Author response:
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public review):
(1) Figure 1B shows the PREDICTED force-extension curve for DNA based on a worm-like chain model. Where is the experimental evidence for this curve? This issue is crucial because the F-E curve will decide how and when a catch-bond is induced (if at all it is) as the motor moves against the tensiometer. Unless this is actually measured by some other means, I find it hard to accept all the results based on Figure 1B.
The Worm-Like-Chain model for the elasticity of DNA was established by early work from the Bustamante lab (Smith et al., 1992) and Marko and Siggia (Marko and Siggia, 1995), and was further validated and refined by the Block lab (Bouchiat et al., 1999; Wang et al., 1997). The 50 nm persistence length is the consensus value, and was shown to be independent of force and extension in Figure 3 of Bouchiat et al (Bouchiat et al., 1999). However, we would like to stress that for our conclusions, the precise details of the Force-Extension relationship of our dsDNA are immaterial. The key point is that the motor stretches the DNA and stalls when it reaches its stall force. Our claim of the catch-bond character of kinesin is based on the longer duration at stall compared to the run duration in the absence of load. Provided that the motor is indeed stalling because it has stretched out the DNA (which is strongly supported by the repeated stalling around the predicted extension corresponding to ~6 pN of force), then the stall duration depends on neither the precise value for the extension nor the precise value of the force at stall.
(2) The authors can correct me on this, but I believe that all the catch-bond studies using optical traps have exerted a load force that exceeds the actual force generated by the motor. For example, see Figure 2 in reference 42 (Kunwar et al). It is in this regime (load force > force from motor) that the dissociation rate is reduced (catch-bond is activated). Such a regime is never reached in the DNA tensiometer study because of the very construction of the experiment. I am very surprised that this point is overlooked in this manuscript. I am therefore not even sure that the present experiments even induce a catch-bond (in the sense reported for earlier papers).
It is true that Kunwar et al measured binding durations at super-stall loads and used that to conclude that dynein does act as a catch-bond (but kinesin does not) (Kunwar et al., 2011). However, we would like to correct the reviewer on this one. This approach of exerting super-stall forces and measuring binding durations is in fact less common than the approach of allowing the motor to walk up to stall and measuring the binding duration. This ‘fixed trap’ approach has been used to show catch-bond behavior of dynein (Leidel et al., 2012; Rai et al., 2013) and kinesin (Kuo et al., 2022; Pyrpassopoulos et al., 2020). For the non-processive motor Myosin I, a dynamic force clamp was used to keep the actin filament in place while the myosin generated a single step (Laakso et al., 2008). Because the motor generates the force, these are not superstall forces either.
(3) I appreciate the concerns about the Vertical force from the optical trap. But that leads to the following questions that have not at all been addressed in this paper:
(i) Why is the Vertical force only a problem for Kinesins, and not a problem for the dynein studies?
Actually, we do not claim that vertical force is not a problem for dynein; our data do not speak to this question. There is debate in the literature as to whether dynein has catch bond behavior in the traditional single-bead optical trap geometry - while some studies have measured dynein catch bond behavior (Kunwar et al., 2011; Leidel et al., 2012; Rai et al., 2013), others have found that dynein has slip-bond or ideal-bond behavior (Ezber et al., 2020; Nicholas et al., 2015; Rao et al., 2019). This discrepancy may relate to vertical forces, but not in an obvious way.
(ii) The authors state that "With this geometry, a kinesin motor pulls against the elastic force of a stretched DNA solely in a direction parallel to the microtubule". Is this really true? What matters is not just how the kinesin pulls the DNA, but also how the DNA pulls on the kinesin. In Figure 1A, what is the guarantee that the DNA is oriented only in the plane of the paper? In fact, the DNA could even be bending transiently in a manner that it pulls the kinesin motor UPWARDS (Vertical force). How are the authors sure that the reaction force between DNA and kinesin is oriented SOLELY along the microtubule?
We acknowledge that “solely” is an absolute term that is too strong to describe our geometry. We softened this term in our revision to “nearly parallel to the microtubule” (Line 464). In the Geometry Calculations section of Supplementary Methods, we calculate that if the motor and streptavidin are on the same protofilament, the vertical force will be <1% of the horizontal force. We also note that if the motor is on a different protofilament, there will be lateral forces and forces perpendicular to the microtubule surface, except they are oriented toward rather than away from the microtubule. The DNA can surely bend due to thermal forces, but because inertia plays a negligible role at the nanoscale (Howard, 2001; Purcell, 1977), any resulting upward forces will only be thermal forces, which the motor is already subjected to at all times.
