An update of /usr/include/sysexits.h allocates previously unused exit codes from 64 - 78. It may be anticipated that the range of unallotted exit codes will be further restricted in the future. The author of this document will not do fixups on the scripting examples to conform to the changing standard. This should not cause any problems, since there is no overlap or conflict in usage of exit codes between compiled C/C++ binaries and shell scripts.
Eh, 0 and 64 - 78 are the only codes it defines. So if it had different codes defined before, what on earth were those codes before? Was only 0 "used"/defined here before? Nothing defined from 1-128? Or were the codes defined there different ones, like 20-42 and then they arbitrarily shifted these up to 64-78 one day? This is very unclear to me.
Also unclear whether this is saying it won't update for any future changes after this, or if he hasn't even updated to align with this supposed "change". (Unclear because I can't figure out whether his "proposes restricting user-defined exit codes to the range 64 - 113 (in addition to 0, for success), to conform with the C/C++ standard" statement is actually conforming or rejecting the sysexits.h standard.)
It seems that he's overreacting a bit here. It's hard to imagine there has been or will be any major changes to the sysexits.h. I would only imagine there being additions to, but not changes to because backwards compatibility would be of utmost concern.
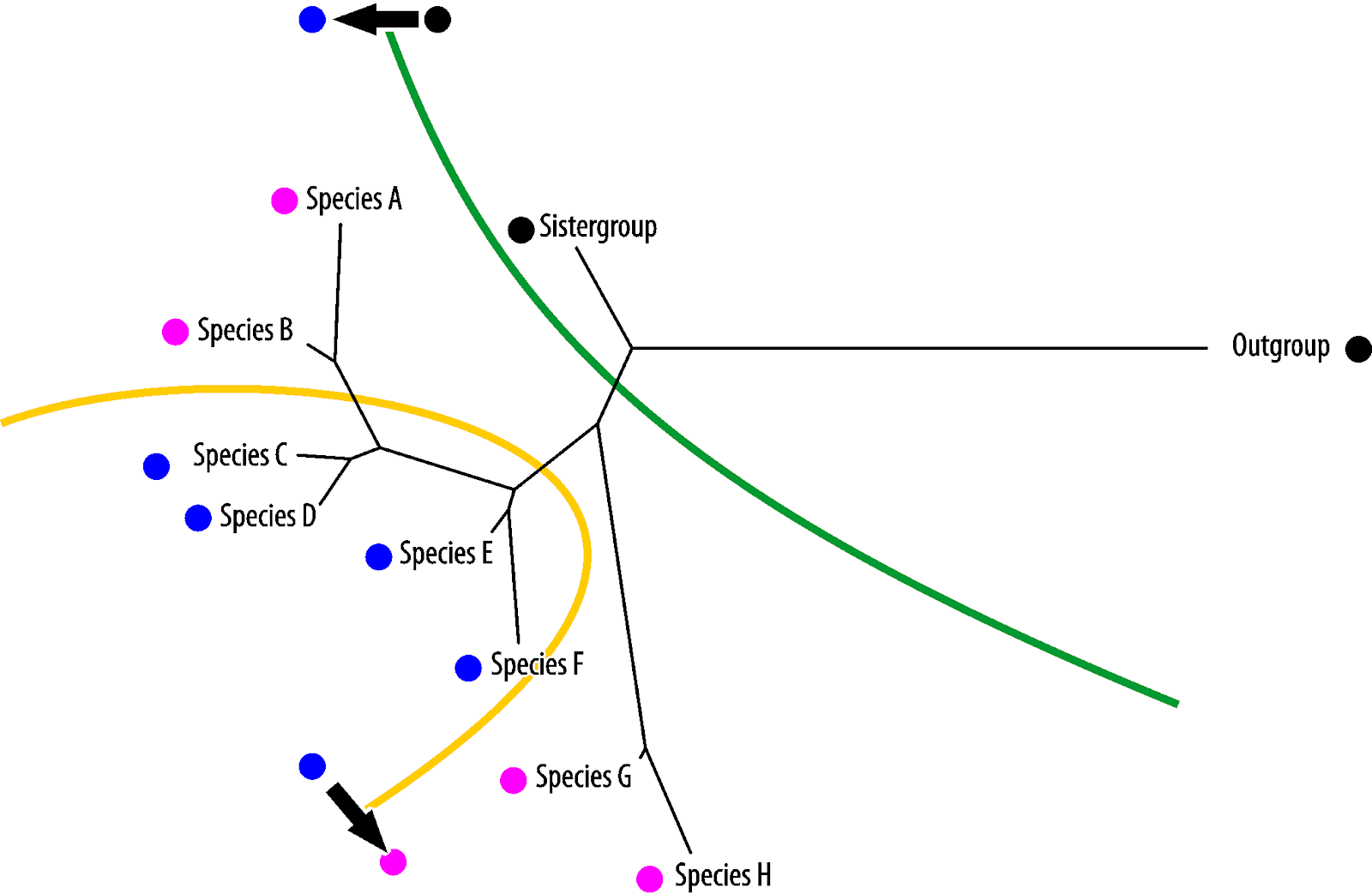
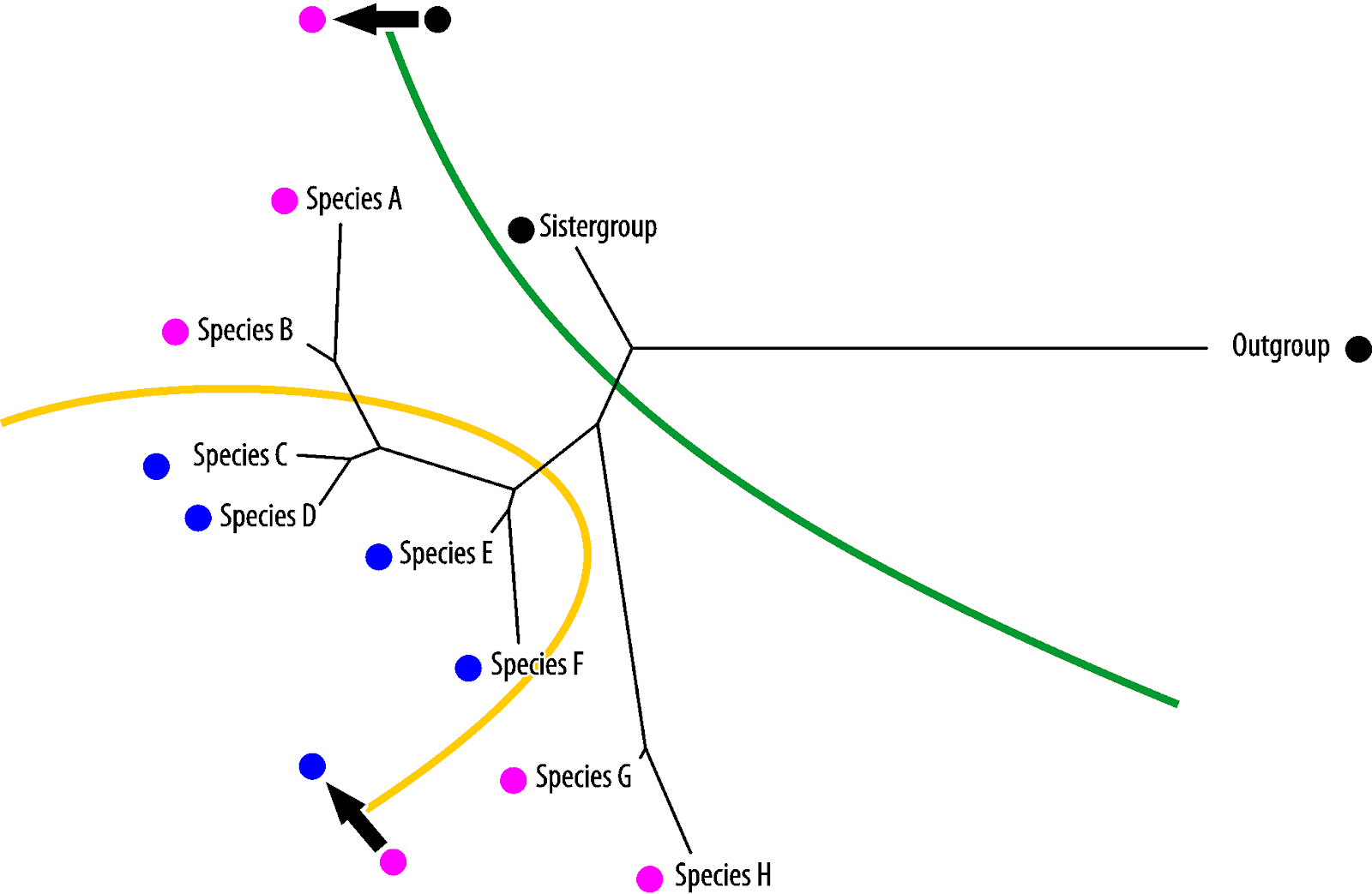 Vice versa, blue is the symplesiomorphy, pink the homoiology found only in the most derived (in absolute evolutionary terms) taxa
Vice versa, blue is the symplesiomorphy, pink the homoiology found only in the most derived (in absolute evolutionary terms) taxa