Format of HTML bookmarks file | Firefox Support Forum
to
What an abomination

Format of HTML bookmarks file | Firefox Support Forum
to
What an abomination

Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Recommendations For the Authors):
(1) While not absolutely necessary - it would be nice to see at least at the in-situ level what happens to the handful of other HC-important transcription factors in the Rbm24 KO (IKZF2, Barlh1, RFX) as the authors did look at Insm1.
Reply: Thanks for your suggested experiments. We agree that knowing whether the genes that are known to be involved in cell survival regulation are changed will provide insights into the mechanisms underlying cell death of Rbm24-/- HCs. Our data showed that Ikzf2 seemed to be upregulated when in the Rbm24-/- HCs, relative to Rbm24+/+ HCs at P5. We also tested Barlh1 and RFX, but we did not obtain confident data to present. Nonetheless, following the reviewer’s logic, we further tested Gata3, another gene involved in HC survival, and found that Gata3 was down-regulated in Rbm24 -/- HCs, compared to Rbm24+/+ HCs. Please refer to the text on lines 12-22 on page 12 and lines 1-10 on page 13, and Figure 3-figure supplement 1.
(2) Major comments: The nomenclature for mouse gene vs. mouse protein needs to be addressed throughout the manuscript. The nomenclature when referring to a mouse gene: gene symbols are italicized, with only the first letter in upper-case (e.g. Rbm24).
The nomenclature when referring to a mouse protein: Protein symbols are not italicized, and all letters are in upper-case (e.g. RBM24).
Reply: Thanks for pointing it out. In the entire manuscript, we have followed the reviewer’s comments to list gene and protein.
(3) Supplemental Figure 2D: Individual data points should be displayed on the bar graph via dots. SEM is not appropriate for this graph as SEM precision with only 3 samples is low. Furthermore, readers are more interested in knowing the variability within samples and not proximity of mean to the population mean, therefore standard deviation (SD) should be used instead.
Reply: We have edited the Figure 1-figure supplement 2D, as suggested. The Figure 1figure supplement 2 legend was updated, too. Please refer to line 21-22 on page 32.
(4) Red/Green should be avoided, especially when both are on the same image (merged immunofluorescence images that are found throughout the manuscript). I highly recommend changing to a color-blind friendly color scheme (such as cyan/green/magenta, cyan/magenta/yellow, etc.) for inclusivity.
Reply: Thanks for pointing it out. We have changed the red to magenta in all our Figures and figure supplements.
(5) Minor comments: As CRISPR-stop is a major method used throughout the paper, a brief explanation is needed for readers to understand what this methodology entails and why it was used. Something along the lines of," The CRISPR-stop technique allows for the introduction of early stop codons without the induction of DNA damage via Cas9 which can cause deleterious effects".
Reply: We have further elaborated how CRISPR-stop works and its advantages. Please refer to lines 8-13 on page 5.
(6) Page 5; line 5 - "Phenotypes occur earlier..." Grammar
Reply: The grammar error was corrected. Please refer to line 4, page 5.
(7) Page 5; line 5 - "Given Pou4f3 is the upstream regulator..." Not proven, rephrase
Reply: We have rephrased this sentence. Please refer to lines 5-6 on page 5.
(8) Supplemental 1A: Fine, Proof of knockout, I wouldn't mention INSM1 being "irregular"
Reply: We have rephrased this sentence. Please refer to lines 2-3 on page 6.
(9) Page 5; line21 - "Alignment of Insm1+ OHCs was not as regular..." Not a good description
Reply: We have rephrased this sentence. Please refer to lines 2-3 on page 6.
(10) Page 6; line11 - "Rbm24 was completely absent.." Redundancy with line 9
Reply: Thanks for pointing it out, and we have removed the redundant sentence.
(11) Page 7 - HA tag should be indicated originally as: Hemaglutinin (HA)
Reply: We have switched “HA” to “Hemaglutinin (HA)”. Please refer to line 15, page 7.
(12) Page 9, line 11- "Determine if autonomous/noncell autonomous." Disagree, cells still clustered in supplemental fig 4.
Reply: We have removed this sentence.
Reviewer #2 (Recommendations For The Authors):
The writing of the manuscript is adequate, but it would certainly be improved by professional editing.
Reply: Thanks for the reviewer’s encouraging comments. The revised version of our manuscript has been edited by an English native speaker.
Author Response
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1:
Summary:
In this study, Yan et al. investigate the molecular bases underlying mating type recognition in Tetrahymena thermophila. This model protist possesses a total of 7 mating types/sexes and mating occurs only between individuals expressing different mating types. The authors aimed to characterize the function of mating type proteins (MTA and MTB) in the process of self- and non-self recognition, using a combination of elegant phenotypic assays, protein studies, and imaging. They showed that the presence of MTA and MTB in the same cell is required for the expression of concavalin-A receptors and for tip transformation - two processes that are characteristic of the costimulation phase that precedes cell fusion. Using protein studies, the authors identify a set of additional proteins of varied functions that interact with MTA and MTB and are likely responsible for the downstream signaling processes required for mating. This is a description of a fascinating self- and non-self-recognition system and, as the authors point out, it is a rare example of a system with numerous mating types/sexes. This work opens the door for the further understanding of the molecular bases and evolution of these complex recognition systems within and outside protists.
The results shown in this study point to the unequivocal requirement of MTA and MTB proteins for mating. Nevertheless, some of the conclusions regarding the mode of functioning of these proteins are not fully supported and require additional investigation.
Strengths:
(1) The authors have established a set of very useful knock-out and reporter lines for MT proteins and extensively used them in sophisticated and well-designed phenotypic assays that allowed them to test the role of these proteins in vivo.
(2) Despite their apparent low abundance, the authors took advantage of a varied set of protein isolation and characterization techniques to pinpoint the localization of MT proteins to the cell membrane, and their interaction with multiple other proteins that could be downstream effectors. This opens the door for the future characterization of these proteins and further elucidation of the mating type recognition cascade.
Weaknesses:
The manuscript is structured and written in a very clear and easy-to-follow manner. However, several conclusions and discussion points fall short of highlighting possible models and mechanisms through which MT proteins control mating type recognition:
(1) The authors dismiss the possibility of a "simple receptor-ligand system", even though the data does not exclude this possibility. The model presented in Figure 2 S1, and on which the authors based their hypothesis, assumes the independence of MTA and MTB proteins in the generation of the intracellular cascade. However, the results presented in Figure 2 show that both proteins are required to be active in the same cell. Coupled with the fact that MTA and MTB proteins interact, this is compatible with a model where MTA would be a ligand and MTB a receptor (or vice-versa), and could thus form a receptor-ligand complex that could potentially be activated by a non-cognate MTA-MTB receptor-ligand complex, leading to an intracellular cascade mediated by the identified MRC proteins. As it stands, it is not clear what is the proposed working model, and it would be very beneficial for the reader for this to be clarified by having the point of view of the authors on this or other types of models.
We are very grateful that Reviewer #1 proposed the possibility that MTA and MTB form a receptor-ligand complex in which one acting as the ligand and the other as the receptor. We considered this hypothesis when asking how dose MTRC function, too. However, our current results do not support this idea. For instance, if MTA were a ligand and MTB a receptor, we would expect a mating signal upon treatment with MTAxc protein, but not with MTBxc. Contrary to this expectation, our experiments revealed that both MTAxc and MTBxc exhibit very similar effects (Figure 5, green and blue), and their combined treatment produces a stronger effect (Figure 5, teal). This suggests a mixed function for both proteins. (We incorporated this discussion into the revised version [line 120-121, 240-244].) It is pity that our current knowledge does not provide a detailed molecular mechanism for this intricate system. We are actively investigating the protein structures of MTA, MTB, and the entire MTRC, hoping to gain deeper insights into the molecular functions of MTA and MTB.
Additionally, we also realized that the expression we used in the previous version, “simple receptor-ligand model”, is not clearly defined. As Reviewer #1 pointed out, in this section, we examined whether the individual proteins of MTA and MTB act as a couple of receptor and ligand. We think this is the simplest possibility as a null hypothesis for Tetrahymena mating-type recognition. We have clarified it in the revised version (line 90-91, 104-106). According to this section, we proposed that MTA and MTB may form a complex that serves as a recognizer (functioning as both ligand and receptor) (line 117-118).
(2) The presence of MTA/MTB proteins is required for costimulation (Figure 2), and supplementation with non-cognate extracellular fragments of these proteins (MTAxc, or MTBxc) is a positive stimulator of pairing. However, alone, these fragments do not have the ability to induce costimulation (Figure 5). Based on the results in Figures 5 and 6 the authors suggest that MT proteins mediate both self and non-self recognition. Why do MTAxc and MTBxc not induce costimulation alone? Are any other components required? How to reconcile this with the results of Figure 2? A more in-depth interpretation of these results would be very helpful, since these questions remain unanswered, making it difficult for the reader to extract a clear hypothesis on how MT proteins mediate self- and non-self-recognition.
Several factors could contribute to the inability of MTA/Bxc to induce costimulation. It is highly likely that additional components are necessary, given that MTA/B form a protein complex with other proteins. Moreover, the expression of MTA/Bxc in insect cells, compared with Tetrahymena, might result in differences in post-translational modifications. Additionally, there are variations in protein conditions; on the Tetrahymena membrane, these proteins are arranged regularly and concentrated in a small area, while MTA/Bxc is randomly dispersed in the medium. The former condition could be more efficient. If there is a threshold required to stimulate a costimulation marker, MTA/Bxc may fail to meet this requirement. Much more studies are needed to fully answer this question. We acknowledged this limitation in the revised version (line 244-248).
Reviewer #2:
This manuscript reports the discovery and analysis of a large protein complex that controls mating type and sexual reproduction of the model ciliate Tetrahymena thermophila. In contrast to many organisms that have two mating types or two sexes, Tetrahymena is multi-sexual with 7 distinct mating types. Previous studies identified the mating type locus, which encodes two transmembrane proteins called MTA and MTB that determine the specificity of mating type interactions. In this study, mutants are generated in the MTA and MTB genes and mutant isolates are studied for mating properties. Cells missing either MTA or MTB failed to co-stimulate wild-type cells of different mating types. Moreover, a mixture of mutants lacking MTA or MTB also failed to stimulate. These observations support the conclusion that MTA and MTB may form a complex that directs mating-type identity. To address this, the proteins were epitope-tagged and subjected to IP-MS analysis. This revealed that MTA and MTB are in a physical complex, and also revealed a series of 6 other proteins (MRC1-6) that together with MTA/B form the mating type recognition complex (MTRC). All 8 proteins feature predicted transmembrane domains, three feature GFR domains, and two are predicted to function as calcium transporters. The authors went on to demonstrate that components of the MTRC are localized on the cell surface but not in the cilia. They also presented findings that support the conclusion that the mating type-specific region of the MTA and MTB genes can influence both self- and non-self-recognition in mating.
Taken together, the findings presented are interesting and extend our understanding of how organisms with more than two mating types/sexes may be specified. The identification of the six-protein MRC complex is quite intriguing. It would seem important that the function of at least one of these subunits be analyzed by gene deletion and phenotyping, similar to the findings presented here for the MTA and MTB mutants. A straightforward prediction might be that a deletion of any subunit of the MRC complex would result in a sterile phenotype. The manuscript was very well written and a pleasure to read.
Thanks for the valuable comments and suggestions. We are currently in the process of constructing deletion strains for these genes. As of now, we have successfully obtained ΔMRC1-3 and MRC4-6 knockdown strains. Our preliminary observations indicate that ΔMRC1-3 strains are unable to undergo mating. However, we prefer not to include these results in the current manuscript, as we believe that more comprehensive studies are still needed.
Reviewer #3:
The authors describe the role, location, and function of the MTA and MTB mating type genes in the multi-mating-type species T. thermophila. The ciliate is an important group of organisms to study the evolution of mating types, as it is one of the few groups in which more than two mating types evolved independently. In the study, the authors use deletion strains of the species to show that both mating types genes located in each allele are required in both mating individuals for successful matings to occur. They show that the proteins are localized in the cell membrane, not the cilia, and that they interact in a complex (MTRC) with a set of 6 associated (non-mating type-allelic) genes. This complex is furthermore likely to interact with a cyclin-dependent kinase complex. It is intriguing that T. thermophila has two genes that are allelic and that are both required for successful mating. This coevolved double recognition has to my knowledge not been described for any other mating-type recognition system. I am not familiar with experimental research on ciliates, but as far as I can judge, the experiments appear well performed and mostly support the interpretation of the authors with appropriate controls and statistical analyses.
The results show clearly that the mating type genes regulate non-self-recognition, however, I am not convinced that self-recognition occurs leading to the suppression of mating. An alternative explanation could be that the MTA and MTB proteins form a complex and that the two extracellular regions together interact with the MTA+MTB proteins from different mating types. This alternative hypothesis fits with the coevolution of MTA and MTB genes observed in the phylogenetic subgroups as described by Yan et al. (2021 iScience). Adding MTAxc and/or MTBxc to the cells can lead to the occupation of the external parts of the full proteins thereby inhibiting the formation of the complex, which in turn reduces non-self interactions. Self-recognition as explained in Figure 2S1 suggests an active response, which should be measurable in expression data for example. This is in my opinion not essential, but a claim of self-recognition through the MTA and MTB should not be made.
We express our gratitude to Reviewer #3 for proposing the occupation model and have incorporated this possibility into the manuscript. We believe it is possible that occupation may serve as the molecular mechanism through which self-recognition negatively regulates mating. If there is a physical interaction between mating-type proteins of the same type, but this interaction blocks the recognition machinery rather than initiating mating, it can be considered a form of self-recognition. This aligns with the observation that strains expressing MTA/B6 and MTB2 mate normally with WT cells of all mating types except for VI and II (line 203-204). A concise discussion on this topic is included in the manuscript (line 288-293, 659-661). We are actively investigating the downstream aspects of mating-type recognition, and we hope to provide further insights into this question soon.