(4) For this study to be really impactful and for some of the above concerns to be addressed, the data should also have included DNA tensiometer experiments with Dynein. I wonder why this was not done?
As much as we would love to fully characterize dynein here, this paper is about kinesin and it took a substantial effort. The dynein work merits a stand-alone paper.
While I do like several aspects of the paper, I do not believe that the conclusions are supported by the data presented in this paper for the reasons stated above.
The three key points the reviewer makes are the validity of the worm-like-chain model, the question of superstall loads, and the role of DNA bending in generating vertical forces. We hope that we have fully addressed these concerns in our responses above.
Reviewer #2 (Public review):
Major comments:
(1) The use of the term "catch bond" is misleading, as the authors do not really mean consistently a catch bond in the classical sense (i.e., a protein-protein interaction having a dissociation rate that decreases with load). Instead, what they mean is that after motor detachment (i.e., after a motor protein dissociating from a tubulin protein), there is a slip state during which the reattachment rate is higher as compared to a motor diffusing in solution. While this may indeed influence the dynamics of bidirectional cargo transport (e.g., during tug-of-war events), the used terms (detachment (with or without slip?), dissociation, rescue, ...) need to be better defined and the results discussed in the context of these definitions. It is very unsatisfactory at the moment, for example, that kinesin-3 is at first not classified as a catch bond, but later on (after tweaking the definitions) it is. In essence, the typical slip/catch bond nomenclature used for protein-protein interaction is not readily applicable for motors with slippage.
We acknowledge that our treatment of kinesin-3 was confusing. In response, we deleted any reference to kinesin-3 catch-bond in the Results section, and restricted it to the Discussion where it is interpretation. In Line 635 in the Discussion, we softened the statement of catch-bond activity to “…all three dominant kinesin transport families display catch-bond like behavior at stall…”. We acknowledge that, classically, the catch/slip bond nomenclature refers to simple protein-protein interactions and is easier to interpret there. However, the term ‘catch-bond’ has been used in the literature for myosin, dynein and kinesin, and thus we feel that it is sufficiently established to use it here.
(2) The authors define the stall duration as the time at full load, terminated by >60 nm slips/detachments. Isn't that a problem? Smaller slips are not detected/considered... but are also indicative of a motor dissociation event, i.e., the end of a stall. What is the distribution of the slip distances? If the slip distances follow an exponential decay, a large number of short slips are expected, and the presented data (neglecting those short slips) would be highly distorted.
The reviewer brings up a good point that there may be undetected slips. To address this question, we plotted the distribution of slip distances for kinesin-3, which by far had the most slip events. As the reviewer suggested, it is indeed an exponential distribution, and we calculated a corrected kinesin-3 stall duration due to these undetected slips. This data and analysis are included as a new Supplementary Figure S8. In the main text on Lines 283-293 we included the following text:
“It was notable that the kinesin-3 stall durations at high load are longer than the ramp durations at low load, because this indicates that the kinesin-3 off-rate slows with increasing load. However, because kinesin-3 had the most slip events at stall, we were concerned that there may be undetected slip events below the 60 nm threshold of detection that led to an overestimation of the kinesin-3 stall duration. To test this hypothesis, we plotted the distribution of kinesin-3 slip distances at stall, fit an exponential, and calculated the fraction of missed slip events (Fig. S8). From this analysis, we calculated a correction factor of 1.42 that brought the kinesin-3 stall duration down 1.33 s. Notably, this stall duration value is still well above the kinesin-3 ramp duration value of 0.75 s in Fig. 3C and thus does not qualitatively change our conclusions.”
We thank the reviewer for this suggestion.
(3) Along the same line: Why do the authors compare the stall duration (without including the time it took the motor to reach stall) to the unloaded single motor run durations? Shouldn't the times of the runs be included?
The elastic force of the DNA spring is variable as the motor steps up to stall, and so if we included the entire run duration then it would be difficult to specify what force we were comparing to unloaded. More importantly, if we assume that any stepping and detachment behavior is history independent, then it is mathematically proper to take any arbitrary starting point (such as when the motor reaches stall), start the clock there, and measure the distribution of detachments durations relative to that starting point. More importantly, what we do in Fig. 3 is to separate out the ramps from the stalls and, using a statistical model, we compute a separate duration parameter (which is the inverse of the off-rate) for the ramp and the stall. What we find is that the relationship between ramp, stall, and unloaded durations is different for the three motors, which is interesting in itself.
