PMID: 9698455, PMID: 18478159) and we here visualised the actin in high resolution.
The transport system employs different transport machinery on these MTs Proteomics data indicates the presence of multiple motor proteins in the manchette, while cryo-EM data corroborates this by revealing morphologically distinct densities associated with the MTs. However, the nature of only one of these MT-associated densities has been confirmed-specifically, dynein, as identified through STA. The presence of kinesin or myosin in the EM data remains unconfirmed based on just the cryo-ET density, and therefore it is unclear whether these proteins are actively involved in cargo transport, as this cannot be supported by just the proteomics data. In summary, we recommend that the authors rephrase this conclusion and avoid using the term "employ".
We agree that our cryo-ET only confirmed the motor protein dynein. As such, we removed the term employ and rephrased our claims regarding the active transport and accordingly changed the title.
Dynein mediated transport (Line 225-227) The data shows that dynein is present in the manchette; however, whether it plays and active role in transport cannot be determined from the cryo-ET data provided in the manuscript, as it does not clearly display a dynein-dynactin complex attached to cargo. The attachment to cargo is also not revealed via proteomics as no adaptor proteins that link dynein-dynactin to its cargo have been shown.
A list of cargo adaptor proteins were found in our proteomics data but we agree that cryo-ET and proteomics alone cannot prove active transport. As such we toned down the discussion about active transport (lines 212-220).
-> Should the authors qualify some of their claims as preliminary or speculative, or remove them altogether?
F-actin • In the abstract, the authors state that F-actin provides tracks for transport as well as having structural and mechanical roles. However, the manuscript does not include experiments demonstrating a mechanical role. The authors appear to base this statement on literature where actin bundles have been shown to play a mechanical role in other model systems. We suggest they clarify that the mechanical role the authors suggest is speculative and add references if appropriate.
__ ____We removed the claim about the mechanical role of the actin from the abstract and rephrased this in the discussion to suggest this role for the F-actin (lines 242-243).__
• Lines 15,92, 180 and 255: The statement "Filamentous actin is an integral part of the manchette" is misleading. While the authors show that F-actin is present in their purified manchette structures, whether it is integral has not been tested. Authors should rephrase the sentence.
We removed the word integral.
• To support the claim that F-actin plays a role in transport within the manchette, the authors present only one instance where an unidentified density is attached to an actin filament. This is insufficient evidence to claim that it is myosin actively transporting cargo. Although the proteomics data show the presence of myosin, we suggest the authors exercise more caution with this claim.
We agree that our data do not demonstrate active transport as such we removed that claim. We mention the possibility of cargo transport in the discussion (lines 250-255).
• The authors mention the presence of F-actin bundles but do not show direct crosslinking between the F-actin filaments. They could in principle just be closely packed F-actin filaments that are not necessarily linked, so the term "bundle" should be used more cautiously.
We do not assume that a bundle means that the F-actin filaments are crosslinked. A bundle simply indicates the presence of multiple F-actin filaments together. We rephrased it to call them actin clusters.
Observations of dynein • Relating to Figure 2B: From the provided image it is not clear whether the density corresponds to a dynein complex, as it does not exhibit the characteristic morphological features of dynein or dynactin molecules.
We indeed do not claim that the densities in this figure are dynein or dynactin. __We revised this paragraph and hope that it is now more clear (lines 135-137). __
• Lines 171-172 and Figure 4: It is well established that dynein is a dimer and should always possess two motor domains. The authors have incorrectly assumed they observed single motor heads, except possibly in Figure 4A (marked by an arrow). In all other instances, the dynein complexes show two motor domains in proximity, but these have not been segmented accurately. Furthermore, the "cargos" shown in grey are more likely to represent dynein tails or the dynactin molecule, based on comparisons with in vitro structures of these complexes (see references 1-3).
We thank the reviewer for this correction. We improved the annotations in the figure and revised the text to clarify that we identified dimers of dynein motor heads (lines 140-144). We further added a projection of a dynein dynactin complex to compare to the observation on the manchette (new Fig. 5E). We further changed claims on the presence of protein cargo to the presence of dynein/dynactin that allows cargo tethering based on the presence of cargo adaptors in the proteomics data.
• Lines 21, 173, and 233 mention cargos, but as noted above, it seems to be parts of the dynein complex the authors are referring to.
This was corrected as mentioned above.