The authors discuss that T. thermophila has special mating-type proteins that are large, while those of other groups are generally small (lines 157-160 and discussion). The complex formed is very large and in the discussion, they argue that this might be due to the "highly complex process, given that there are seven mating types in all". There is no argument given why large is more complex, if this is complex, and whether more mating types require more complexity. In basidiomycete fungi, many more mating types than 7 exist, and the homeodomain genes involved in mating types are relatively small but highly diverse (Luo et al. 1994 PMID: 7914671). The mating types associated with GPCR receptors in fungi are arguably larger, but again their function is not that complex, and mating-type specific variations appear to evolve easily (Fowler et al 2004 PMID: 14643262; Seike et al. 2015 PMID: 25831518). The large protein complex formed is reminiscent of the fusion patches that develop in budding or fission yeasts. In these species, the mating type receptors are activated by ligand pheromones from the opposite mating type that induce polarity patch formation (see Sieber et al. 2023 PMID: 35148940 for a recent review). At these patches, growth (shmooing) and fusion occur, which is reminiscent (in a different order) of the tip transformation in T. thermophilia. The fusion of two cells is in all taxa a dangerous and complex event that requires the evolution of very strict regulation and the existence of a system like the MTRC and cyclin-dependent complex to regulate this process is therefore not unexpected. The existence of multiple mating types should not greatly complicate the process, as most of the machinery (except for the MTA and MTB) is identical among all mating types.
We are very grateful that Reviewer #3 provide this insightful view and relevant papers. In response to the feedback, we removed the sentences regarding “multiple mating types greatly complicate the process” in the revised version. Instead, we have introduced a discussion section comparing the mating systems of yeasts and Tetrahymena (line 279-286).
The Tetrahymena/ciliate genetics and lifecycle could be better explained. For a general audience, the system is not easy to follow. For example, the ploidy of the somatic nucleus with regards to the mating type is not clear to me. The MAC is generally considered "polyploid", but how does this work for the mating type? I assume only a single copy of the mating type locus is available in the MAC to avoid self-recognition in the cells. Is it known how the diploid origin reduces to a single mating type? This does not become apparent from Cervantes et al. 2013.
In T. thermophila, the MIC (diploid) contains several mating-type gene pairs (mtGP, i.e., MTA and MTB) organized in a tandem array at the mat locus on each chromosome. In sexual reproduction, the new MAC of the progeny develops from the fertilized MIC through a series of genome editing events, and its ploidy increases to ~90 by endoreduplication. During this process, mtGP loss occurs, resulting in only one mtGP remaining on the MAC chromosome. The mating-type specificity of mtGPs on each chromosome within one nucleus becomes relatively pure through intranuclear coordination. After multiple assortments (possibly caused by MAC amitosis during cell fission), only mtGPs of one mating-type specificity exist in each cell, determining the cell’s mating type.
It is pity that the exact mechanisms involved in this complicated process remain a black box. The loss of mating-type gene pairs is hypothesized to involve a series of homologous recombination events, but this has not been completely proven. Furthermore, there is no clear understanding of how intranuclear coordination and assortment are achieved. While we have made observations confirming these events, a breakthrough in understanding the molecular mechanism is yet to be achieved.
We included more information in the revised version (line 672-683). Given the complexity of these unusual processes, we recommend an excellent review by Prof. Eduardo Orias (PMID: 28715961), which offers detailed explanations of the process and related concepts (line 685-686).
Also, the explanation of co-stimulation is not completely clear (lines 49-60). Initially, direct cell-cell contact is mentioned, but later it is mentioned that "all cells become fully stimulated", even when unequal ratios are used. Is physical contact necessary? Or is this due to the "secrete mating-essential factors" (line 601)? These details are essential, for interpretation of the results and need to be explained better.
Sorry that we didn’t realize the term “contact” is not precise enough. In Tetrahymena, physical contact is indeed necessary, but it can refer to temporary interactions. Unlike yeast, Tetrahymena cells exhibit rapid movement, swimming randomly in the medium. Occasionally, two cells may come into contact, but they quickly separate instead of sticking together. Even newly formed loose pairs often become separated. As a result, one cell can come into contact with numerous others and stimulate them. We have clarified this aspect in the revised version (line 50-51, 57).
Abstract and introduction: Sexes are not mating types. In general, mating types refer to systems in which there is no obvious asymmetry between the gametes, beyond the compatibility system. When there is a physiological difference such as size or motility, sexes are used. This distinction is of importance because in many species mating types and sexes can occur together, with each sex being able to have either (when two) or multiple mating types. An example are SI in angiosperms as used as an example by the authors or mating types in filamentous fungi. See Billiard et al. 2011 [PMID: 21489122] for a good explanation and argumentation for the importance of making this distinction.
We have clarified the expression in the revised version (line 20, 38, 40, 45).
Recommendations for the authors:
Reviewer #1:
I really enjoyed reading this manuscript and I think a few tweaks in the writing/data presentation could greatly improve the experience for the reader:
(1) The information about your previous work in identifying downstream proteins CDK19, CYC9, and CIP1 (lines 170-173) could be directly presented in the introduction.
We have moved this information in the introduction in the revised version (line 74-77).
(2) For a reader who is not familiar with Tetrahymena, a few more details on how reporter and knock-out lines are generated would be beneficial.
We introduced the knock-out method in Figure 2 – figure supplement 1B, HA-tag method in Figure 3A, and MTB2-eGFP construction method in Figure 4E. In addition, we introduced how co-stimulation markers observed in Materials and Methods (line 404-410)
(3) Figures 5 and 6: clarify the types of pairing and treatments that were done directly in the figure (eg. adding additional labels). As of now, it is necessary to go through the text and legend to try and understand in detail what was done.
Cell types and treatments were directly introduced in the revised figure (Figure 5 and 6).
(4) The logical transition in lines 136-142 is hard to follow.
We rewrote this paragraph in the revised version (lines 143-156). Additionally, we added a figure to illustrate the theoretical mating-type recognition model between WT cells and ΔCDK19, ΔCYC9 cells, MTAxc, MTBxc proteins, and ΔMTA, ΔMTB cells (Figure 2 – figure supplement 1D-G).
(5) Lines 191-196: the fact that cells expressing multiple mating types can self goes against an active self-rejection system - if this is the case there should be self-rejection among all expressed mating types. Unless non-self recognition is an active process and self-recognition is simply the absence of non-self recognition. The authors briefly mention this in lines 263-265, but it would be interesting to expand and clarify this.
We appreciate that Reviewer #1 notice the interesting selfing phenotype of the MTB2-eGFP (MTVI background) strain. We further discussed it in the revised manuscript (line 298-306).
(6) The authors briefly mention the possibility of different mating types using different recognition mechanisms (lines 255-260), based on the big differences in the size of the mating-specific region of MT proteins. Following this and the weakness nr. 2, I think it would be pertinent to gather and present more information on the properties and structures of the mating-type specific regions of MT proteins. Simple in silico analysis of motifs, structure, etc. could help clarify the role of these regions. It seems more parsimonious that MT proteins would have variable mating type specific regions that account for the recognition of the different mating types, and conserved cytoplasmic functions that could trigger a single downstream signaling cascade. It would be interesting to know the authors' opinion on this.
We are very grateful for this suggestion. Actually, we are currently working on determining the 3D structure of MTRC. The Alphafold2 prediction indicates that the MT-specific region is comprised of seven global β-sheets, resembling the structure of immunoglobulins (Ig). Our most recent cryo-EM results have revealed a ~15Å structure, aligning well with the prediction. However, the main challenge lies in the low expression levels, both in Tetrahymena and insect/mammal cells. We anticipate obtaining more detailed results soon. Therefore, we prefer to present the MT recognition model with robust experimental evidence in the future, and didn’t discuss too much on this aspect in the current manuscript.
(7) Adding a figure including a proposed model, as well as expanding the discussion on the points presented as "weaknesses" would help clarify the ideas/hypothesis on how the mating recognition works. I think this would really elevate the paper and help highlight the results.
We added a figure to introduce the model and the weaknesses in the revised version (Figure 7, line 656-665).
(8) Line 202-203: It is far-fetched to infer subcellular localization based on the data presented here, couterstaining with other dyes and antibodies specific to certain cell components, as well as negative control images, are required.
Thanks for the suggestion. We attempted to stain cell components using various dyes and antibodies. Unfortunately, we found that cell surface and cilia (especially oral cilia) is very easy to give a false positive signal. We think this issue seriously affects the credibility of the results. It may seem like splitting hairs, but we are trying to be precise.
Meanwhile, we still believe the mating-type proteins localizes to cell surface because MTA-HA is identified in the isolated cell surface proteins.
Regarding negative control, as shown in Fig. 4G, where a MTB2-eGFP cell is pairing with a WT cell, no GFP signal is observed in the WT cell.
(9) Lines 131: clarify the sentence - expression of Con-A receptors requires both MTA and MTB (MTA to receive the signal).
We modified the sentence in the revised version (line 139-140).
Reviewer #2:
Minor points.
(1) Line 194-196. Why are these cells able to self?
These cells able to self may because the MTRC contain heterotypic mating-type proteins (MTA6 and MTB2), which activate mating when they interact with another heterotypic MTRC (line 207-208).
(2) Line 232. What do the authors mean by the term synergistic effect here? Definition and statistics?
Sorry about the confusion. The synergistic effect refers to the effect of MTAxc and MTBxc become stronger when using together. We clarified it in the revised version (line 232).
(3) For Figure 4 panel D, are there antibodies that are available as a control for cilia? If so, then blotting this membrane would show that cilia-associated proteins are in the cilia preparation, which is a standard control for sub-cellular fractionation.
Thanks for the suggestion. Unfortunately, we didn’t find a suitable cilia-specific antibody yet. Instead, we employed MS analysis to confirm the presence of cilia proteins in this sample (line 195-196, Figure 4–Source data 1). We also observed the sample under the microscope, which directly revealed the presence of cilia (Figure 4C).
(4) At least one reference cited in the text was not present in the reference list. The authors should go through the references cited to ensure that all have made it into the reference list.
We have checked all the references.
Some minor edits:
(1) MTA and MTB are presented in both roman and italics (e.g. line 209) in the manuscript. Maybe all should be in italics? Or is this a distinction between the gene and the protein?
The italics word (MTA) refers to gene, and non-italics word (MTA) refers to protein.
(2) Line 251. Change "achieving" to "achieve".
We have corrected this word (line 266).
Reviewer #3:
Line 101. It would help to explain this expectation earlier in this paragraph.
We explained the expectation in the revised version (line 92-97, 104-106).
Line 109. How is a co-receptor different from the MTRC complex?
We have rewritten the relevant sentences to enhance clarity (line 116-119). The molecular function of the MTRC complex could involve acting as a co-receptor or recognizer (functioning as both ligand and receptor). Based on the results presented in this section, we propose that MTA and MTB may function as a complex, but the confirmation of this hypothesis (MTRC) is provided in a later section. Therefore, we did not use the term “MTRC” here. These sentences briefly discuss the molecular function of this complex and explain why MTRC does not appear to function as a co-receptor.
Line 251: which "dual approach" is referred to?
Dual approach is referred to both self and non-self recognition. We explained it in the revised version (line 265-266).
Line 258: what "different mechanisms" do the authors have in mind? Why would a different mechanism be expected? The different sizes could have evolved for (coevolutionary?) selection on the same mechanism.
Sorry about the confusion. We clarified it in the revised version (line 269-278).
What we intended to express is that we are uncertain whether the mating-type recognition model we discovered in T. thermophila is applicable to all Tetrahymena species due to significant differences in the length of the mating-type-specific region. We believe it is important to highlight this distinction to avoid potential misinterpretations in future studies involving other Tetrahymena species. At the same time, we look forward to future research that may provide insights into this question.
Fig 2 C&D. Is it correct that these figures show the strains only after 'preincubation'? This is not apparent from the caption of the text. Additionally, the order of the images is very confusing. Write in the figures (so not just in the caption) what the sub-script means.
These panels are re-organized in the revised version (Fig. 2C&D). There are three kinds of pictures: “not incubated”, “WT pre-incubated by mutant” and “mutant pre-incubated by WT”.
The methods used to generate Figure 5 are not clearly described. I understand that the obtained xc proteins were added to the cells, and then washed, after which a test was performed mixing WT-VI and WT-VII cells. Were both cells treated? Or only one of the strains? The explanation for the reused washing medium is not clear and the method is not indicated.
Both cells are treated. More details are provided in the revised manuscript (line 230-231, 633-634, 637-639, Fig. 5). To prepare the starvation medium containing mating-essential factors, cells were starved in fresh starvation medium for ~16 hours. Subsequently, cells were removed by three rounds of centrifugation (1000 g, 3 min) (line 330-332).
In general, the figures are difficult to understand without repeated inquiries in the captions. Give more information in the figures themselves.
More information is introduced in the figure (Fig. 2C, Fig. 3B, Fig. 4A, B, D, Fig. 5 and Fig. 6).
mártires por vocación
tag yourself soy ésta
Author Response
Note to the editor and reviewers.
All the authors would like to thank the editorial team and the two anonymous reviewers for their efforts and thoughtfulness in assessing our manuscript. We very much appreciate it and we all believe that the manuscript has been much improved in addressing the comments and suggestions made.
General considerations on the revised manuscript
We have applied extensive modifications to the manuscript with our main goal being the improvement of clarity. The Introduction has been changed mainly to introduce precisely our terminology and we have stuck to it in the rest of the manuscript. The Results section has been divided up into more defined sections. The discussion has been extensively re-written to improve clarity, following the suggestion of the reviewers. Main figures 1 and 4 have been modified with clearer schematics. Supplementary figures and legends have been modified and several supplementary schematic figures have been added to clearly present our interpretations for various data. We have added a Supplementary Discussion where the most detailed technical parts of our discussion are presented to avoid unnecessarily weighing down the main discussion, where our main conclusions are outlined. We have presented our mass photometry mixing experiment in a new supplementary figure, with detailed explanation. We have also expanded our discussion of in vivo and general relevance of our study.