(4) At many places, it appears too simple that for the biologically relevant processes, mainly/only the load-dependent off-rates of the motors matter. The stall forces and the kind of motor-cargo linkage (e.g., rigid vs. diffusive) do likely also matter. For example: "In the context of pulling a large cargo through the viscous cytoplasm or competing against dynein in a tug-of-war, these slip events enable the motor to maintain force generation and, hence, are distinct from true detachment events." I disagree. The kinesin force at reattachment (after slippage) is much smaller than at stall. What helps, however, is that due to the geometry of being held close to the microtubule (either by the DNA in the present case or by the cargo in vivo) the attachment rate is much higher. Note also that upon DNA relaxation, the motor is likely kept close to the microtubule surface, while, for example, when bound to a vesicle, the motor may diffuse away from the microtubule quickly (e.g., reference 20).
We appreciate the reviewer’s detailed thinking here, and we offer our perspective. As to the first point, we agree that the stall force is relevant and that the rigidity of the motor-cargo linkage will play a role. The goal of the sentence on pulling cargo that the reviewer highlights is to set up our analysis of slips, which we define as rearward displacements that don’t return to the baseline before force generation resumes. We revised this sentence to the following: “In the context of pulling a large cargo through the viscous cytoplasm or competing against dynein in a tug-of-war, these slip events enable the motor to continue generating force after a small rearward displacement, rather than fully detaching and ‘resetting’ to zero load.” (Line 339-342)
It should be noted that, as shown in the model diagram in Fig. 5, we differentiate between the slip state (and recovery from this slip state) and the detached state (and reattachment from this detached state). This delineation is important because, as the reviewer points out, if we are measuring detachment and reattachment with our DNA tensiometer, then the geometry of a vesicle in a cell will be different and diffusion away from the microtubule or elastic recoil perpendicular to the microtubule will suppress this reattachment.
Our evidence for a slip state in which the motor maintains association with the microtubule comes from optical trapping work by Tokelis et al (Toleikis et al., 2020) and Sudhakar et al (Sudhakar et al., 2021). In particular, Sudhakar used small, high index Germanium microspheres that had a low drag coefficient. They showed that during ‘slip’ events, the relaxation time constant of the bead back to the center of the trap was nearly 10-fold slower than the trap response time, consistent with the motor exerting drag on the microtubule. (With larger beads, the drag of the bead swamps the motor-microtubule friction.) Another piece of support for the motor maintaining association during a slip is work by Ramaiya et al. who used birefringent microspheres to exert and measure rotational torque during kinesin stepping (Ramaiya et al., 2017). In most traces, when the motor returned to baseline following a stall, the torque was dissipated as well, consistent with a ‘detached’ state. However, a slip event is shown in S18a where the motor slips backward while maintaining torque. This is best explained by the motor slipping backward in a state where the heads are associated with the microtubule (at least sufficiently to resist rotational forces). Thus, we term the resumption after slip to be a rescue from the slip state rather than a reattachment from the detached state.
To finish the point, with the complex geometry of a vesicle, during slip events the motor remains associated with the microtubule and hence primed for recovery. This recovery rate is expected to be the same as for the DNA tensiometer. Following a detachment, however, we agree that there will likely be a higher probability of reattachment in the DNA tensiometer due to proximity effects, whereas with a vesicle any elastic recoil or ‘rolling’ will pull the detached motor away from the microtubule, suppressing reattachment. To address this point, we added in the Discussion on lines 654-656:
“Additionally, any ‘rolling’ of a spherical cargo following motor detachment will tend to suppress the motor reattachment rate.”
(5) Why were all motors linked to the neck-coil domain of kinesin-1? Couldn't it be that for normal function, the different coils matter? Autoinhibition can also be circumvented by consistently shortening the constructs.
We chose this dimerization approach to focus on how the mechoanochemical properties of kinesins vary between the three dominant transport families. We agree that in cells, autoinhibition of both kinesins and dynein likely play roles in regulating bidirectional transport, as will the activity of other regulatory proteins. The native coiled-coils may act as ‘shock absorbers’ due to their compliance, or they might slow the motor reattachment rate due to the relatively large search volumes created by their long lengths (10s of nm). These are topics for future work. By using the neck-coil domain of kinesin-1 for all three motors, we eliminate any differences in autoinhibition or other regulation between the three kinesin families and focus solely on differences in the mechanochemistry of their motor domains.
(6) I am worried about the neutravidin on the microtubules, which may act as roadblocks (e.g. DOI: 10.1039/b803585g), slip termination sites (maybe without the neutravidin, the rescue rate would be much lower?), and potentially also DNA-interaction sites? At 8 nM neutravidin and the given level of biotinylation, what density of neutravidin do the authors expect on their microtubules? Can the authors rule out that the observed stall events are predominantly the result of a kinesin motor being stopped after a short slippage event at a neutravidin molecule?