• Panel 4B appears to show a dynein-dynactin complex, but whether there is a cargo is unclear and if there is it should be labelled accordingly. To assessment of whether there is any cargo bound to the dynein-dynactin complex a larger crop of the panel would be helpful In summary, we recommend that the authors revisit their segmentations in Figures 2B and 4, revise their text based on these observations, and perform quantification of the data (as suggested in the next section).
We thank the reviewers for sharing their expertise on dynein-dynactin complexes. We have revised the text as detailed above and excluded the assignment of any cargo, as we cannot (even from larger panels) see a clear association of cargo. We have made clear that we only refer to dynein dynactin with the capability of linking cargo based on the presence of proteomics data. We have removed claims on active transport with dynein.
Dynein versus kinesin-based transport The calculation presented in lines 147-151 does not account for the fact that both the dynein-dynactin complex and kinesin proteins require cargo adaptors to transport cargo. Additionally, the authors overlook the possibility that multiple motors could be attached to a single cargo. If the authors did not observe this, they should explicitly mention it to support their argument. In short, the calculations are based on an incorrect premise, rendering the comparison inaccurate. Unless the authors have identified any dynein-dynactin or kinesin cargo adaptors in their proteomics data which could be used for such a comparison, we believe the authors lack sufficient data to accurately estimate the "active transport ratio" between dynein and kinesin.
Even though we detect cargo adaptors in our proteomics, we agree that calculating relative transport based only on the proteomics can be inaccurate as such we removed absolute quantification and comparison between dynein and kinesin-based IMT.
- Would additional experiments be essential to support the claims of the paper?
F-actin distance and length distribution • To support the claim that F-actin is bundled (line 189), could the authors provide the distance between each F-actin filament and its neighbours? Additionally, could they compare the average distance to the length of actin crosslinkers found in their proteomics data, or compare it to the distances between crosslinked F-actin observed in other research studies?
We measured distances between the actin filaments and added a plot to new Fig 6.
• While showing that F-actin is important for the manchette would require cellular experiments, authors could provide quantification of how frequently these actin structures are observed in comparison to MTs to support their claims that these actin filaments could be important for the manchette structure.
We agree that claims on the role and function of actin in the manchette require cellular experiments that are beyond the scope of this study. Absolute quantification of the ratio between MTs and actin from cryoET is very hard and will be inaccurate as the manchette cannot be imaged as a whole due to its size and thickness. The ratio we have is based on the relative abundance provided by the proteomics (Fig. 5F).
• In line 193, the authors claim that the F-actin in bundles appears too short for transport. Could they provide length distributions for these filaments? This might provide further support to their claim that individual F-actin filaments can serve as transport tracks (line 266).
__In addition to the limitation mentioned in the previous point, quantification of length from high magnification imaging will likely be inaccurate as the length of the actin in most cases is bigger than the field of view that is captured. Nevertheless, we removed the claim about the actin being too short for transport. __
• Could the authors also quantify the abundance of individual F-actin filaments observed, compared to MTs and F-actin bundles, to support the idea that they could play a role in transport?
As explained for the above points absolute quantification of the ratio between MTs and actin is not feasible from cryoET data that cannot capture all of the manchette in high enough resolution to resolve the actin.
• In the discussion, the authors mention "interactions between F-actin singlets and mMTs" (line 269), yet they report observing only one instance of this interaction (lines 210 and 211). Given the limited data, they should refer to this as a single interaction in the discussion. The scarcity of data raises questions about how representative this event truly is.
We agree that one example is not enough. In the new Fig. 6E-K, we provide a gallery of more examples as also requested by reviewers 1 and 2. We have also revised the text to reflect the point that these observations are still rare (Lines 190-194).
Quantifications for judgement of representativity The authors should quantify how often they observed vesicles with a stick-like connection to MTs (lines 106-107); this would strengthen the interpretation of the density, as currently only one example is shown in the manuscript (Figure 4A). If possible, they could show how many of them are facing towards the MT plus end.
__As mentioned in the text (lines 135-137), the linkers connecting vesicles to MTs were irregular and so we could not interpret them further this is in contrast to dynein that were easily recognisable but were not associated with vesicles. __
Dynein quantifications • The authors are recommended to quantify how many dynein molecules per micron of MT they observe and how often they are angled with their MT binding domain towards the minus-end.
As the manchette is large and highly dense any quantification will likely be biased towards parts of the manchette that are easier to image, for example the periphery. As such we do not think quantifying the dynein density will yield meaningful insight.