Response to manuscript evaluation
Our manuscript has been evaluated as a valuable study and presenting solid experimental evidence. We appreciate the recognition of our work.
Two weaknesses were identified by reviewers: 1) our experiments do not completely exclude the possibility of an alternative nucleophile. This relates to the evaluation of our experimental evidence. 2) Our study does not address the in vivo relevance of the interface swapping phenomenon, which relate to the value of the study for the community.
Response to the evaluation of experimental evidence (Weakness #1):
We argued in the original manuscript that we have excluded completely the presence of an alternative nucleophile. This conclusion is based on a series of experiments which were presented in the originally submitted manuscript. These experiments are not discussed by the reviewers in relation to this main conclusion and therefore we suggest that they have not been properly evaluated. We believe our conclusion to be appropriately supported by these data (see our response to reviewer #1). In addition, the criticism of our gel-filtration data by reviewer #2 was based on a misinterpretation of Supplementary figure 1 b. We accept of course that the way the data was presented could be misleading and we assume responsibility for this. We have attempted to correct this by changing the main text and the figures legends and annotation. In conclusion, we believe that the evaluation of experimental evidence as presented in the revised manuscript could be upgraded to “convincing”.
Response to our study general relevance evaluation (weakness #2):
We agree with both reviewers about the in vivo relevance of our observation being an important question, not addressed so far. Indeed, the value of our study would be greatly increased by in vivo data and be of interest to a wider audience. However, we would like to argue that our study would interest a wider audience than initially stated for the following reasons: 1) Our study is the first evidence of interface swapping in vitro and will constitute a base to investigate this phenomenon both in vivo and in vitro. It will therefore interest a wide audience due to the potential involvement of interface swapping in a wide range of processes, such as recombination, evolution, and drug targeting (see also below). 2) DNA cleavage is the central mode of action of antibiotics targeting bacterial type II topoisomerases (i.e. topoisomerases “poisons”). This already established target is one of the few having produced new scaffolds and too few new antibacterial are in production to fulfill medical needs. The role of interface stability is also emerging as a modulator of the efficiency of topoisomerase poisons. See for instance (Germe, Voros et al. 2018, Bandak, Blower et al. 2023). By shedding light on interface dynamics, our study will be of interest to scientist interested in the development of these drugs. In addition, the heterodimer system can potentially produce detailed mechanistic information (Gubaev, Weidlich et al. 2016, Hartmann, Gubaev et al. 2017, Stelljes, Weidlich et al. 2018) not only on gyrase but also on other, dimeric type II topoisomerases or even other dimeric enzyme in general. We have amended the manuscript to make these points clearer. Therefore, we believe that the evaluation of the revised manuscript’s relevance could be upgraded to “important”.
Point-by-point response to the reviewer
Reviewer #1 (Public Review):
Germe and colleagues have investigated the mode of action of bacterial DNA gyrase, a tetrameric GyrA2GyrB2 complex that catalyses ATP-dependent DNA supercoiling. The accepted mechanism is that the enzyme passes a DNA segment through a reversible double-stranded DNA break formed by two catalytic Tyr residues-one from each GyrA subunit. The present study sought to understand an intriguing earlier observation that gyrase with a single catalytic tyrosine that cleaves a single strand of DNA, nonetheless has DNA supercoiling activity, a finding that led to the suggestion that gyrase acts via a nicking closing mechanism. Germe et al used bacterial co-expression to make the wild-type and mutant heterodimeric BA(fused). A complexes with only one catalytic tyrosine. Whether the Tyr mutation was on the A side or BA fusion side, both complexes plus GyrB reconstituted fluoroquinolone-stabilized double-stranded DNA cleavage and DNA supercoiling. This indicates that the preparations of these complexes sustain double strand DNA passage. Of possible explanations, contamination of heterodimeric complexes or GyrB with GyrA dimers was ruled out by the meticulous prior analysis of the proteins on native Page gels, by analytical gel filtration and by mass photometry. Involvement of an alternative nucleophile on the Tyr-mutated protein was ruled unlikely by mutagenesis studies focused on the catalytic ArgTyrThr triad of residues. Instead, results of the present study favour a third explanation wherein double-strand DNA breakage arises as a consequence of subunit (or interface/domain) exchange. The authors showed that although subunits in the GyrA dimer were thought to be tightly associated, addition of GyrB to heterodimers with one catalytic tyrosine stimulates rapid DNA-dependent subunit or interface exchange to generate complexes with two catalytic tyrosines capable of double-stranded DNA breakage. Subunit exchange between complexes is facilitated by DNA bending and wrapping by gyrase, by the ability of both GyrA and GyrB to form higher order aggregates and by dense packing of gyrase complexes on DNA. By addressing a puzzling paradox, this study provides support for the accepted double strand break (strand passage) mechanism of gyrase and opens new insights on subunit exchange that may have biological significance in promoting DNA recombination and genome evolution.
The conclusions of the work are mostly well supported by the experimental data.
Strengths:
The study examines a fundamental biological question, namely the mechanism of DNA gyrase, an essential and ubiquitous enzyme in bacteria, and the target of fluoroquinolone antimicrobial agents.
The experiments have been carefully done and the analysis of their outcomes is comprehensive, thoughtful and considered.
The work uses an array of complementary techniques to characterize preparations of GyrA, GyrB and various gyrase complexes. In this regard, mass photometry seems particularly useful. Analysis reveals that purified GyrA and GyrB can each form multimeric complexes and highlights the complexities involved in investigating the gyrase system.
The various possible explanations for the double-strand DNA breakage by gyrase heterodimers with a single catalytic tyrosine are considered and addressed by appropriate experiments.
The study highlights the potential biological importance of interactions between gyrase complexes through domain-or subunit-exchange
We thank the reviewer for their support, effort, and comments. The above is a great summary.
Weaknesses:
The mutagenesis experiments described do not fully eliminate the perhaps unlikely participation of an alternative nucleophile.
We agree that the mutagenesis experiment on its own does not fully eliminate the possibility of an alternative nucleophile. The number of residues mutated is limited, and therefore it is possible we have missed a putative alternative nucleophile.
However, we have other data and experiments supporting the conclusion that no alternative nucleophile exists. Therefore, we want to stress that our conclusion that no such alternative exist is based on these extra data. These data and experiments are not discussed by either reviewer despite being present in the original manuscript. This puzzled us and we have modified the manuscript and the figures in the hope that they, and their significance, would not be missed.
Briefly:
1) We have performed cleavage-based labeling of the nucleophile responsible for cleavage. This experiment is depicted in Figure 4. The nucleophilic activity of the residue involved results in covalent link between the polypeptide (that includes the residue) and radiolabeled DNA. Therefore, a polypeptide that includes an active nucleophile will be radiolabeled and visible, whereas a polypeptide that is missing an active nucleophile will remain unlabeled and invisible. We can distinguish the BA and the A polypeptide from their size. In the case of the BA.A complex both the BA polypetide and the A polypetide are radiolabeled and therefore both have an active nucleophile. In the case of the BAF.A complex, the unmutated A polypeptide is labeled, meaning that a nucleophile is still active. In contrast, the BAF polypeptide shows no detectable labeling. This result means that removing the hydroxyl group from the catalytic tyrosine abolishes any protein-DNA covalent link, suggesting that no other nucleophile from the BA polypetidic chain can substitute for the catalytic tyrosine hydroxyl group. This experiment excludes the possibility of an alternative nucleophile coming from the polypeptidic chain of either GyrA or GyrB. This experiment, described in figure 4, is not discussed by the reviewer. This experiment is similar in principle to early experiments identifying catalytic tyrosine in topoisomerases. See for instance, (Shuman, Kane et al. 1989).
2) The experiment above does not exclude a nucleophile coming from the solvent. To exclude this possibility, we have used T5 exonuclease (which needs a free 5’ DNA end to digest) and ExoIII (which need a free 3’ DNA end to digest). We have shown the reconstituted cleavage is not sensitive to T5 and sensitive to ExoIII. This shows that the 5’ end of the cleaved sites are protected by a bulky polypeptide impairing T5 activity, which is active in our reaction as shown by the digestion of a control DNA fragment. This experiment shows that the reconstituted cleavage is very unlikely to come from a small nucleotide potentially provided by the solvent. This experiment is described in the main text and the results are shown in supplementary figure 5. It is not mentioned by either reviewer.
3) Finally, we would like to emphasize our experiment comparing the BAF.A59 to BALLL.A59. The BALLL.A59 complex displays increased cleavage compared to BAF.A59. If this increased cleavage was due to an alternative nucleophile on the BALLL side, we would expect an accompanying increase in supercoiling activity since the BALLL.A59 possesses one CTD, which is sufficient for supercoiling. The fact that no increased supercoiling activity is observed strongly suggests subunit exchange reconstituting an A59 dimer, inactive for supercoiling but active for cleavage. We believe this somewhat complex observation to be quite significant and we have attempted to clarify the manuscript and discuss its full significance in several places.
Reviewer #1 (Recommendations For The Authors):
An interesting paper on DNA gyrase that explains a puzzling paradox in terms of the double-strand break mechanism.
Major points
1) The authors consider several mechanisms that could potentially explain their data. On page 15, the authors present the evidence against the nicking closing mechanism proposed by Gubaev et al. Throughout the manuscript, they indicate where their experimental results agree with this earlier work but should also indicate and account for differences. For example, Gubaev et al describe cross linking experiments that they claim rule out subunit exchange. These aspects should be clearly explained.
Thank you for the suggestion. We have re-written the discussion to address this point. We are extensively discussing experiments from (Gubaev, Weidlich et al. 2016), and offer our interpretation of apparently conflicting results. We suggest that their experiments are basically consistent with our data when correctly interpreted. To keep the main manuscript clear, we have added a supplementary discussion where experiments from (Gubaev, Weidlich et al. 2016) are discussed further in relation to our data.
2) Page 9. The experiments done to rule out the perhaps unlikely alternative nucleophile hypothesis relate to the possible role of the Arg and Threonine of the RYT triad. These residues are close to the DNA and therefore are prime candidates and attractive targets for mutagenesis. However, strictly speaking, the mutant enzyme data presented do not rule all possibilities. For example, Serine is often the nucleophile used by resolvases to effect DNA recombination via subunit exchange. The ideal experiment to rule out/rule in other nucleophiles would be to identify the residue(s) that become attached to DNA in the cleavage reaction.
Please see above. We have effectively ruled an alternative nucleophile with our cleavage-based labeling experiment and others that were present and discussed in the original manuscript but were missed. We have modified the manuscript and figures in order to make this point clearer than before.
3) p17. The readout for subunit exchange used by the authors is double-stranded DNA cleavage. Attempts to directly detect the formation of the DNA cleaving complexes GyrA2B2 and (GyrBA)2 (arising from subunit exchange between heterodimers) by mass photometry were not successful. Perhaps FRET would have been another approach to try as it could also detect interface and domain interchanges.
Directly detecting interface exchange directly by proximity experiment would be extremely useful. FRET would have to be done in the BAF.A + GyrB configuration where the amount of interface exchange is important. Now, we do not have the tools to do that and developing them would be outside the scope of the study. We propose cross linking experiment to be done in the future. We argue that the manuscript is convincing without these for now. This will be addressed in the future. This point, and other possible future experiments are now discussed in the discussion section.
4) The underlying canvas of this paper is the strand passage mechanism of gyrase. It would seem appropriate to include the papers first proposing it - Brown P.O and Cozzarelli N.R. (1979) and Mizuuchi K et al (1980).
We very much agree. These papers have now been added in the introduction as appropriate, highlighting the relationship between double-strand cleavage and the strand-passage mechanism.
5) Figure 1. The quality of the insets is poor. It is difficult to pick out the key catalytic residues and their disposition vis-a-vis DNA.
We agree, Figure 1 has been re-done and the schematic theme has been harmonized throughout the whole manuscript. We very much hope that clarity has improved. Thank you for the suggestion.
6) The experimental work is a very detailed analysis of a specific feature of engineered gyrase heterodimers. Making the work accessible to the general reader will be important. Using shorter paragraphs each with a specific theme might help. In particular, the second paragraph of the Results on p7, the section on p9 and bottom of p11, p13 and the first paragraph of the Discussion on p14 are each a page or more long. A shorter manuscript that avoids overinterpretation of the smaller details would also help.
We agree. We have now split long paragraphs into individual sections, with titles, in the Results. This structure is recapitulated at the beginning of the discussion, and we have split the discussion into shorter paragraphs, each with a unique point being made.
7) The impact of the Gubaev et al (2016) paper for the field in general, and as the catalyst for the present work should be better documented. Mention of this earlier paper and its significance at the beginning of the Abstract and elsewhere e.g in the Introduction might also help with a more logical organization of the current findings and result in a shorter paper (which would be easier to read).
We have added a reference to (Gubaev, Weidlich et al. 2016) in the abstract and have expanded our introduction
Minor points
1) Legends for Figs 2 and 6; Supplementary Figs 1 and 8. The designation of subfigures as a, b, c, d , e etc appears to be incorrect. Check throughout and in the text.
The manuscript has been checked for such errors.
2) Figure 2, and first paragraph p8. Peaks in Fig 2c should be labelled to facilitate discussion on p8.
Agreed, this has been done.
3) Supplementary Fig 4 and elsewhere in the manuscript. A variety of notations are used to denote phenylalanine mutants e.g. AsubscriptF, AsuperscriptF and AF. Check and use one format throughout.
Done
4) Figures showing gels include the label '+EtBr, +cipro'. This is somewhat confusing because EtBr was contained in the gel (not the samples) whereas cipro was included in the reaction. Modify or describe in the legend..
We have re-written the figure legend.