(7) Also, the unloaded runs should be performed on the same microtubules as in the DNA experiments, i.e., with neutravidin. Otherwise, I do not see how the values can be compared.
To address the question of neutravidin acting as a roadblock, we did the following. Because of the sequence of injections used to assemble the tensiometer in the flow cell, there are often some residual GFP-kinesin motors that aren’t attached to DNA and thus serve as internal controls for unloaded motility on the neutravidin-functionalized Mt. We quantified the run durations of these free kinesin-GFP and found that their run duration was 0.92 s (95% CI: 0.79 to 1.04 by MEMLET). This is slightly lower but not statistically different from the 1.04 s [0.78, 1.31] on control microtubules in Fig 2A. This result is included in Figure S6 in the revised manuscript.
We don’t have a precise estimate for the amount of neutravidin on the microtubules. Based on Fig. 3C of Korten and Diez (Korten and Diez, 2008), the reduction in the unloaded run duration that we see corresponds to a ~2% biotinylation ratio. We polymerize Mt with 10% biotinylated tubulin and add 8 nM neutravidin to the flow cell, so in principle the microtubules could be 10% biotin-streptavidin coated. However, there are a number of uncertainties that push this estimate lower – a) the precise degree of biotinylation, b) whether the %biotinylated tubulin in polymerized microtubules is lower than the mixing ratio due to unequal incorporation, and 3) what fraction of the biotinylated tubulin are occupied by the neutravidin when using this neutravidin flow-in method. Thus, our best estimate is ~2% biotin-streptavidin functionalization.
The ramp durations in Fig. 3 provide another argument that biotinylated microtubules are not affecting the motors. Compared to unloaded durations for each motor, the kinesin-1 ramps were longer, the kinesin-2 ramps were the same, and the kinesin-3 ramps were shorter duration. That argues against any systematic effect of biotinylation on motor run durations, with the caveat that family-dependent differences could in principle be masking an effect. The fact that ramp durations aren’t systematically longer or shorter than the unloaded run durations also argues that the stalls we see, which are at the expected extension length of the dsDNA, are not caused by neutravidin roadblocks.
The final point the reviewer brings up is whether neutravidin may be contributing to the rescues from slips events that we observe. This is difficult to fully rule out. However, because the unloaded run durations aren’t significantly altered by the biotin-streptavidin on the microtubules, we don’t expect the rescue events following a slip to be significantly affected. In principle, we could systematically increase and decrease the biotinylation and see whether the slip rescues change, but we haven’t done this.
(8) If, as stated, "a portion of kinesin-3 unloaded run durations were limited by the length of the microtubules, meaning the unloaded duration is a lower limit." corrections (such as Kaplan-Meier) should be applied, DOI: 10.1016/j.bpj.2017.09.024.
(9) Shouldn't Kaplan-Meier also be applied to the ramp durations ... as a ramp may also artificially end upon stall? Also, doesn't the comparison between ramp and stall duration have a problem, as each stall is preceded by a ramp ...and the (maximum) ramp times will depend on the speed of the motor? Kinesin-3 is the fastest motor and will reach stall much faster than kinesin-1. Isn't it obvious that the stall durations are longer than the ramp duration (as seen for all three motors in Figure 3)?
The reviewer rightly notes the many challenges in estimating the motor off-rates during ramps. To estimate ramp off-rates and as an independent approach to calculating the unloaded and stall durations, we developed a Markov model coupled with Bayesian inference methods to estimate a duration parameter (equivalent to the inverse of the off-rate) for the unloaded, ramp, and stall duration distributions. With the ramps, we have left censoring due to the difficulty in detecting the start of the ramps in the fluctuating baseline, and we have right censoring due to reaching stall (with different censoring of the ramp duration for the three motors due to their different speeds). The Markov model assumes a constant detachment probability and history-independence, and thus is robust even in the face of left and right censoring (details in the Supplementary section). This approach is preferred over Kaplan-Meier because, although non-parametric methods such as K-M make no assumptions for the distribution, they require the user to know exactly where the start time is.