• Could the authors quantify how many dynein densities they found to be attached to a (vesicle) cargo, if any (line 175)? They could show these observations in a supplementary figure.
We did not observe any case of a connection between a vesicle and dynein motors, we edited this sentence to be more clear on that.
• For densities that match the size and location of dynein but lack clear dynein morphology (as seen in Figure 2B), could the authors quantify how many are oriented towards the MT minus end?
We had many cases where the connection did not have a clear dynein morphology, and as the morphology is not clear, it is impossible to make a claim about whether they are oriented towards the minus end.
Artefacts due to purification: Authors should discuss if the purification could have effects on visualizing components of the manchette. For example, if it has effect on the MTs and actin structure or the abundance/structure of the motor protein complexes (bound to cargo or isolated).
We have followed a protocol that was published before and showed the overall integrity of the manchette. Nevertheless, losing connections between manchette and other cellular organelles are expected. To address this point, we added in-situ data (new Fig 1) showing manchette in intact spermatids interacting with vesicles and mitochondria, as well as overviews of manchettes (new Fig 2), the text was revised accordingly.
- Are the experiments adequately replicated and statistical analysis adequate? The cryo-ET data presented in the manuscript is collected using two separate sample preparations. Along with the quantifications of the different observations suggested above which will help the reader assess how abundant and representative these observations are, the authors could further strengthen their claims by acquiring data from a third sample preparation and then analysing how consistent their observations are between different purifications. This however could be time consuming so it is not a major requirement but recommended if possible within a short time frame.
We regret not explicitly mentioning our data set size, it was added now to the revised version. In essence, the data is based on three biological replicas (3 animals). We collected more than 100 tomograms of different regions of the manchettes. We provided in the revised version more observations (new Fig 1, 2, 4B-C and 6E-K).
- Are the suggested experiments realistic in terms of time and resources? It would help if you could add an estimated cost and time investment for substantial experiments.
Most of the comments deal with either modifying the text or analysing the data already presented, so the revision could be done with 1-3 months.
Minor comments: - Specific experimental issues that are easily addressable. 1) Could the authors state how many tilt series were collected for each dataset/independent sample preparation? We recommend that they upload their raw data or tomograms to EMPAIR.
We added this information in the material and methods.
2) It is not clear to me if the same sample was used for cryo-ET and proteomics. Could the authors clarify how comparable the sample preparation for the cryo-ET and proteomics data is or if the same sample was used for both. If there is a discrepancy between these preparations, they would need to discuss how this can affect comparing observations from cryo-ET and mass spectrometry. Ideally both samples should be the same.
After sample preparation the manchettes were directly frozen on grids. The rest of the samples was used for proteomics. Consequently, EM and MS data were acquired on the same samples. We clarified this in the text (lines 327-328).
- Are prior studies referenced appropriately? We recommend including additional references to support the claim that F-actin has a mechanical role (line 242). Could the authors compare their proteomics data to other mass spectrometry studies conducted on the Manchette (for example, see reference 4)?
We added the comparison but it is important to point out that in reference 4 the manchettes were isolated from mice testes.
- Are the text and figures clear and accurate? Text: We do not see the necessity of specifying the microtubules (MTs) in the data as "manchette MTs" or "mMTs" rather than simply "MTs". However, we recommend that the authors use either "MT" or "mMT" consistently throughout the manuscript.
We changed to only MTs.
The authors appear to refer to both dynein-1 (cytoplasmic dynein) and dynein-2 (axonemal dynein or IFT dynein). To avoid confusion, it is important that the authors clearly specify which dynein they are referring to throughout the text. This is particularly relevant as the study aims to demonstrate that IFT is not part of the manchette transport system.
• Introduction: In the third paragraph (lines 59-75), the authors should specify that they are referring to dynein-2, which is distinct from cytoplasmic dynein discussed in the previous paragraph (lines 44-58).
We specify the respective dyneins in the text (line 66,140-141,145).
• Figure 4D: The authors could fit a dynein-1 motor domain instead of a dynein-2 into the density to stay consistent with the fact that the density belongs to cytoplasmic dynein-1.
__We changed the figure and fitted a cytosolic dynein-1 structure (5nvu) instead. __
Figures: • Figure 2B: The legend mentions a large linker complex; however, this may correspond to two or three separate densities.
We have addressed this and changed the wording.
• Figure 4: please revisit the segmentation of this whole figure based on previous comments.