5) Supplementary Fig 4b describes a small effect on the ratio of linear to nicked DNA for the triple LLL mutant. Is this significant? How many times was the measurement made?
This has been addressed in the original manuscript in the supplementary data. In term of quantification, the experiment has been done 3 times for each prep, with the same GyrB prep and concentration. The standard error is displayed on the figure. This result is very reproducible and have been reproduced more than 3 times. No LLL cleavage assay showed more single-strand than double-strand cleavage. For the phenylalanine mutant, no cleavage assay showed more double-strand than single-strand cleavage.
6) Supplementary Fig 5 legend. Should 'L' read 'size markers' (and give their sizes)?
Yes indeed, we have modified the figure to clarify.
7) p11 line 5. Is this statement correct?
Yes, it is correct. Although we hope we are on the same line. When the Tyrosine is mutated on one side only of the heterodimer, both single- and double-strand cleavage are protected from T5 exonuclease digestion.
8) 12 last line should read...and supercoiling activity (not shown)..were
Thank you, done.
There are a number of typos throughout the text, for example:
Page 3 line..Difficult to conclude...what?
Page 3 para 3...Lopez....and Blazquez
We have corrected these typos and checked the whole manuscript.
Reviewer #2 (Public Review):
DNA gyrase is an essential enzyme in bacteria that regulates DNA topology and has the unique property to introduce negative supercoils into DNA. This enzyme contains 2 subunits GyrA and GyrB, which forms an A2B2 heterotetramer that associates with DNA and hydrolyzes ATP. The molecular structure of the A2B2 assembly is composed of 3 dimeric interfaces, called gates, which allow the cleavage and transport of DNA double stranded molecules through the gates, in order to perform DNA topology simplification. The article by Germe et al. questions the existence and possible mechanism for subunit exchange in the bacterial DNA gyrase complex.
The complexes are purified as a dimer of GyrA and a fusion of GyrB and GyrA (GyrBA), encoded by different plasmids, to allow the introduction of targeted mutations on one side only of the complex. The conclusion drawn by the authors is that subunit exchange does happen, favored by DNA binding and wrapping. They propose that the accumulation of gyrase in higher-order oligomers can favor rapid subunit exchange between two active gyrase complexes brought into proximity.
The authors are also debating the conclusions of a previous article by Gubaev, Weidlich et al 2016 (https://doi.org/10.1093/nar/gkw740). Gubaev et al. originally used this strategy of complex reconstitution to propose a nicking-closing mechanism for the introduction of negative supercoils by DNA gyrase, an alternative mechanism that precludes DNA strand passage, previously established in the field. Germe et al. incriminate in this earlier study the potential subunit swapping of the recombinant protein with the endogenous enzyme, that would be responsible for the detected negative supercoiling activity.
Accordingly, the authors also conclude that they cannot completely exclude the presence of endogenous subunits in their samples as well.
Strengths
The mix of gyrase subunits is plausible, this mechanism has been suggested by Ideka et al, 2004 and also for the human Top2 isoforms with the formation of Top2a/Top2b hybrids being identified in HeLa cells (doi: 10.1073/pnas.93.16.8288).
Germe et al have used extensive and solid biochemical experiments, together with thorough experimental controls, involving :
the purification of gyrase subunits including mutants with domain deletion, subunit fusion or point mutations.
DNA relaxation, cleavage and supercoiling assays
biophysical characterization in solution (size exclusion chromatography, mass photometry, mass spectrometry)
Together the combination of experimental approaches provides solid evidence for subunit swapping in gyrase in vitro, despite the technical limitations of standard biochemistry applied to such a complex macromolecule.
We thank the reviewer for their supportive and considered comments.
Weaknesses
The conclusions of this study could be strengthened by in vivo data to identify subunit swapping in the bacteria, as proposed by Ideka et al, 2004. Indeed, if shown in vivo, together with this biochemical evidence, this mechanism could have a substantial impact on our understanding of bacterial physiology and resistance to drugs.
Thank you for this comment. Indeed, whether this interface exchange can happen in vivo and lead to recombination is a very important question. However, we believe that this is outside the scope of this study simply because of the amount of work one can fit into one paper. Proving that interface exchange can happen in vitro has already necessitated a number of non-trivial experiments and likewise investigating interface exchange in vivo will require a careful, long-term study (see our reply to reviewer #2 comment, who also raised this point). We can’t address it with one additional experiment with the tools we have. However, we very much hope to do it in the future.
Reviewer #2 (Recommendations For The Authors):
Specific questions and comments for the authors:
1) Complex identification during purification
The statement line 236-237 that "Our heterodimer preparation showed a single-peak on a gel-filtration column, distinct from the GyrA dimer peak" is not entirely clear. In Fig supp 1 b, how can the authors conclude from the superose 6 that GyrBA is separated from the GyrA dimer? Since they seem close in size 160/180kDa, they are unlikely to be well separated in a superose 6 gel filtration column. The SDS-PAGE seems to show both species in the same fractions #15-17 therefore it would not be possible to distinguish GyrBA. A from A2.
There appears to be some confusion about what Supp Fig. 1b shows. First, in all our gel filtration conditions both GyrBA and GyrA can’t exist as monomers at a significant concentration. Therefore, we can never observe the GyrBA monomer on a gel filtration column. Supp Fig. 1b shows the gel filtration profile of the BA.A heterodimer only. This is the output of the last, polishing step in the reaction. We analyze these results using SDS-PAGE. Therefore, the BA.A heterodimer will be denatured and separated into 2 polypeptides: GyrBA and GyrA, which migrates according to their size in an SDS-PAGE and forms two bands. These two bands do not represent two separate species in solution. They represent the separation of one species only, the BA.A heterodimer into its two, denatured, subunits: GyrA and GyrBA. We do not conclude from Supp Fig. 1 as a whole that GyrBA and the GyrA dimer are well separated, and this is not stated in the manuscript. We conclude that the BA.A dimer is fairly well separated from the GyrA dimer. They have significant different size (~260 kDa and ~180 kDa respectively) and form different peaks on a gel filtration column. The BA.A heterodimer has a GyrA subunit and therefore will shows a GyrA band on an SDS-PAGE, like the GyrA dimers but the two are obviously distinct in their quaternary structure. We are hoping that our new schematics and re-write of some of the results and figure legends will clarify this.
Panel 6 shows a different elution volume for the 2 species BA.A and A2 on an analytical S200 column, which appears better at separating the complexes in this size range.
Did the authors consider using a S200 column instead of superose 6 for the sample preparation, to optimize the separation of GyrBA. A from A2?
This is not a necessarily true statement (see above). We have not run the GyrA dimer on a Superose 6 column. The analysis was done on an s200 because extensive data for the GyrA dimer was already available with this, already calibrated column. We do not expect the Superose 6 to be worse in this size range. In fact, it might even be better. The Superose 6 profile in Supp. Fig. 1b shows BA.A only and no GyrA dimer. We have clarified the annotations in the figure to make this clearer.
Regarding the analytical gel filtration experiment, there is however an overlap in the elution volume in the analytical column, therefore how can the authors ensure there is no excess free A2 complex in the GyrBA. A sample?
Indeed, there is an overlap, but we argue that it is overstated. The important part of the overlap is where the maximum height of the GyrA peak is positioned compared to the BA.A trace, not where the traces intersect. This overlap is minimal. If a contaminating GyrA peak was hidden in the BA.A peak, it would have to be at least 10 times less intense than the BA.A peak. Since BA.A and GyrA dimer have roughly the same extinction coefficient, this means that a contamination would detectable at 10 % or even less. Our mass photometry further excludes such contamination.
Alternatively, the addition of a larger (cleavable) tag at the C-terminal end of the BA construct (therefore not disturbing dimer association) could allow to better distinguish the 2 populations already at the size exclusion step.
This is true and could allow cleaner purification. There are also other ways to achieve cleaner purification, like adding a secondary tag. However, like we argue in the manuscript, our contaminations are already minimal. It is questionable what benefits could be gained in changing the protocol. We also argue that the tandem tag method does not completely exclude contamination (Supplementary Discussion) and therefore we are not sure if this would be worth the time and expenditure.
2) GyrA and GyrB Oligomers:
In the mass photometry experiment, the authors explain that the low concentration of the proteins promotes dissociation of GyrA dimers, hence the detection of GyrA monomers instead of GyrA dimers, which are also detected in the GyrBA.A sample.
However, it cannot be concluded that the GyrA dimer is not formed in the condition of the gel filtration chromatography, at higher concentration.
In our mass photometry experiment, The BA.A sample is not as diluted as the GyrA dimer and much closer to our experimental condition. Since we have calculated the dissociation constant, we can calculate the expected level of dissociation (or reassociation). The level of dissociation is minimal in these conditions. If some dissociation is expected from the BA.A heterodimers, a very low amount of GyrBA monomer should also be present and yet they are not observed. We presume that it is because mass photometry is much more sensitive to GyrA (see our mixing mass photometry experiment that we have added). If the GyrA would reassociate at higher concentration, it would do so either with itself (forming a GyrA dimer) or with the GyrBA monomer, reforming the heterodimer. Assuming both GyrA dimer and heterodimer have the same dissociation constant, roughly one third of the GyrA monomer would reassociate with themselves. Assuming even complete reassociation of the GyrA dimer, this would leave only GyrA dimer accounting for 2% of the prep.
Another interpretation would be to assume that GyrBA monomers are not present at all and that GyrA monomer are reassociating only with themselves. This is not valid because of the following thermodynamic reason:
Since the profile for the GyrA dimer are collected at equilibrium, we should expect a ratio between GyrA monomer and dimers that follow the dissociation constant. In other words, if the GyrA monomer were in equilibrium with GyrA dimer we should expect a much higher dimer concentration already as the GyrA monomers are not as dilute. We do not observe a GyrA dimer peak in the BA.A profile, even though we can detect a low amount of GyrA dimer mixed with BA.A. Therefore, we conclude that the observed GyrA monomer must be in equilibrium with another dimerization partner, which is most probably the GyrBA monomer (see above). Therefore, only a minimal amount of GyrA dimer is expected to be formed at higher concentration by direct reassociation. This could probably increase if we let this solution-based exchange carry on for a long time at dissociation equilibrium. We have actually shown that this solution-based exchange is very slow and take several days because of the low dissociation at equilibrium.
The mass spectrometry analysis in Fig 2 confirms the presence of (monomeric) GyrA in the sample, despite different experimental conditions.
The concentration of heterodimer in the mass spectrometry experiment is actually higher than in the mass photometry experiment. This shows that self-reassociation of the GyrA monomer as suggested above is undetectable with mass spectrometry at higher concentration.
We considered that the “GyrA monomer” peak could be a contaminating GyrB monomer, which is ~90 kDa, which would explain the lack of reassociation. However, the mass spectrometry peak shows precisely the expected molecular weight of GyrA so we interpret this peak as arising from very limited dissociation of the BA.A heterodimer. The reassociation is limited at high concentration due simply to the fact that the difference in concentration between the mass photometry and our other experimental conditions is not that high. The GyrA dimer had to be diluted 400 times to see significant dissociation and yet even at this very low concentration the dissociation is far from complete.
Our general conclusions on the couple of point above is that we cannot completely exclude the presence of GyrA dimers being present, although they are undetectable in our working conditions either by mass photometry (lower concentration), Mass spectrometry (higher concentration) and even gel filtration (even higher concentration, see above). For the mass photometry, we have established that our detection threshold for a contamination is very low (see our mixing experiment).
Figure 2A: the authors state in the introduction that GyrB is a monomer in solution and then explain that the upper bands in the native gel are multimer of GyrB. Could the authors comment and provide the size exclusion profile of the Gyr B purification?
We have expanded our discussion of this. However, we have not been successful in collecting a gel filtration profile for GyrB. This is likely due to excessive oligomerization at the concentration we are using for gel filtration. We suggest that our mass photometry and Blue-Native PAGE experiment shows clearly that GyrB can be detected as a monomer in solution at the appropriate dilution. However, GyrB tends to oligomerize in a regular fashion (Consider especially Supp Fig. 8a), which suggest that it could align heterodimers on DNA in a linear, regular orientation. We have added a discussion of this.
Together the relevance of the oligomeric state of purified GyrA or GyrB should be clarified, relative to their role in subunit swapping.
We have added explanation in our discussion, while also trying to not be too speculative. Basically, we believe that GyrB oligomerization is likely to be involved. It is difficult to conclude for GyrA since no experiment has allowed us to test it. Therefore, the role of GyrA oligomerization, if any, is unclear. The GyrA tetramer is very prominent though and forms very easily. GyrB on the contrary forms longer oligomers more readily than GyrA and we surmise that this would help interface exchange. However, the structure of these GyrA and GyrB oligomers is not clear, which make it difficult to go beyond speculation on this. It would be a very interesting experiment if we were able to suppress GyrB oligomerization whilst conserving its ability to promote strand-passage and cleavage. Same goes for GyrA. Unfortunately, we are unable to do that at this time.
4) Subunit exchange
Line 320: the concept of subunit exchange in this context should be clearly explained. If one understands correctly, the authors mean that the BAF polypeptide, part of the BAF.A complex, could be replaced by a combination of B+A therefore forming a fully functional WT A2B2 gyrase complex.
Thank you for the suggestion. We have harmonized and clearly defined our terminology for interface swapping and subunit exchange in the introduction and attempted to be much more rigorous when referring to it.
A great effort has been done in this study to explain all the pros and cons of the experimental design but the length of the explanations may prevent readers outside of the field to fully appreciate the conclusions. This article would benefit from the addition of a few schematics to summarize the working hypothesis.
Thanks for the suggestion. We have added a series of schematics to illustrate our interpretation for each construct. As mentioned above the terminology has been more rigorously defined and updated throughout the manuscript.
5) Presence of endogenous GyrA
Line 419-425: it is quite difficult to follow the explanations regarding the possible contamination of the sample by endogenous GyrA.