Regarding the potential underestimate of the kinesin-3 unloaded run duration due to finite microtubule lengths. The first point is that the unloaded duration data in Fig. 2C are quite linear up to 6 s and are well fit by the single-exponential fit (the points above 6 s don’t affect the fit very much). The second point is that when we used our Markov model (which is robust against right censoring) to estimate the unloaded and stall durations, the results agreed with the single-exponential fits very well (Table S2). Specifically, the single-exponential fit for the kinesin-3 unloaded duration was 2.74 s (2.33 – 3.17 s 95% CI) and the estimate from the Markov model was 2.76 (2.28 – 3.34 s 95% CI). Thus, we chose not to make any corrections to the kinesin-3 unloaded run durations due to finite microtubule lengths. To address this point in the revision, we added the following note in Table S2: “* Because the Markov-Bayesian model, which is unaffected by left and right censoring of data gave same unloaded run durations for kinesin-3 as the MEMLET fit, we did not the kinesin-3 unloaded run durations for any right censoring due to finite microtubule lengths.” We also added the following point in the legend of Fig. S1: “A fraction of kinesin-3 unloaded run durations were limited by the length of the microtubules, but fitting to a model that took into account missed events gave a similar mean duration as an exponential fit, and so no correction was made (Table S2).”
(10) It is not clear what is seen in Figure S6A: It looks like only single motors (green, w/o a DNA molecule) are walking ... Note: the influence of the attached DNA onto the stepping duration of a motor may depend on the DNA conformation (stretched and near to the microtubule (with neutravidin!) in the tethered case and spherically coiled in the untethered case).
In Figure S6 kymograph, the green traces are GFP-labeled kinesin-1 without DNA attached (which are in excess) and the red diagonal trace is a motor with DNA attached. We clarified this in the revised Figure S6 legend. We agree that the DNA conformation will differ if it is attached and stretched (more linear) versus simply being transported (random coil), but by its nature this control experiment is only addressing random coil DNA.
(11) Along this line: While the run time of kinesin-1 with DNA (1.4 s) is significantly shorter than the stall time (3.0 s), it is still larger than the unloaded run time (1.0 s). What do the authors think is the origin of this increase?
We addressed this point in lines 200-212 of the revised manuscript:
“We carried out two additional control experiments. First, to confirm that the neutravidin used to link the DNA to the microtubule wasn’t affecting kinesin motility, we analyzed the run durations of kinesin-1 motors on neutravidin-coated microtubules and found no change compared to unlabeled microtubules (Fig. S6). Second, we measured the run duration of kinesin-1 linked to a DNA tether that was not bound to the microtubule and thus was being transported (Fig. S6). The kinesin-DNA run duration was 1.40 s, longer than the 1.04 s of motors alone (Fig. 2A). We interpret this longer duration to reflect the slower diffusion constant of the dsDNA relative to the motor alone, which enables motors to transiently detach and rebind before the DNA cargo has diffused away, thus extending the run duration (Block et al., 1990). Notably, this slower diffusion constant should not play a role in the DNA tensiometer geometry because if the motor transiently detaches, it will be pulled backward by the elastic forces of the DNA and detected as a slip or detachment event.“
(12) "The simplest prediction is that against the low loads experienced during ramps, the detachment rate should match the unloaded detachment rate." I disagree. I would already expect a slight increase.
Agreed. We changed this text (Lines 265-267) to: “The prediction for a slip bond is that against the low loads experienced during ramps, the detachment rate should be equal to or faster than the unloaded detachment rate.”
(13) Isn't the model over-defined by fitting the values for the load-dependence of the strong-to-weak transition and fitting the load dependence into the transition to the slip state?
Essentially, yes, it is overdefined, but that is essentially by design and the model is still very useful. Our goal here was to make as simple a model as possible that could account for the data and use it to compare model parameters for the different motor families. Ignoring the complexity of the slip and detached states, a model with a strong and weak state in the stepping cycle and a single transition out of the stepping cycle is the simplest formulation possible. And having rate constants (k<sub>S-W</sub> and k<sub>slip</sub> in our case) that vary exponentially with load makes thermodynamic sense for modeling mechanochemistry (Howard, 2001). Thus, we were pleasantly surprised that this bare-bones model could recapitulate the unloaded and stall durations for all three motors (Fig. 5C-E).
(14) "When kinesin-1 was tethered to a glass coverslip via a DNA linker and hydrodynamic forces were imposed on an associated microtubule, kinesin-1 dissociation rates were relatively insensitive to loads up to ~3 pN, inconsistent with slip-bond characteristics (37)." This statement appears not to be true. In reference 37, very similar to the geometry reported here, the microtubules were fixed on the surface, and the stepping of single kinesin motors attached to large beads (to which defined forces were applied by hydrodynamics) via long DNA linkers was studied. In fact, quite a number of statements made in the present manuscript have been made already in ref. 37 (see in particular sections 2.6 and 2.7), and the authors may consider putting their results better into this context in the Introduction and Discussion. It is also noteworthy to discuss that the (admittedly limited) data in ref. 37 does not indicate a "catch-bond" behavior but rather an insensitivity to force over a defined range of forces.