__We revised as suggested. __
• Figures 1, 2, 4, 5, and 6: It would be helpful to state in the legends that the tomograms are denoised. There are stripe-like densities visible in the images (e.g., in the vesicle in Figure 2B). Do these artefacts also appear in the raw data?
As stated in the Methods section, tomograms were generally denoised with CryoCare for visualisation purposes. The “stripe-like densities” are artefacts of the gold fiducials used for tomogram alignment and appear in the raw data (before denoising).
- Do you have suggestions that would help the authors improve the presentation of their data and conclusions? We suggest revising the paragraph title "Dynein-mediated cargo along the manchette" (line 165) to "Dynein-mediated cargo transport along the manchette".
__We have changed this in the revised version. __
We recommend that the authors provide additional evidence to support the interpretation that the observed EM densities correspond to motor proteins. Specifically: • Include scale bars or reference lines indicating the known dimensions of motor proteins, based on previous data, to demonstrate that the observed densities match the expected size.
The dynein structure is provided for reference. We also added the cytosolic dynein–dynactin as a reference (Fig 5E).
• Make direct comparisons to existing EM data and highlight morphological similarities.
We have added a comparison to existing data (Fig 5E).
In the discussion (lines 249-254), the authors could speculate on alternative roles for the IFT components in the manchette, particularly if they are not part of the IFT trains. We also suggest rephrasing the claim in line 266 to make it more speculative in tone.
__We have addressed this in the revised version (lines 221-230). __
Finally, a schematic overview of the manchette ultrastructure in a spermatid would greatly aid the reader in understanding the material presented.
We now include a graphical abstract and overviews of isolated manchettes on cryo-EM grids.
References: 1. Chowdhury, S., Ketcham, S., Schroer, T. et al. Structural organization of the dynein-dynactin complex bound to microtubules. Nat Struct Mol Biol 22, 345-347 (2015). https://doi.org/10.1038/nsmb.2996
Grotjahn, D.A., Chowdhury, S., Xu, Y. et al. Cryo-electron tomography reveals that dynactin recruits a team of dyneins for processive motility. Nat Struct Mol Biol 25, 203-207 (2018). https://doi.org/10.1038/s41594-018-0027-7
Chaaban, S., Carter, A.P. Structure of dynein-dynactin on microtubules shows tandem adaptor binding. Nature 610, 212-216 (2022).https://doi.org/10.1038/s41586-022-05186-y
W. Hu, R. Zhang, H. Xu, Y. Li, X. Yang, Z. Zhou, X. Huang, Y. Wang, W. Ji, F. Gao, W. Meng, CAMSAP1 role in orchestrating structure and dynamics of manchette microtubule minus-ends impacts male fertility during spermiogenesis, Proc. Natl. Acad. Sci. U.S.A. 120 (45) e2313787120, https://doi.org/10.1073/pnas.2313787120 (2023).
Reviewer #3 (Significance (Required)):
This study employs cryo-electron tomography (cryo-ET) and proteomics to elucidate the architecture of the manchette. It advances our understanding of the components involved in intracellular transport within the manchette and introduces the following technical and conceptual innovations:
a) Technical Advances: The authors have visualized the manchette at high resolution using cryo-ET. They optimized a purification pipeline capable of retaining, at least partially, the transport machinery of the manchette. Notably, they observed dynein and putative kinesin motors attached to microtubules-a significant achievement that, to our knowledge, has not been reported previously.
b) Conceptual Advances: This study provides novel insights into spermatogenesis. The findings suggest that intraflagellar transport (IFT) is unlikely to play a role at this stage of sperm development while shedding light on alternative transport systems. Importantly, the authors demonstrate that actin filaments organize in two distinct ways: clustering parallel to microtubules or forming single filaments.
This work is likely to be of considerable interest to researchers in sperm development and structural biology. Additionally, it may appeal to scientists studying motor proteins and the cytoskeleton.
We thank the reviewers for appreciating the significance and novelty of our study.