Maybe these points should rather be addressed in the discussion, when debating the conclusions of Gubaev et al.
We agree. We have re-organized the Discussion doing just that. We added a Supplementary Discussion in which we further discuss the contamination problem in relation to (Gubaev, Weidlich et al. 2016).
Production of the subunits in another (non bacterial) expression system or a cell free system may prevent the association of endogenous protein.
Absolutely. We are planning on addressing this in the future, using the yeast expression system.
6) Mechanism for subunit swapping
Lines 588-595: As described by the authors the BA fusion shows decreased activity when compared with the WT probably due to limited conformational flexibility in absence of an additional linker sequence between the fused subunits.
The affinity of BA for A may possibly be reduced compared to the free A2B2 complex, due to a relative stiffness of the fusion upon full association with a free B subunit, as rightfully pointed by the authors.
If subunit exchange do happen in vitro, at least in the conditions of this study, the authors could assess the affinity of BA for A, when compared to the association of free B and A subunits
Experiments using analytical ultracentrifugation or surface plasmon resonance (SPR) may allow to determine the relative affinity of the BA +(A+B) compared to the A2B2 complex. This could be done also for the BALLL mutant and association with A59.
It would be extremely useful to measure the affinity of BA for A. However, this is difficult because of the high affinity of the interface. To measure a dissociation constant, one has to be able to measure the concentration of the monomer and the dimer at equilibrium. Because of this, the complex must be diluted enough to see any dissociation, making detection difficult. In practice, this also means that we cannot purify monomeric versions of these subunits. We therefore can’t perform “on-rate” study on an SPR surface, which would require flowing monomers on its partner subunit tethered to the SPR surface. However, we could perform “off-rate” studies, but the dissociation time is likely to be very long, making the measurement difficult. We have not tried it though, and it could turn out to be informative. An analysis of antibodies off-rate done in the past could provide a guideline for us to perform this experiment. Analytical ultracentrifugation is an excellent technique and could in theory provide information. In practice however it would be still necessary to dilute the complex enough to obtain significant dissociation at equilibrium, making detection difficult. As far as we are aware, analytical ultracentrifugation rely on UV absorbance for protein detection and therefore we probably would not detect our material at the necessary dilution. We are however open-minded about technique with very sensitive detection methods that could be used.
9) In vivo relevance
The study does not conclude on the subunits exchange in vivo, which have been suggested by earlier studies by Ikeda et al. To elaborate further on the relevance of such mechanism in the bacteria, experiments involving the fluorescent labeling of endogenous / exogenous mutant subunits may be required to provide further information on this phenomenon.
We completely agree that the in vivo relevance of such phenomena is the central question. Addressing this directly is not trivial though. Expressing both BA and A in vivo will results in random partnering and lead to a mix of dimers: A2 (1/4), BA2(1/4) and BA.A (1/2), assuming equal interface affinity. Therefore, to see subunit exchange in the same way as in vitro, one would have to get rid of the BA2 and A2 dimer together, or the BA.A dimer only. Our initial strategy to do that would be to engineer a specific dimer as being uniquely targeted for degradation. This could allow us to “get rid” of for instance the BA.A dimer. Subsequently, we would turn off the degradation and translation together and observe the rate of subunit exchange. This is not trivial though and would be the subject of a further study.
10) Figure 3: I guess the "intact" label refers to the supercoiled DNA (SC) ? It also appears as "uncleaved" in supp Figure 6. The same label for this topoisomer should be used throughout.
Thank you for pointing that out. It has now been corrected.
Bandak, A. F., T. R. Blower, K. C. Nitiss, R. Gupta, A. Y. Lau, R. Guha, J. L. Nitiss and J. M. Berger (2023). "Naturally mutagenic sequence diversity in a human type II topoisomerase." Proceedings of the National Academy of Sciences 120(28).
Germe, T., J. Voros, F. Jeannot, T. Taillier, R. A. Stavenger, E. Bacque, A. Maxwell and B. D. Bax (2018). "A new class of antibacterials, the imidazopyrazinones, reveal structural transitions involved in DNA gyrase poisoning and mechanisms of resistance." Nucleic Acids Res.
Gubaev, A., D. Weidlich and D. Klostermeier (2016). "DNA gyrase with a single catalytic tyrosine can catalyze DNA supercoiling by a nicking-closing mechanism." Nucleic Acids Res 44(21): 10354-10366.
Hartmann, S., A. Gubaev and D. Klostermeier (2017). "Binding and Hydrolysis of a Single ATP Is Sufficient for N-Gate Closure and DNA Supercoiling by Gyrase." J Mol Biol 429(23): 3717-3729. Shuman, S., E. M. Kane and S. G. Morham (1989). "Mapping the active-site tyrosine of vaccinia virus DNA topoisomerase I." Proc Natl Acad Sci U S A 86(24): 9793-9797.
Stelljes, J. T., D. Weidlich, A. Gubaev and D. Klostermeier (2018). "Gyrase containing a single C-terminal domain catalyzes negative supercoiling of DNA by decreasing the linking number in steps of two." Nucleic Acids Res.
eLife assessment
This study presents two useful new mouse models that individually tag proteins from the SMAD family to identify distinct roles during early pregnancy. Solid evidence is provided that SMAD1 and SMAD5 target many of the same genomic regions as each other and the progesterone receptor. Given the broad effect of these signaling pathways in multiple systems, these new tools will most likely interest readers across biological disciplines.
"bevor die AFD die verfassung abschafft, müssen wir die verfassung abschaffen" -- klingt doch logisch...<br /> die sind einfach voll hängen geblieben in ihrer opferrolle, und erfinden jeden tag neue strohmänner, neue false flag attacks, neue lügen... 9/11 ist quasi zum dauerzustand geworden<br /> aber die grundprobleme bleiben: pazifismus und übervölkerung
Reviewer #2 (Public Review):
Summary:<br /> Verma et al. provide a short technical report showing that endogenously tagged dynein and dynactin molecules localize to growing microtubule plus-ends and also move processively along microtubules in cells. The data are convincing, and the imaging and movies very nicely demonstrate their claims. I don't have any large technical concerns about the work. It is perhaps not surprising that dynein-dynactin complexes behave this way in cells due to other reports on the topic, but the current data are among some of the nicest direct demonstrations of this phenomenon. It may be somewhat controversial since a separate group has reported that dynein does not move processively in mammalian cells (https://www.biorxiv.org/content/10.1101/2021.04.05.438428v3). Because of this, it might be nice for the authors to comment on this discrepancy in the field, although the aforementioned work is still in pre-print form.
Strengths:<br /> Using state-of-the-art methods to endogenously tag dynein/dynactin subunits and performing live-cell imaging is convincing and useful for the field.
Weaknesses:<br /> The claims are perhaps not surprising or novel given the extensive data already published in the field. However, there aren't many similar studies using endogenously tagged subunits to date.
Reviewer #3 (Public Review):
Summary:<br /> In this manuscript, Verma et al. set out to visualize cytoplasmic dynein in living cells and describe their behaviour. They first generated heterozygous CRISPR-Cas9 knock-ins of DHC1 and p50 subunit of dynactin and used spinning disk confocal microscopy and TIRF microscopy to visualize these EGFP-tagged molecules. They describe robust localization and movement of DHC and p50 at the plus tips of MTs, which was abrogated using SiR tubulin to visualize the pool of DHC and p50 on the MTs. These DHC and p50 punctae on the MTs showed similar, highly processive movement on MTs. Based on comparison to inducible EGFP-tagged kinesin-1 intensity in Drosophila S2 cells, the authors concluded that the DHC and p50 punctae visualized represented 1 DHC-EGFP dimer+1 untagged DHC dimer and 1 p50-EGFP+3 untagged p50 molecules.
Strengths:<br /> The idea and motivation behind this work are commendable.
Weaknesses:<br /> There are several major issues with the characterization of the knock-in lines generated, the choice of imaging and analysis methods, and inadequate discussion of prior findings.
The specific points are below:
1. CRISPR-edited HeLa clones:<br /> (i) The authors indicate that both the DHC-EGFP and p50-EGFP lines are heterozygous and that the level of DHC-EGFP was not measured due to technical difficulties. However, quantification of the relative amounts of untagged and tagged DHC needs to be performed - either using Western blot, immunofluorescence or qPCR comparing the parent cell line and the cell lines used in this work.<br /> (ii) The localization of DHC predominantly at the plus tips (Fig. 1A) is at odds with other work where endogenous or close-to-endogenous levels of DHC were visualized in HeLa cells and other non-polarized cells like HEK293, A-431 and U-251MG (e.g.: OpenCell (https://opencell.czbiohub.org/target/CID001880), Human Protein Atlas (https://www.proteinatlas.org/ENSG00000197102-DYNC1H1/subcellular#human), https://www.biorxiv.org/content/10.1101/2021.04.05.438428v3). The authors should perform immunofluorescence of DHC in the parental cells and DHC-EGFP cells to confirm there are no expression artifacts in the latter. Additionally, a comparison of the colocalization of DHC with EB1 in the parental and DHC-EGFP and p50-EGFP lines would be good to confirm MT plus-tip localisation of DHC in both lines.<br /> (iii) It would also be useful to see entire fields of view of cells expressing DHC-EGFP and p50-EGFP (e.g. in Spinning Disk microscopy) to understand if there is heterogeneity in expression. Similarly, it would be useful to report the relative levels of expression of EGFP (by measuring the total intensity of EGFP fluorescence per cell) in those cells employed for the analysis in the manuscript.<br /> (iv) Given that the authors suspect there is differential gene regulation in their CRISPR-edited lines, it cannot be concluded that the DHC-EGFP and p50-EGFP punctae tracked are functional and not piggybacking on untagged proteins. The authors could use the FKBP part of the FKBP-EGFP tag to perform knock-sideways of the DHC and p50 to the plasma membrane and confirm abrogation of dynein activity by visualizing known dynein targets such as the Golgi (Golgi should disperse following recruitment of EGFP-tagged DHC-EGFP or p50-EGFP to the PM), or EGF (movement towards the cell center should cease).
2. TIFRM and analysis:<br /> (i) What was the rationale for using TIRFM given its limitation of visualization at/near the plasma membrane? Are the authors confident they are in TIRF mode and not HILO, which would fit with the representative images shown in the manuscript?<br /> (ii) At what depth are the authors imaging DHC-EGFP and p50-EGFP?<br /> (iii) The authors rely on manual inspection of tracks before analyzing them in kymographs - this is not rigorous and is prone to bias. They should instead track the molecules using single particle tracking tools (eg. TrackMate/uTrack), and use these traces to then quantify the displacement, velocity, and run-time.<br /> (iv) It is unclear how the tracks that were eventually used in the quantification were chosen. Are they representative of the kind of movements seen? Kymographs of dynein movement along an entire MT/cell needs to be shown and all punctae that appear on MTs need to be tracked, and their movement quantified.<br /> (v) What is the directionality of the moving punctae?<br /> (vi) Since all the quantification was performed on SiR tubulin-treated cells, it is unclear if the behavior of dynein observed here reflects the behavior of dynein in untreated cells. Analysis of untreated cells is required.
3. Estimation of stoichiometry of DHC and p50<br /> Given that the punctae of DHC-EGFP and p50 seemingly bleach on MT before the end of the movie, the authors should use photobleaching to estimate the number of molecules in their punctae, either by simple counting the number of bleaching steps or by measuring single-step sizes and estimating the number of molecules from the intensity of punctae in the first frame.
4. Discussion of prior literature<br /> Recent work visualizing the behavior of dyneins in HeLa cells (DOI: 10.1101/2021.04.05.438428), which shows results that do not align with observations in this manuscript, has not been discussed. These contradictory findings need to be discussed, and a more objective assessment of the literature in general needs to be undertaken.
Reviewer #2 (Public Review):
Summary:<br /> Eaton and colleagues use targeted protein degradation coupled with nascent transcription mapping to highlight a role for the integrator component INST11 in terminating antisense transcription. They find that upon inhibition of CDK9, INST11 can terminate both antisense and sense transcription - leading to a model whereby INST11 can terminate antisense transcription and the activity of CDK9 protects sense transcription from INST11-mediated termination. They further develop a new method called sPOINT which selectively amplifies nascent 5' capped RNAs and find that transcription initiation is more efficient in the sense direction than in the antisense direction. This is an excellent paper that uses elegant experimental design and innovative technologies to uncover a novel regulatory step in the control of transcriptional directionality.
Strengths:<br /> One of the major strengths of this work is that the authors endogenously tag two of their proteins of interest - RBBP6 and INST11. This tag allows them to rapidly degrade these proteins - increasing the likelihood that any effects they see are primary effects of protein depletion rather than secondary effects. Another strength of this work is that the authors immunoprecipitate RNAPII and sequence extracted full-length RNA (POINT-seq) allowing them to map nascent transcription. A technical advance from this work is the development of sPOINT which allows the selective amplification of 5' capped RNAs < 150 nucleotides, allowing the direction of transcription initiation to be resolved.
Weaknesses:<br /> While the authors provide strong evidence that INST11 and CDK9 play important roles in determining promoter directionality, their data suggests that when INST11 is degraded and CDK9 is inhibited there remains a bias in favour of sense transcription (Figures 4B and C). This suggests that there are other unknown factors that promote sense transcription over antisense transcription and future work could look to identify these.
Tag text will be in white
Indigo
Tag text will be in white
SBI Life
Text on the tag will be in white.
And You Can Join Us For The Year For Just $490
I know you're trying to steer them towards the annual membership... but could you lead with the monthly price tag here to make the mental hurdle even lower?