The reviewer misquoted our sentence. The actual wording of the sentence was: “When kinesin-1 was connected to micron-scale beads through a DNA linker and hydrodynamic forces parallel to the microtubule imposed, dissociation rates were relatively insensitive to loads up to ~3 pN, inconsistent with slip-bond characteristics (Urbanska et al., 2021).” The sentence the reviewer quoted was in a previous version that is available on BioRxiv and perhaps they were reading that version. Nonetheless, in the Discussion of the revision, we added text to note that this behavior is indicative of an ideal bond (not a catch-bond) on Lines 480-483: “When kinesin-1 was connected to micron-scale beads through a DNA linker and hydrodynamic forces parallel to the microtubule imposed, dissociation rates were relatively insensitive to loads up to ~3 pN, inconsistent with slip-bond characteristics and instead characteristic of an ideal-bond.” We also added a sentence in the Introduction highlighting this work, Lines 84-87: “Fourth, when kinesin-1 was connected to a bead through a micron-long segment of DNA and hydrodynamic forces were imposed on the bead, motor interaction times were insensitive to hindering loads up to 3 pN, indicative of an ideal-bond.”
Reviewer #3 (Public review):
The authors attribute the differences in the behaviour of kinesins when pulling against a DNA tether compared to an optical trap to the differences in the perpendicular forces. However, the compliance is also much different in these two experiments. The optical trap acts like a ~ linear spring with stiffness ~ 0.05 pN/nm. The dsDNA tether is an entropic spring, with negligible stiffness at low extensions and very high compliance once the tether is extended to its contour length (Fig. 1B). The effect of the compliance on the results should be addressed in the manuscript.
This is an interesting point. We added the following paragraph in Lines 101-111 in the Geometry Consideration section of the Supplementary Methods.
“Another consideration when comparing the DNA tensiometer to optical trap measurements is the relative stiffness of the trap and dsDNA. Optical trap stiffnesses are generally in the range of 0.05 pN/nm [12,13]. To calculate the predicted stiffness of the dsDNA spring, we computed the slope of theoretical force-extension curve in Fig. 1B. The stiffness is highly nonlinear and is <0.001 pN/nM below 650 nm extension. At the predicted stall force of 6 pN (960 nm extension), the dsDNA stiffness ~0.2 pN/nm, which is stiffer than most optical traps, but it is similar to the estimated 0.3 pN/nm stiffness of kinesin motors themselves[12,13]. An 8 nm step at this stiffness leads to a 1.6 pN jump in force, so it is reasonable to expect that motors are dynamically stepping at stall. Therefore, there is no reason to expect that stiffness differences between optical traps and the dsDNA spring are affecting the motor detachment kinetics.”
Compared to an optical trapping assay, the motors are also tethered closer to the microtubule in this geometry. In an optical trap assay, the bead could rotate when the kinesin is not bound. The authors should discuss how this tethering is expected to affect the kinesin reattachment and slipping. While likely outside the scope of this study, it would be interesting to compare the static tether used here with a dynamic tether like MAP7 or the CAP-GLY domain of p150glued.
Please see our response to Reviewer #2 Major Comment #4 above, which asks this same question in the context of intracellular cargo. In response to the point from Reviewer #3, we added the following sentence on Lines 654-656: “Additionally, any ‘rolling’ of a spherical cargo following motor detachment will tend to suppress the motor reattachment rate.”
Regarding a dynamic tether, we agree that’s interesting – there are kinesins that have a second, non-canonical binding site that achieves this tethering (e.g. ncd and Cin8); p150glued likely does this naturally for dynein-dynactin-activator complexes; and we speculated in a review some years ago (Hancock, 2014) that during bidirectional transport kinesin and dynein may act as dynamic tethers for one another when not engaged, enhancing the activity of the opposing motor.
In the single-molecule extension traces (Figure 1F-H; S3), the kinesin-2 traces often show jumps in position at the beginning of runs (e.g., the four runs from ~4-13 s in Fig. 1G). These jumps are not apparent in the kinesin-1 and -3 traces. What is the explanation? Is kinesin-2 binding accelerated by resisting loads more strongly than kinesin-1 and -3?