The reviewers possess extensive expertise in in situ cryo-electron tomography and single-particle microscopy, including work on dynein-based complexes. Collectively, they have significant experience in the field of cytoskeleton-based transport.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary:
The manchette is a temporary microtubule (MT)-based structure essential for the development of the highly polarised sperm cell. In this study, the authors employed cryo-electron tomography (cryo-ET) and proteomics to investigate the intra-manchette transport system. Cryo-EM analysis of purified rat manchette revealed a high density of MTs interspersed with actin filaments, which appeared either bundled or as single filaments. Vesicles were observed among the MTs, connected by stick-like densities that, based on their orientation relative to MT polarity, were inferred to be kinesins. Subtomogram averaging (STA) confirmed the presence of dynein motor proteins. Proteomic analysis further validated the presence of dynein and kinesins and showed the presence of actin crosslinkers that could bundle actin filaments. Proteomics data also indicated the involvement of actin-based transport mediated by myosin. Importantly, the data indicated that the intraflagellar transport (IFT) system is not part of the intra-manchette transport mechanism. The visualisation of motor proteins directly from a biological sample represents a notable technical advancement, providing new insights into the organisation of the intra-manchette transport system in developing sperm.
Are the key conclusions convincing?
Below we comment on three main conclusions.
MT and F-actin bundles are both constituents of the manchette While the data convincingly shows that MT and F-actin are part of the manchette, one cannot conclude from it that F-actin is an integral part of the manchette. The authors would need to rephrase so that it is clear that they are speculating.
The transport system employs different transport machinery on these MTs Proteomics data indicates the presence of multiple motor proteins in the manchette, while cryo-EM data corroborates this by revealing morphologically distinct densities associated with the MTs. However, the nature of only one of these MT-associated densities has been confirmed-specifically, dynein, as identified through STA. The presence of kinesin or myosin in the EM data remains unconfirmed based on just the cryo-ET density, and therefore it is unclear whether these proteins are actively involved in cargo transport, as this cannot be supported by just the proteomics data. In summary, we recommend that the authors rephrase this conclusion and avoid using the term "employ".
Dynein mediated transport (Line 225-227) The data shows that dynein is present in the manchette; however, whether it plays and active role in transport cannot be determined from the cryo-ET data provided in the manuscript, as it does not clearly display a dynein-dynactin complex attached to cargo. The attachment to cargo is also not revealed via proteomics as no adaptor proteins that link dynein-dynactin to its cargo have been shown.
Should the authors qualify some of their claims as preliminary or speculative, or remove them altogether?
F-actin
Observations of dynein
Dynein versus kinesin-based transport
The calculation presented in lines 147-151 does not account for the fact that both the dynein-dynactin complex and kinesin proteins require cargo adaptors to transport cargo. Additionally, the authors overlook the possibility that multiple motors could be attached to a single cargo. If the authors did not observe this, they should explicitly mention it to support their argument. In short, the calculations are based on an incorrect premise, rendering the comparison inaccurate. Unless the authors have identified any dynein-dynactin or kinesin cargo adaptors in their proteomics data which could be used for such a comparison, we believe the authors lack sufficient data to accurately estimate the "active transport ratio" between dynein and kinesin.
Would additional experiments be essential to support the claims of the paper?
F-actin distance and length distribution
Quantifications for judgement of representativity
The authors should quantify how often they observed vesicles with a stick-like connection to MTs (lines 106-107); this would strengthen the interpretation of the density, as currently only one example is shown in the manuscript (Figure 4A). If possible, they could show how many of them are facing towards the MT plus end.
Dynein quantifications
Artefacts due to purification: Authors should discuss if the purification could have effects on visualizing components of the manchette. For example, if it has effect on the MTs and actin structure or the abundance/structure of the motor protein complexes (bound to cargo or isolated).
Are the experiments adequately replicated and statistical analysis adequate?
The cryo-ET data presented in the manuscript is collected using two separate sample preparations. Along with the quantifications of the different observations suggested above which will help the reader assess how abundant and representative these observations are, the authors could further strengthen their claims by acquiring data from a third sample preparation and then analysing how consistent their observations are between different purifications. This however could be time consuming so it is not a major requirement but recommended if possible within a short time frame.
Are the suggested experiments realistic in terms of time and resources? It would help if you could add an estimated cost and time investment for substantial experiments.
Minor comments:
Specific experimental issues that are easily addressable.
Are prior studies referenced appropriately?
We recommend including additional references to support the claim that F-actin has a mechanical role (line 242). Could the authors compare their proteomics data to other mass spectrometry studies conducted on the Manchette (for example see reference 4)?
Are the text and figures clear and accurate?
Text: We do not see the necessity of specifying the microtubules (MTs) in the data as "manchette MTs" or "mMTs" rather than simply "MTs". However, we recommend that the authors use either "MT" or "mMT" consistently throughout the manuscript.