I'd also include that it's month-to-month without lock in contract
Die Bauernproteste haben zu Revisionen von Maßnahmen zur Dekarbonisierung (und Pestizidreduktion) in europäischen Ländern und auf EU-Ebene geführt, obwohl die Klimaziele der EU ohne eine deutliche Reduktion der Emissionen der Landwirtschaft nicht zu erreichen sind. Der Arikel der New York Times beschäftigt sich mit der besonderen Rolle der Landwirtschaft in der EU-Politik und mit der Notwendigkeit, Klimapolitik als just transition zu gestalten. https://www.nytimes.com/2024/02/06/climate/europe-farming-protests-policy.html
Mehr zu den EU-Emissionszielen für 2040: https://hypothes.is/search?q=tag%3A%22EU%20emission%20goals%202040%22
Die Europäische Kommission plant, die Treibhausgas-Emissionen bis 2040 um 90% im Vergleich zu 1990 zu reduzieren. Dabei soll CCS eine große Rolle spielen. Der Plan ist eine Empfehlung für die nächste Kommission. Aufgrund der Proteste von Landwirt:innen wurden einige der vorgesehenen Regelungen, etwa zu Methan und Lachgas, in letzter Minute zurückgenommen. https://www.theguardian.com/environment/2024/feb/06/eu-lays-out-plan-to-cut-greenhouse-emissions-by-90-by-2040
Mehr dazu: https://hypothes.is/search?q=tag%3A%22EU%20emission%20goals%202040%22
Die Hitzeperioden des vergangenen Sommers haben in Frankreich 5167 Todesopfer gefordert. Das ist eine der höchsten Zahlen der vergangenen Jahre. https://www.liberation.fr/environnement/climat/plus-de-5-000-morts-lies-a-la-chaleur-lors-de-lete-2023-20240208_UFBI7UKESFFJ7MHLRKWS4DPUM4/
Mehr zu Hitzetoten: https://hypothes.is/search?q=tag%3A%22heat%20deaths%22
Kurzer Überblicksartikel der taz zur Krise in der Windindustriebranche. Sie hängt unter anderem mit Lieferkettenproblemen, Preissteigerungen und Genehmigungsverfahren zusammen, aber auch mit eigenen Fehlern der drei westlichen Marktführer #Siemens Energy, #Ørsted und #Vestas. https://taz.de/Windenergiekonzerne-in-der-Bredouille/!5987469/
Mehr zu Ørsted: https://hypothes.is/search?q=tag%3A%22Orsted%22
Mehr zu Siemens Energy: https://hypothes.is/search?q=tag%3A%22Siemens%20Energy%22
Mehr zu Vestas: https://hypothes.is/search?q=tag%3A%22Vestas%22
Protean e
Flag‐tag CDK9
DOI: 10.1002/ctm2.1531
Resource: Addgene (RRID:SCR_002037)
Curator: @olekpark
SciCrunch record: RRID:SCR_002037
The vernacular tag, at least in Berlin, of Reichskristallnacht, or“night of shattered glass,” to designate what should be considered anationwide pogrom against more than 300,000 Germans Jews inNovember 1938, was inflected with sardonic humor, which mockedthe pretentiousness of Nazi vocabulary in which Reich-this andReich-that puffed up the historical moment of the regime
Consolidated peer review report (22 January 2024)
GENERAL ASSESSMENT
Nanobodies (Nbs) are small antibody fragments that function similarly to antibodies. The smaller size of nanobodies makes them useful tools for studying biology and potentially useful as therapeutics. Nanobodies have had a significant impact on research related to the structure and function of G protein-coupled receptors (GPCRs), a family of proteins that are the target of approximately 30% of approved drugs. Nanobodies fused to peptide agonists can potentially increase the potency and selectivity of ligands.
The manuscript by Nayara Braga Emidio and Ross Cheloha describes the fusion of peptide agonists to Nbs to create chimeric ligands that differentially modulate the molecular pharmacology of the Neurokinin 1 receptor (NK1R), a potential therapeutic target for the treatment of pain. The authors observe that Nb-peptide fusions display divergent pharmacology to that of the unfused peptides via extensive characterisation at multiple signalling pathways (cAMP, Ca2+ mobilization, direct Gq TruPATH measurements, and β-arrestin recruitment), receptor binding assays, and measures of downstream transcriptional activation. The pharmacology results show that these conjugates exhibit diverse and unexpected signalling properties, including enhanced receptor binding, high potency partial agonism, prolonged cAMP production, and altered transcriptional outputs.
However, the degree to which signalling was altered was highly dependent on the location of the epitope tag and the utilized Nbs with small alterations in the relative distance and orientation between the Nb epitopes and peptide binding sites, causing significantly different outcomes. These findings highlight the potential of nanobody conjugation for creating compounds with biased agonism, extended duration of action, and improved transcriptional responses, suggesting their promise for research on GPCR signal transduction mechanisms. This study also lays the groundwork for important considerations regarding optimising nanobody-peptide fusions. Importantly, for peptide discovery, these approaches may afford improved properties regarding selectivity and duration of action. Overall, this work suggests an opportunity to create long-acting agonists with enhanced signalling properties using nanobody-peptide conjugates. However, this would require further experiments to validate the mechanism of the altered pharmacology responses of the Nb conjugates.
RECOMMENDATIONS
Essential revisions:
Because no known Nbs bind to WT NK1R, the authors have fused the epitopes of three different Nbs (6e, alpha, BC2) to the N-terminus of NK1R. These epitope tags could alter the pharmacology of the endogenous ligand Substance P (SP) with respect to the WT receptor. A comparison of signalling for WT versus the epitope-tagged NK1R in Figure 1C would alleviate these concerns.
The expected masses of the Nb conjugates after sortagging are sometimes over or under the expected masses (Table S2). Could the authors clarify the reason for these differences in the text.
Based on the results of Figure 2, all Nb conjugates, including the negative control NbGFP-peptide, negatively impact signaling by reducing efficacy in cAMP production (which is completely abolished for Nb-SP6-11) and reducing potency in β-arrestin and Gq activation. This could be due to differences in the binding of conjugated NKA compared to non-conjugated NKA due to conformational constraints or by hindering access of the peptides to the orthosteric pocket. In addition, it is unclear if the Nbs alter NK1R signaling on their own or if they act as allosteric modulators. These concerns could be experimentally addressed in both functional and binding experiments using 1) unconjugated Nbs and 2) unconjugated Nbs and peptides.
Compared to the Nbalpha-NKA and Nb6e-NKA conjugates, NbBC2-NKA has no effect in the cAMP assays (no increase in potency or effect in washout experiments). This is despite NbBC2-NKA having the greatest effect in binding experiments (Figure 3A,B). Can the authors discuss these differences, particularly with respect to the conclusion that a bitopic binding mechanism may contribute to prolonged signalling.
Regarding biased agonism as a potential advantage of Nb-peptide conjugates, the kinetics of β-arrestin recruitment or activation should also be measured (Figure 3) to determine if there is prolonged arrestin activation or receptor internalization.
The impact of Nb conjugation on ligand competition binding assays was assessed in Figures 3A and 3B. However, it would be useful to include the unconjugated Nbs as a control to determine if the enhanced inhibition is due to increased hindrance to the orthosteric pocket (see comment #3) or due to increased binding of the Nb-peptide conjugates as suggested. Similarly, in Fig S10, the lack of inhibition by spantide with the Nb6e-NKA could be due to reduced access of spantide to the orthosteric pocket in the presence of the Nb conjugate due to steric hindrance and testing with unconjugated Nb6e would strengthen the results.
The kinetics of cAMP signaling are assessed in Figures 3C and 3D. An EC10 concentration of G3-NKA was compared to an EC100 concentration of the other ligands, which may not be appropriate for comparisons of kinetics in this washout experiment. Do the authors have an explanation for comparing different concentrations?
In the cAMP washout experiments, cAMP production was still increased after washout (Figure 3C). Can the authors discuss why this was observed (Figure 3C).
In the transcriptional reporter assays (Figure 4), can the authors clarify why ~35nM was chosen as the concentration of peptides?
There are significant caveats with the Figure 5A model provided that the authors do not mention or address. Importantly, whilst AlphaFold 2 is useful for predicting the structure of well-ordered proteins, the relative location & orientation of these domains is unreliable when there are large flexible linkers between them; as is the case with the NK1R N-terminus. It would be at least worth mentioning this, and at best doing additional MD simulations to show the relative orientation of these two "domains". In addition, the authors should discuss the effects various linker lengths between the Nbs and peptide would have.
If the author's suggestions are true about the relative position of the epitope tag in relation to the peptide binding pocket, this could be demonstrated by making a construct where the epitope positions are swapped. Alternatively, instead of using 3x epitope-tagged constructs, single-tagged epitope NK1R constructs should demonstrate this.
Optional suggestions:
In several instances, the authors have chosen to show a representative concentration-response curve rather than showing data that are grouped across multiple replicates (e.g. Fig S4). Consider grouping data, as is often done in the field.
In Figure S2, it is unclear which receptor construct was used in these experiments to validate the signalling of the G3-peptides. In some of the concentration-response curves (Gs Glo, Gq TRUPATH), the maximal response is not reached, and this could affect the estimation of EC50 values reported. In any case, the authors report that G3-SP6-11 has a 100-fold increased potency, indicating that the truncation of SP might already be unfavourable for signalling. Untruncated SP could be added for comparison and may have been a better choice of ligand to see whether Nb conjugation can, in fact, improve the natural NK1R agonist.
In Figure 1D, the binding of the unconjugated Nbs to the tagged receptor was tested. It would be useful to compare the binding of unconjugated Nbs with peptide-conjugated Nbs to see whether a second binding point by the peptide increases total binding.
Fig S5 shows a lot of variability between replicates in the association of the unconjugated Nbs, is this of concern?
The opposite effect of nanobody-fusion with SP6-11 in regard to the washout experiments compared to NKA are striking but somewhat confusing. Ideally, a longer linker between the fusion should be used to show that this is indeed due to steric restraint altering the peptide binding pose.
Despite the quicker washout of the Nb-fused SP6-11 peptide, there was no significant decrease in Gq-mediated transcriptional response (Fig S11). This is difficult to reconcile, given the conclusions drawn for Nb-fused NKA (opposing effects). Is this a dose issue? The authors should explain this further.
At the end of the article, there is an emphasis on the potential usefulness of translational therapeutics. It would be ideal if the authors could further expand upon the likelihood and criteria for a nanobody that would recognize the WT NK1R and could act as a peptide fusion tool i.e. how much solvent accessible surface away from the peptide binding pocket is available for NK1R, and how likely it would provide the relative distance given the findings of this work.
REVIEWING TEAM
Reviewed by:
Reviewer #1: molecular pharmacology of pain-related GPCRs
Reviewer #2: structure and pharmacology of GPCRs
Reviewer #3: structure and pharmacology of pain-related GPCRs
Curated by:
David Thal, Senior Research Fellow, Monash University, Australia
(This consolidated report is a result of peer review conducted by Biophysics Colab on version 1 of this preprint. Comments concerning minor and presentational issues have been omitted for brevity.)
Your zettelkasten, having a perfect memory of your "past self" acts as a ratchet so that when you have a new conversation on a particular topic, your "present self" can quickly remember where you left off and not only advance the arguments but leave an associative trail for your "future self" to continue on again later.
Many thoughts and associations occur when you're having conversations with any text, whether it's with something you're reading by another author or your own notes in your zettelkasten or commonplace book. For more conversations on this topic, perhaps thumb through: https://hypothes.is/users/chrisaldrich?q=tag%3A%27conversations+with+the+text%27
If you view conversations broadly as means of finding and collecting information from external sources and naturally associating them together, perhaps you'll appreciate this quote:
No piece of information is superior to any other. Power lies in having them all on file and then finding the connections. There are always connections; you have only to want to find them.—Umberto Eco in Foucault's Pendulum (Secker & Warburg)
(Reply to u/u/Plastic-Lettuce-7150 at https://www.reddit.com/r/Zettelkasten/comments/1ae2qf4/communicating_with_a_zettelkasten/)
Should I post this photo? Are you sure? I feel like my stomach looks too big. Can I post this selfie, or are those no longer in? What is your username, I will tag you. Would it be strange to post a video on my main account? These are just a few of the hectic questions that pop into the heads of teenagers on a daily basis in regards to social media.
When looking at these questions I would say that for certain people these are questions that would plague someone's mind. It most certainly depend on the person though. For those who's entire life revolves around social media these are critical questions and I can definitely see when looking at the different areas of social media and how people would respond to the same picture differently across different social media
Reviewer #3 (Public Review):
Summary:<br /> It has been proposed in the literature, that the ATP release channel Panx1 can be activated in various ways, including by tyrosine phosphorylation of the Panx1 protein. The present study reexamines the commercial antibodies used previously in support of the phosphorylation hypothesis and the presented data indicate that the antibodies may recognize proteins unrelated to Panx1. Consequently, the authors caution about the use and interpretation of results obtained with these antibodies.
Strengths:<br /> The manuscript by Ruan et al. addresses an important issue in Panx1 research, i.e. the activation of the channel formed by Panx1 via protein phosphorylation. If the authors' conclusions are correct, the previous claims for Panx1 phosphorylation on the basis of the commercial anti-phospho-Panx1 antibodies would be in question.
This is a very detailed and comprehensive analysis making use of state-of-the-art techniques, including mass spectrometry and phos-tag gel electrophoresis.
In general, the study is well-controlled as relating to negative controls.
The value of this manuscript is, that it could spawn new, more function-oriented studies on the activation of Panx1 channels.
Weaknesses:<br /> Although the manuscript addresses an important issue, the activation of the ATP-release channel Panx1 by protein phosphorylation, the data provided do not support the firm conclusion that such activation does not exist. The failure to reproduce published data obtained with commercial anti-phospho Panx1 antibodies can only be of limited interest for a subfield.
1. The title claiming that "Panx1 is NOT phosphorylated..." is not justified by the failure to reproduce previously published data obtained with these antibodies. If, as claimed, the antibodies do not recognize Panx1, their failure cannot be used to exclude tyrosine phosphorylation of the Panx1 protein. There is no positive control for the antibodies.