We agree that at first glance those jumps are puzzling. To investigate this question the first thing we did was to go back to our tensiometer dataset and look systematically at jumps for all three motors. We found roughly 4-6 large jumps like these for all three motors (kinesin-1: 250 +/- 99 nm (mean +/- SD; N=5); kinesin-2: 249 +/- 165 nm (N=6); kinesin-3: 490 +/- 231 nm (N=4)). Thus, although the apparent jumps may be more pronounced due to the specific rebinding kinetics of kinesin-2, this behavior is not unique to this motor. (Note that the motor binding position distribution in Fig. S2 is taken from initial binding positions that follow a clear period of detachment; thus, not all jumps are captured there.)
Our interpretation is that these apparent jumps are simply a reflection of the long length and high compliance of the dsDNA tether. For instance, below 650 nm extension the stiffness, k <0.001 pN/nM (see Reviewer #3, point #1 above). Thus, we expect large fluctuations of the tethered motor when not bound to the microtubule. One reason that these events look like ‘jumps’ is that the sub-ms fluctuations during detached periods are not captured by the ~25 fps movies (40 ms frame acquisition time). Instead, the fitted Qdot position represents the average position during the acquisition window. Actually, due to these rapid fluctuations (and the limited depth of the TIRF illumination field) the position often can’t be determined during these periods of fluctuation (e.g. see gaps at ~2.5 s, 11 s and 24 s in Fig. 1F).
When comparing the durations of unloaded and stall events (Fig. 2), there is a potential for bias in the measurement, where very long unloaded runs cannot be observed due to the limited length of the microtubule (Thompson, Hoeprich, and Berger, 2013), while the duration of tethered runs is only limited by photobleaching. Was the possible censoring of the results addressed in the analysis?
Yes. Please see response to Reviewer #2 points (8) and (9) above.
The mathematical model is helpful in interpreting the data. To assess how the "slip" state contributes to the association kinetics, it would be helpful to compare the proposed model with a similar model with no slip state. Could the slips be explained by fast reattachments from the detached state?
In the model, the slip state and the detached states are conceptually similar; they only differ in the sequence (slip to detached) and the transition rates into and out of them. The simple answer is: yes, the slips could be explained by fast reattachments from the detached state. In that case, the slip state and recovery could be called a “detached state with fast reattachment kinetics”. However, the key data for defining the kinetics of the slip and detached states is the distribution of Recovery times shown in Fig. 4D-F, which required a triple exponential to account for all of the data. If we simplified the model by eliminating the slip state and incorporating fast reattachment from a single detached state, then the distribution of Recovery times would be a single-exponential with a time constant equivalent to t<sub>1</sub>, which would be a poor fit to the experimental distributions in Fig. 4D-F.
Recommendations for the authors:
Reviewing Editor Comments:
The reviewers are in agreement with the motivation and approach of this study. The use of DNA tethers is an important advance in tethering motor proteins to gain insight into how motors respond to load. However, all 3 reviewers express reservations on how well the results support the claims. In particular, the use of the term catch bond was problematic, with Reviewer #2 suggesting some alternative nomenclature. Reviewer #1 expressed concern with experimental evidence for the predicted force-extension curve shown in Figure 1. I agree with the reviewers that additional experimental evidence would be required to conclude the catch-bond detachment kinetics of kinesin.
Recommendations for the authors:
Reviewer #2 (Recommendations for the authors):
(1) By eye, the run lengths, e.g., of kin-1 look very long in Figure S1 ... certainly above the expected 1 µm. Please check and comment.
We agree that the long runs do stick out by eye in this figure. To address this point, we analyzed the run lengths and run times from the kymograph shown in Fig. S1. Fitting the run duration distribution gave t = 1.31 s with a 95% CI of 0.96 to 1.67. This is slightly longer than the 1.04 s duration in Fig. 2A, but the 95% CI include this population mean, and so the S1 data are not statistically significantly different. The run time distribution from the S1 kymograph is given in Author response image 1.
Author response image 1.
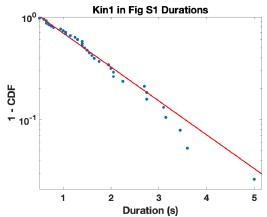
(2) The upper right kymograph in Figure 4A does not show a motor return to the baseline. Also, the scale bars, etc., are unreadable. Please modify.
Our purpose for showing the kymographs in Fig. 4A was to show the specific features of slips and fast and slow reattachment. Because we blew up the kymographs to show those specific features, it precluded us from showing the entire return to baseline. As suggested, we magnified the scale bars and the labels on the kymograph labels to make them readable.
Reviewer #3 (Recommendations for the authors):
(1) The frequent references to 95% confidence intervals disrupt the flow of the text. Perhaps the confidence intervals could be listed in a table rather than in the body of the text.
We deleted those from the text; they are shown in Fig. 2D and listed in Table S2.