The authors appear to refer to both dynein-1 (cytoplasmic dynein) and dynein-2 (axonemal dynein or IFT dynein). To avoid confusion, it is important that the authors clearly specify which dynein they are referring to throughout the text. This is particularly relevant as the study aims to demonstrate that IFT is not part of the manchette transport system.
Do you have suggestions that would help the authors improve the presentation of their data and conclusions?
We suggest revising the paragraph title "Dynein-mediated cargo along the manchette" (line 165) to "Dynein-mediated cargo transport along the manchette".
We recommend that the authors provide additional evidence to support the interpretation that the observed EM densities correspond to motor proteins. Specifically:
References:
This study employs cryo-electron tomography (cryo-ET) and proteomics to elucidate the architecture of the manchette. It advances our understanding of the components involved in intracellular transport within the manchette and introduces the following technical and conceptual innovations:
a) Technical Advances:
The authors have visualized the manchette at high resolution using cryo-ET. They optimized a purification pipeline capable of retaining, at least partially, the transport machinery of the manchette. Notably, they observed dynein and putative kinesin motors attached to microtubules-a significant achievement that, to our knowledge, has not been reported previously.
b) Conceptual Advances:
This study provides novel insights into spermatogenesis. The findings suggest that intraflagellar transport (IFT) is unlikely to play a role at this stage of sperm development while shedding light on alternative transport systems. Importantly, the authors demonstrate that actin filaments organize in two distinct ways: clustering parallel to microtubules or forming single filaments.
This work is likely to be of considerable interest to researchers in sperm development and structural biology. Additionally, it may appeal to scientists studying motor proteins and the cytoskeleton.
The reviewers possess extensive expertise in in situ cryo-electron tomography and single-particle microscopy, including work on dynein-based complexes. Collectively, they have significant experience in the field of cytoskeleton-based transport.
Reviewer #1 (Public review):
Lipid transfer proteins (LTPs) play a crucial role in the intramembrane lipid exchange within cells. However, the molecular mechanisms that govern this activity remain largely unclear. Specifically, the way in which LTPs surmount the energy barrier to extract a single lipid molecule from a lipid bilayer is not yet fully understood. This manuscript investigates the influence of membrane properties on the binding of Ups1 to the membrane and the transfer of phosphatidic acid (PA) by the LTP. The findings reveal that Ups1 shows a preference for binding to membranes with positive curvature. Moreover, coarse-grained molecular dynamics simulations indicate that positive curvature decreases the energy barrier associated with PA extraction from the membrane. Additionally, lipid transfer assays conducted with purified proteins and liposomes in vitro demonstrate that the size of the donor membrane significantly impacts lipid transfer efficiency by Ups1-Mdm35 complexes, with smaller liposomes (characterized by high positive curvature) promoting rapid lipid transfer.
This study offers significant new insights into the reaction cycle of phosphatidic acid (PA) transfer by Ups1 in mitochondria. Notably, the authors present compelling evidence that, alongside negatively charged phospholipids, positive membrane curvature enhances lipid transfer - an effect that is particularly relevant at the mitochondrial outer membrane. The experiments are technically robust, and my primary feedback pertains to the interpretation of specific results.
(1) The authors conclude from the lipid transfer assays (Figure 5) that lipid extraction is the rate-limiting step in the transfer cycle. While this conclusion seems plausible, it should be noted that the authors employed high concentrations of Ups1-Mdm35 along with less negatively charged phospholipids in these reactions. This combination may lead to binding becoming the rate-limiting factor. The authors should take this point into consideration. In this type of assay, it is challenging to clearly distinguish between binding, lipid extraction, and membrane dissociation as separate processes.
(2) The authors should discuss that variations in the size of liposomes will also affect the distance between them at a constant concentration, which may affect the rate of lipid transfer. Therefore, the authors should determine the average size and size distribution of liposomes after sonication (by DLS or nanoparticle analyzer, etc.).
(3) The authors use NBD-PA in the lipid transfer assays. Does the size of the donor liposomes affect the transfer of NBD-PA and DOPA similarly? Since NBD-labeled lipids are somewhat unstable within lipid bilayers (as shown by spontaneous desorption in Figure 5B), monitoring the transfer of unlabeled PA in at least one setting would strengthen the conclusion of the swap experiments.
(4) The present study suggests that membrane domains with positive curvature at the outer membrane may serve as starting points for lipid transport by Ups1-Mdm35. Is anything known about the mechanisms that form such structures? This should be discussed in the text.