2. The authors claim that exogenous SRC expression does not phosphorylate Y198. DeLalio et al. 2019 show that Panx1 is constitutively phosphorylated at Y198, so an effect of exogenous SRC expression is not necessarily expected.
3. The authors argue that the GFP tag of Panx1at the COOH terminus does not interfere with folding since the COOH modified (thrombin cleavage site) Panx1 folds properly, forming an amorphous glob in the cryo-EM structure. However, they do not show that the COOH-modified Panx1 folds properly. It may not, because functional data strongly suggest that the terminal cysteine dives deep into the pore. For example, the terminal cysteine, C426, can form a disulfide bond with an engineered cysteine at position F54 (Sandilos et al. 2012).
4. The authors dismiss the additional arguments for tyrosine phosphorylation of Panx1 given by the various previous studies on Panx1 phosphorylation. These studies did not, as implied, solely rely on the commercial anti-phospho-Panx1 antibodies, but also presented a wealth of independent supporting data. Contrary to the authors' assertion, in the previous papers the pY198 and pY308 antibodies recognized two protein bands in the size range of glycosylated and partial glycosylated Panx1.
5. A phosphorylation step triggering channel activity of Panx1 would be expected to occur exclusively on proteins embedded in the plasma membrane. The membrane-bound fraction is small in relation to the total protein, which is particularly true for exogenously expressed proteins. Thus, any phosphorylated protein may escape detection when total protein is analyzed. Furthermore, to be of functional consequence, only a small fraction of the channels present in the plasma membrane need to be in the open state. Consequently, only a fraction of the Panx1 protein in the plasma membrane may need to be phosphorylated. Even the high resolution of mass spectroscopy may not be sufficient to detect phosphorylated Panx1 in the absence of enrichment processes.
6. In the electrophysiology experiments described in Figure 7, there is no evidence that the GFP-tagged Panx1 is in the plasma membrane. Instead, the image in Figure 7a shows prominent fluorescence in the cytoplasm. In addition, there is no evidence that the CBX-sensitive currents in 7b are mediated by Panx1-GFP and are not endogenous Panx1. Previous literature suggests that the hPanx1 protein needs to be cleaved (Chiu et al. 2014) or mutated at the amino terminus (Michalski et al 2018) to see voltage-activated currents, so it is not clear that the currents represent hPANX1 voltage-activated currents.
Personal Names
This example has every tag category represented, as well as a healthy list of personal names.
See:mongodbPulls items from the array atomically. Equality is determined by casting the provided value to an embedded document and comparing using the Document.equals() function. Example: doc.array.pull(ObjectId) doc.array.pull({ _id: 'someId' }) doc.array.pull(36) doc.array.pull('tag 1', 'tag 2') To remove a document from a subdocument array we may pass an object with a matching _id. doc.subdocs.push({ _id: 4815162342 }) doc.subdocs.pull({ _id: 4815162342 }) // removed Or we may passing the _id directly and let mongoose take care of it. doc.subdocs.push({ _id: 4815162342 }) doc.subdocs.pull(4815162342); // works The first pull call will result in a atomic operation on the database, if pull is called repeatedly without saving the document, a $set operation is used on the complete array instead, overwriting possible changes that happened on the database in the meantime.
Certainly! Let's break down the explanation step by step.
pull in Mongoose:In Mongoose, the pull method is used to remove items from an array field in a document. It is designed to work atomically, meaning it ensures consistency even if multiple operations are being performed simultaneously.
javascript
doc.array.pull(...args);
Equality Check: The method uses an equality check by casting the provided value to an embedded document and comparing using the Document.equals() function.
Atomic Operation: When you call pull, it performs an atomic operation on the database, ensuring that the removal is done in a single step.
Example:
javascript
doc.array.pull(ObjectId); // Removes an item by matching ObjectId
doc.array.pull({ _id: 'someId' }); // Removes an item by matching the _id field
doc.array.pull(36); // Removes an item by matching the value 36
doc.array.pull('tag 1', 'tag 2'); // Removes items with values 'tag 1' and 'tag 2'
You can use pull to remove items from a subdocument array as well.
// Removing by passing the _id directly doc.subdocs.push({ _id: 4815162342 }); doc.subdocs.pull(4815162342); // works in the same way ```
pull call results in an atomic operation on the database.pull is called repeatedly without saving the document, a $set operation is used on the complete array instead. This means it overwrites any possible changes that happened on the database in the meantime.In Mongoose, you can use pull on an array field of a document. Here's a simple example:
```javascript const mongoose = require('mongoose');
const schema = new mongoose.Schema({ items: [{ type: String }] });
const Model = mongoose.model('Example', schema);
// Example usage Model.findOne({ _id: 'someId' }, (err, doc) => { if (err) throw err;
// Removing 'unwantedItem' from the 'items' array doc.items.pull('unwantedItem');
// Save the document to persist the changes to the database doc.save((saveErr) => { if (saveErr) throw saveErr; console.log('Item removed successfully.'); }); }); ```
In this example, pull is used to remove an item from the 'items' array of a document, and then the changes are saved to the database.
33:50 basisdemokratie, feedback, fragen<br /> warum ist die verfassung unfehlbar?<br /> wenn migranten-invasion und links-extremismus und gift-impfungen und pestizide und schulmedizin und cannabis-verbot und kriege und und und so "falsch" sind, wie sollen wir sonst die globale bevölkerung reduzieren? wie sollen wir sonst die todesrate steigern?<br /> oder, wenn es die übervölkerung gar nicht gibt, was ist dann die "carrying capacity" von diesem planet? aktuell haben wir circa 1E10 menschen global, also circa 5000 m² fläche für landwirtschaft pro person = 1000 m² für pflanzen und 4000 m² für tiere. wären 1000 mal mehr menschen "zu viel"? was dann? warum ist das "recht auf leben" unfehlbar? warum haben tiere und pflanzen kein "recht auf leben"? was ist so toll am pazifismus, der langfristig nur übervölkerung und degeneration bringt? spätestens wenn das erdöl "endlich" aus ist, wird klar, wir haben zu viele menschen.
spoiler: ich sehe mich selber nicht "links" und nicht "rechts", sondern "oben". ich beobachte dieses spiel von "oben" und denke mir: die sind alle irgendwie dumm. einerseits dieses "linke" regime, das den ganzen tag nur lügen verbreitet, und heimlich die bevölkerung austauscht, und einen bürgerkrieg vorbereitet, damit die bevölkerung reduziert wird. andererseits eine "rechte" opposition, die wahrheit und ehrlichkeit fordert, aber die selber keine bessere lösung hat zur depopulation. ich selber habe das gleiche ziel wie die "linken" (also bürgerkrieg, depopulation, deindustrialisierung, zurück zu tribalismus), und ich habe den gleichen weg wie die "rechten" (also ehrlichkeit, transparenz, wahrheit, ...)
zurück zu tribalismus
das ist auch ein thema in meinem buch:<br /> pallas. wer sind meine freunde. gruppenaufbau nach persönlichkeitstyp.<br /> ich suche eine "weltformel" für die frage: wie müssen wir verschiedene persönlichkeitstypen verbinden, damit stabile gruppen entstehen?<br /> spoiler: die konsequenz ist sicher kein pazifismus, sondern tribalismus = kleinstaaten je 150 menschen, und dazu gehören auch regelmäßige stammkriege, für natürliche selektion.<br /> auch diese "unschöne" konsequenz ist ein grund, warum meine hypothese bisher ignoriert wird... die leute wollen es schön haben, aber nichts dafür opfern, und nicht dafür kämpfen, sondern wollen alles geschenkt kriegen. ist wohl so ein merkmal der "boomer" generation, die ihr ganzes leben lang diesen "billige energie" rausch erlebt haben, der "einfach weitergehen" soll
The {@const ...} tag defines a local constant.
Author Response
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public Review):
Nitrogen metabolism is of fundamental importance to biology. However, the metabolism and biochemistry of guanidine and guanidine containing compounds, including arginine and homoarginine, have been understudied over the last few decades. Very few guanidine forming enzymes have been identified. Funck et al define a new type of guanidine forming enzyme. It was previously known that 2-oxogluturate oxygenase catalysis in bacteria can produce guanidine via oxidation of arginine. Interestingly, the same enzyme that produces guanidine from arginine also oxidises 2-oxogluturate to give the plant signalling molecule ethylene. Funck et al show that a mechanistically related oxygenase enzyme from plants can also produce guanidine, but instead of using arginine as a substrate, it uses homoarginine. The work will stimulate interest in the cellular roles of homoarginine, a metabolite present in plants and other organisms including humans and, more generally, in the biochemistry and metabolism of guanidines.
1) Significance
Studies on the metabolism and biochemistry of the small nitrogen rich molecule guanidine and related compounds including arginine have been largely ignored over the last few decades. Very few guanidine forming enzymes have been identified. Funck et al define a new guanidine forming enzyme that works by oxidation of homoarginine, a metabolite present in organisms ranging from plants to humans. The new enzyme requires oxygen and 2oxogluturate as cosubstrates and is related, but distinct from a known enzyme that oxidises arginine to produce guanidine, but which can also oxidise 2-oxogluturate to produce the plant signalling molecule ethylene.
Overall, I thought this was an exceptionally well written and interesting manuscript. Although a 2-oxogluturate dependent guanidine forming enzyme is known (EFE), the discovery that a related enzyme oxidises homoarginine is really interesting, especially given the presence of homoarginine in plant seeds. There is more work to be done in terms of functional assignment, but this can be the subject of future studies. I also fully endorse the authors' view that guanidine and related compounds have been massively understudied in recent times. I would like to see the possibility that the new enzyme makes ethylene explored. Congratulations to the authors on a very nice study.
Response: We thank the reviewer for the positive evaluation of our manuscript. In the revised version, we have emphasized more clearly that we found no evidence for ethylene production by the recombinant enzymes. The other suggestions of the reviewer are also considered in the revised version as detailed below.
Reviewer #2 (Public Review):
In this study, Dietmar Funck and colleagues have made a significant breakthrough by identifying three isoforms of plant 2-oxoglutarate-dependent dioxygenases (2-ODD-C23) as homo/arginine-6-hydroxylases, catalyzing the degradation of 6-hydroxyhomoarginine into 2aminoadipate-6-semialdehyde (AASA) and guanidine. This discovery marks the very first confirmation of plant or eukaryotic enzymes capable of guanidine production.
The authors selected three plant 2-ODD-C23 enzymes with the highest sequence similarity to bacterial guanidine-producing (EFE) enzymes. They proceeded to clone and express the recombinant enzymes in E coli, demonstrating capacity of all three Arabidopsis isoforms to produce guanidine. Additionally, by precise biochemical experiments, the authors established these three 2-ODD-C23 enzymes as homoarginine-6-hydroxylases (and arginine-hydroxylase for one of them). Furthermore, the authors utilized transgenic plants expressing GFP fusion proteins to show the cytoplasmic localization of all three 2-ODD-C23 enzymes. Most notably, using T-DNA mutant lines and CRISPR/Cas9-generated lines, along with combinations of them, they demonstrate the guanidine-producing capacity of each enzyme isoform in planta. These results provide robust evidence that these three 2-ODD-C23 Arabidopsis isoforms are indeed homoarginine-6-hydroxylases responsible for guanidine generation.
The findings presented in this manuscript are a significant contribution for our understanding of plant biology, particularly given that this work is the first demonstration of enzymatic guanidine production in eukaryotic cells. However, there are a couple of concerns and potential ways for further investigation that the authors should (consider) incorporate.
Firstly, the observation of cytoplasmic and nuclear GFP signals in the transgenic plants may also indicate cleaved GFP from the fusion proteins. Thus, the authors should perform Western blot analysis to confirm the correct size of the 2-ODD-C23 fusion proteins in the transgenic protoplasts.
Secondly, it may be worth measuring pipecolate (and proline?) levels under biotic stress conditions (particularly those that induce transcript changes of these enzymes, Fig S8). Given the results suggesting a potential regulation of the pathway by biotic stress conditions (eg. meJA), these experiments could provide valuable insights into the physiological role of guanidine-producing enzymes in plants. This additional analysis may give a significance of these enzymes in plant defense mechanisms.
Response: We thank also reviewer 2 for the positive evaluation and useful suggestions. We performed the proposed GFP Western blot, which indeed indicated the presences of both, fulllength fusion proteins and free GFP, which can explain the partial nuclear localization. We fully agree that further experiments with biotic and abiotic stress will be required to determine the physiological function of the 2-ODD-C23 enzymes. However, the list of potential experiments is long and they are beyond the scope of the present manuscript.
Reviewer #1 (Recommendations For The Authors):
Specific points
Overall, I thought this was a very interesting study, comprising biochemical, cellular, and in vivo studies. Of course more could be done on each of these, and likely will be, but I think the assignment of biochemical function is very strong, across all three approaches. The one new experiment I would like to see is a clear demonstration of whether ethylene is produced - unlikely but should be tested.
We had mentioned our failure to detect ethylene production by the plant enzymes in the previous version and have made it more prominent and reliable by including ethylene production as positive control in the new supplementary figure S5.
Abstract
Delete 'hitherto overlooked' - this is implicit 'but is more likely' to 'is likely'?
Agreed and modified
Introduction
Second sentence - what about relevant small molecule primary metabolites including precursors of proteins/nucleic acids.
We modified the sentence accordingly.
Paragraph 2 - maybe also note EFE produces glutamate semi aldehyde, via arginine C-5 oxidation.
Paragraph 2 has been re-phrased according to your suggestion.
Overall, I thought the introduction was exceptionally well written.
Perhaps either in the introduction, or later, note there are other 2OG oxygenases that oxidise arginine/arginine derivatives in various ways, e.g. clavaminate synthase/arginine hydroxylases/desaturases.
We added a sentence mentioning the arginine hydroxylases VioC and OrfP to the introduction and included VioC into the sequence comparison in supplementary figure 2 to show that these enzymes, as well as NapI, are very different from EFE and the plant hydroxylases.