We appreciate the efforts and helpful suggestions of all three reviewers and the Editor.
References
Block, S.M., L.S. Goldstein, and B.J. Schnapp. 1990. Bead movement by single kinesin molecules studied with optical tweezers. Nature. 348:348-352.
Bouchiat, C., M.D. Wang, J. Allemand, T. Strick, S.M. Block, and V. Croquette. 1999. Estimating the persistence length of a worm-like chain molecule from force-extension measurements. Biophys J. 76:409-413.
Ezber, Y., V. Belyy, S. Can, and A. Yildiz. 2020. Dynein Harnesses Active Fluctuations of Microtubules for Faster Movement. Nat Phys. 16:312-316.
Hancock, W.O. 2014. Bidirectional cargo transport: moving beyond tug of war. Nat Rev Mol Cell Biol. 15:615-628.
Howard, J. 2001. Mechanics of Motor Proteins and the Cytoskeleton. Sinauer Associates, Inc., Sunderland, MA. 367 pp.
Korten, T., and S. Diez. 2008. Setting up roadblocks for kinesin-1: mechanism for the selective speed control of cargo-carrying microtubules. Lab Chip. 8:1441-1447.
Kunwar, A., S.K. Tripathy, J. Xu, M.K. Mattson, P. Anand, R. Sigua, M. Vershinin, R.J. McKenney, C.C. Yu, A. Mogilner, and S.P. Gross. 2011. Mechanical stochastic tug-ofwar models cannot explain bidirectional lipid-droplet transport. Proc Natl Acad Sci U S A. 108:18960-18965.
Kuo, Y.W., M. Mahamdeh, Y. Tuna y J. Howard. 2022. The force required to remove tubulin from the microtubule lattice by pulling on its alpha-tubulin C-terminal tail. Nature communications. 13:3651.
Laakso, J.M., J.H. Lewis, H. Shuman, and E.M. Ostap. 2008. Myosin I can act as a molecular force sensor. Science. 321:133-136.
Leidel, C., R.A. Longoria, F.M. Gutierrez, and G.T. Shubeita. 2012. Measuring molecular motor forces in vivo: implications for tug-of-war models of bidirectional transport. Biophys J. 103:492-500.
Marko, J.F., and E.D. Siggia. 1995. Stretching DNA. Macromolecules. 28:8759-8770.
Nicholas, M.P., F. Berger, L. Rao, S. Brenner, C. Cho, and A. Gennerich. 2015. Cytoplasmic dynein regulates its attachment to microtubules via nucleotide state-switched mechanosensing at multiple AAA domains. Proc Natl Acad Sci U S A. 112:63716376.
Purcell, E.M. 1977. Life at low Reynolds Number. Amer J. Phys. 45:3-11.
Pyrpassopoulos, S., H. Shuman, and E.M. Ostap. 2020. Modulation of Kinesin's Load-Bearing Capacity by Force Geometry and the Microtubule Track. Biophys J. 118:243253.
Rai, A.K., A. Rai, A.J. Ramaiya, R. Jha, and R. Mallik. 2013. Molecular adaptations allow dynein to generate large collective forces inside cells. Cell. 152:172-182.
Ramaiya, A., B. Roy, M. Bugiel, and E. Schaher. 2017. Kinesin rotates unidirectionally and generates torque while walking on microtubules. Proc Natl Acad Sci U S A. 114:10894-10899.
Rao, L., F. Berger, M.P. Nicholas, and A. Gennerich. 2019. Molecular mechanism of cytoplasmic dynein tension sensing. Nature communications. 10:3332.
Smith, S.B., L. Finzi, and C. Bustamante. 1992. Direct mechanical measurements of the elasticity of single DNA molecules by using magnetic beads. Science. 258:11221126.
Sudhakar, S., M.K. Abdosamadi, T.J. Jachowski, M. Bugiel, A. Jannasch, and E. Schaher. 2021. Germanium nanospheres for ultraresolution picotensiometry of kinesin motors. Science. 371.
Toleikis, A., N.J. Carter, and R.A. Cross. 2020. Backstepping Mechanism of Kinesin-1. Biophys J. 119:1984-1994.
Urbanska, M., A. Ludecke, W.J. Walter, A.M. van Oijen, K.E. Duderstadt, and S. Diez. 2021. Highly-Parallel Microfluidics-Based Force Spectroscopy on Single Cytoskeletal Motors. Small. 17: e2007388.
Wang, M.D., H. Yin, R. Landick, J. Gelles, and S.M. Block. 1997. Stretching DNA with optical tweezers. Biophys J. 72:1335-1346.