Results
Paragraph 1 - qualify similarity and refer to/give a structurally informed sequence alignment, including EFE
A new supplemental figure S2 was added with sequence identity values and a structurally informed alignment. The text has been modified accordingly.
Paragraph 2 - briefly state method of guanidine analysis
We included a reference to the M&M section and mentioned LC-MS in paragraph 2.
Figure 1 - trivial point - proteins are not expressed/genes are
We have modified the legend to figure 1. However, we would like to point out that terms like “recombinant protein expression” are widely used in the field. A quick search with google Ngram viewer shows that “protein expression” started to appear in the mid-80ies and its use stayed constantly at 1/8th of “gene expression”.
Define errors clearly in all figure legends, clearly defining biological/technical repeats<br /> Page 6 - was the His-tag cleared to ensure no issues with Ni contamination?
We treat individual plants or independent bacterial cultures as biological replicates. Only in the case of enzyme activity assays with NAD(P)H, technical replicates were used and this has been indicated in the legend of figure 6.
Lower case 'p' in pentafluorobenzyl corrected
In Figure 2 make clear the hydroxylated intermediates are not observed
We now use grey color for the intermediates and have put them in brackets. Additionally we state in the figure legend that these intermediates were not detected.
Pages 6-7 - I may have missed this but it's important to investigate what happens to the 2OG. Is succinate the only product or is ethylene also produced? This possibility should also be considered in the plant studies, i.e. is there any evidence for responses related to perturbed ethylene metabolism. The authors consider a signalling role relating to AASA/P6C, but seem to ignore a potential ethylene connection.
As stated above, we checked for ethylene production with negative result. EFE produced 6 times more guanidine than the plant enzymes under the same condition, but even 100-fold lower ethylene production would have been clearly detected.
Page 12 - 'plants have been shown to....' Perhaps note how hydroxy guanidine is made?
We now mention the canavanine-γ-lyase that cleaves canavanine into hydroxyguanidine and homoserine.
Overall, I thought the discussion was good, but perhaps a bit long/too speculative on pages 12/13 and this detracted from the biochemical assignment of the enzyme. I'd suggest shortening the discussion somewhat - the precise roles of the enzyme can be the subject of future work. As indicated above, some discussion on potential links to ethylene would be appreciated.
Since reviewer 2 wanted more (speculative) discussion on the role of the 2-ODD-C23 enzymes and there was no detectable ethylene production, we took the liberty to leave the discussion largely unaltered.
I'd also like to see some more consideration/metabolic analyses of guanidine related metabolism in the genetically modified plants.
Such analyses will certainly be included in future experiments once we get an idea about the physiological role of the 2-ODD-C23 enzymes.
Page 16 - mass spectrometry
Corrected.
Please add a structurally informed sequence alignment with EFE and other 2OG oxygenases acting on arginine/derivatives.
An excerpt of the alignment is now presented in supplementary figure S2.
Reviewer #2 (Recommendations For The Authors):
I would like to see more discussion in the manuscript about the possible interconnection/roles between 2-ODD-C23 guanidine-producing, lysine- ALD1-Pipecolate producing, and proline metabolism pathways during both biotic and abiotic stresses.
Since we were unable to detect pipecolate in any of our plant samples and also our preliminary results with biotic stress did not produce any evidence for a function of the 2ODD-C23 enzymes in the tested defense responses, we would like to postpone such extended discussion until we find a condition where the physiological function of these enzymes is evident.
Fig. 4: Authors should change colors for Col-0, 0.2 HoArg and ctrl? They look too similar in my pdf file.
We changed the colors in figure 4 and hope that the enhanced contrast is maintained during the production of the final version of our article.
Reviewer #1 (Public Review):
Summary:
Bartolome et al. report adaptation of proximity labeling using BirA and TurboID fusions to proteasome subunits to identify the proteasome-proximal proteome both in cultured cells and also in a newly developed mouse model. Using this approach, the authors demonstrate identification of many known proteasome-interacting proteins, as well as several new proteins, some of which are validated directly. The authors further evaluate the proteasome-proximal proteome in most mouse organs, and find substantial agreement with the proteome identified from cultured cells, as well as between tissues. This represents one of the first studies of the "proteasome-ome" in vivo, and sets the stage for addressing numerous important future questions regarding how the proteasome's environment changes over time, in response to different stimuli, and in distinct disease conditions.
Strengths:
Generally speaking, the approach provided is rigorous and supported by several complementary lines of evidence, such as demonstration that the interactome is enriched for known proteasome-binding proteins and co-purification or co-elution experiments. Similarly, the high agreement between the outcomes in cultured cells and in the mouse model developed by the authors provides further confidence in the results.
Weaknesses:
The major weakness of the work is arguably the choice of proteasome subunits for tagging with biotinylating enzymes. In most cases, the subunits and termini chosen for tagging are known to either protrude toward functionally important regions (such as the substrate-processing pore of the ATPase component), to have important functional roles likely to be disrupted via tagging, or are subunits known to be substituted by others in some conditions. Thus, the interactome reported may conflate those of normal proteasomes with those harboring tag-induced functional or structural defects. Although the authors made a commendable attempt to demonstrate minimal impacts of tagging, the conclusions would be greatly further strengthened by contrasting the impacts of tagging subunits less likely to cause perturbations and by more rigorously demonstrating normal proteolysis of a broader array of known proteasome substrates.
Reviewer #3 (Public Review):
Summary:
Bartolome et al. present ProteasomeID, a novel method to identify components, interactors, and (potentially) substrates of the proteasome in cell lines and mouse models. As a major protein degradation machine that is highly conserved across eukaryotes, the proteasome has historically been assumed to be relatively homogeneous across biological scales (with few notable exceptions, e.g., immunoproteasomes and thymoproteasomes). However, a growing body of evidence suggests that there is some degree of heterogeneity in the composition of proteasomes across cell tissues, and can be highly dynamic in response to physiologic and pathologic stimuli. This work provides a methodological framework for investigating such sources of variation. The authors start by adapting the increasingly popular biotin ligation strategy for labelling proteins coming into close proximity with one of three different subunits of the proteasome, before proceeding with PSMA4 for further development and analysis based on their preliminary labelling data. In a series of well-constructed and convincing validation experiments, the authors go on to show that the tagged PSMA4 construct can be incorporated into functional proteasomes, and is able to label a broad set of known proteasome components and interacting proteins in HEK293T cells. They also attempt to identify novel proteasomal degradation substrates with ProteasomeID; while this was convincing for known substrates with particularly short half-lives (exemplified by the transcription factor c-myc), follow-up validation experiments with other substrates were less clear. One of the most compelling results was from a similar experiment to confirm proteasomal degradation induced by a BRD-targeting PROTAC, which I think is likely to be of keen interest to the targeted degradation community. Finally, the authors establish a ProteasomeID mouse model, and demonstrate its utility across several tissues.
Strengths:
1) ProteasomeID itself is an important step forward for researchers with an interest in protein turnover across biological scales (e.g., in sub-cellular compartments, in cells, in tissues, and whole organisms). I especially see interest from two communities: those studying fundamental proteostasis in physiological and pathologic processes (e.g., ageing; tissue-specific protein aggregation diseases), and those developing targeted protein degradation modalities (e.g., PROTACs; molecular glues). All the datasets generated and deposited here are likely to provide a rich resource to both. The HEK293T cell line data are a valuable proof-of-concept to allow expansion into more biologically-relevant cell culture settings; however, I envision the greatest innovation here to be the mouse model. For example, in the targeted protein degradation space, two major hurdles in early-stage pre-clinical development are (i) evaluation of degradation efficacy across disease-relevant tissues, and (ii) toxicity and safety implications caused by off-target degradation, e.g., of newly-identified molecular glues and/or in particularly-sensitive tissues. The ProteasomeID mouse allows early in vivo assessment of both these questions. The results of the BRD PROTAC experiment in 293T cells provides an excellent in vitro proof-of-concept for this approach.
2) The mass spectrometry-based proteomics workflows used and presented throughout the manuscript are robust, rigorous, and convincing. For example, the algorithm the authors use for defining enrichment score cut-offs are logical and based on rational models, rather than on arbitrary cut-offs that are common for similar proteomics studies. The construction (and subsequent validation) of both BirA*- and miniTurbo- tagged PSMA4 variants also increases the utility of the method, allowing researchers to choose the variant with the labelling time-scale required for their particular research question.
3) The optimised BioID and TurboID protocol the authors develop (summarised in Fig. S2A) and validate (Fig. S2B-D) is likely to be of broad interest to cell and molecular biologists beyond the protein degradation field, given that proximity labelling is a current gold-standard in global protein:protein interaction profiling.
Limitations:
I think the authors do an excellent job in highlighting the limitations of ProteasomeID throughout the Results and Discussion. I do have some specific comments that might provide additional context for the reader.
1) The authors do a good job in showing that a substantial proportion of PSMA4-BirA* is incorporated into functional proteasome particles; however, it is not immediately clear to me how much background (false-positive IDs) might be contributed by the ~40 % of PSMA4-BirA* that is not incorporated into the mature core particle (based on the BirA* SEC-MS traces in Fig. 2b and S3b, i.e., the large peak ~ fraction 20). Are there any bands lower down in the native gel shown in Fig. 2c, i.e., corresponding to lower molecular weight complexes or monomeric PSMA4-BirA*? The enrichment of proteasome assembly factors in all the ProteasomeID experiments might suggest the presence of assembly intermediates, which might themselves become substrates for proteasomal degradation (as has been shown for other incompletely-assembled protein complexes, e.g., the ribosome, TRiC/CCT).
2) Although the authors attempt to show that BirA* tagging of PSMA4 does not interfere with proteasome activity (Fig. 2e-f), I think the experimental evidence for this is incomplete. They show that the overall chymotrypsin-like activity (attributable to PSMB5) in cells expressing PSMA4-BirA* is not markedly reduced compared with control BirA*-expressing cells. However, they do not show that the activity of the specific proteasome sub-population that contains PSMA4-BirA* is unaffected (e.g., by purifying this sub-population via the Flag tag). The proteasome activity of the sub-population of wild-type proteasome complexes that do not contain the PSMA4-BirA* (~50%, based on the earlier immunoblots) could account for the entire chymotrypsin-like activity-especially in the context of HEK293T cells, where steady-state proteasome levels are unlikely to be limiting. It would also be useful to assess any changes in tryspin- and caspase- like activities, especially as tagging of PSMA4 could conceivably interfere with the activity of some PSMB subunits, but not others.
3) I was left unsure of the general utility of ProteasomeID for identifying novel proteasomal substrates in homeostatic or stressed conditions. The immunoblots for the two candidates the authors follow up in Fig. 4g was not especially clear; the reduction in the bands are modest, at best. Furthermore, classifying candidates based on enrichment following proteasome inhibition with MG-132 have the potential to lead to a high number of false positives. ProteasomeID's utility in identifying potential substrates in more targeted settings (e.g., molecular glues, off-target PROTAC substrates) is far more apparent.
Reviewer #1 (Public Review):
Summary:<br /> In their manuscript, Zhou et al. analyze the factors controlling the activation and maintenance of a sustained cell cycle block in response to persistent DNA DSBs. By conditionally depleting components of the DDC using auxin-inducible degrons, the authors verified that some DDC proteins are only required for the activation (e.g., Dun1) or the maintenance (e.g., Chk1) of the DSB-dependent cell cycle arrest, while others such as Ddc2, Rad24, Rad9 or Rad53 are required for both processes. Notably, they further demonstrate that after a prolonged arrest (>24 h) in a strain carrying two DSBs, the DDC becomes dispensable and the mitotic block is then maintained by SAC proteins such as Mad1, Mad2, or the mitotic exit network (MEN) component Bub2.
Strengths:<br /> The manuscript dissects the specific role that different components of the DDC and the SAC have during the induction of a cell cycle arrest induced by DNA damage, as well as their contribution to the short-term and long-term maintenance of a DNA DSB-induced mitotic block. Overall, the experiments are well described and properly executed, and the data in the manuscript are clearly presented. The conclusions drawn are also generally well supported by the experimental data. The observations contribute to drawing a clearer picture of the relative contribution of these factors to the maintenance of genome stability in cells exposed to permanent DNA damage.
Weaknesses:<br /> The main weakness of the study is that it is fundamentally based only on the use of the auxin-inducible degron (AID) strategy to deplete proteins. This is a widely used method that allows a very efficient depletion of proteins. However, the drawback is that a tag is added to the protein, which can affect the functionality of the targeted protein or modify its capacity to interact with others. In fact, three of the proteins that are depleted using the AID systems are shown to be clearly hypomorphic. Verification of at least some of the results using an alternative manner to eliminate the proteins would help to strengthen the conclusions of the manuscript.
[Banken-Rettung um jeden Preis]
19:39<br /> Glauben Sie nicht, dass die Notenbanken bald gegensteuern?<br /> Weil sie vielleicht sogar gezwungen werden?
19:58<br /> Danke für die Frage.<br /> Es ist nicht nur so, dass die Frage berechtigt ist,<br /> "glauben Sie nicht, Herr Krall, dass die Banken gegensteuern werden?"<br /> Ich bin sicher, dass sie gegensteuern werden!<br /> Darauf läuft die ganze Sache ja hinaus.
20:07<br /> Schon beim nächsten Mal?
Die steuern ja schon gegen.<br /> Die FED [Federal Reserve] hat ja gerade zwei Billionen [US-Dollar]<br /> Liquiditätsspritze ins Bankensystem gegeben.
20:16<br /> Ist das jetzt QE [Quantitative Easing]?
Im Grunde genommen ist das QE [Quantitative Easing].<br /> Natürlich ist das QE.<br /> Wenn ich zwei Billionen zur Verfügung stelle,<br /> dann muss sie im Zweifel ja auch<br /> tatsächlich "drucken" oder elektronisch schaffen.<br /> Das heißt<br /> die Fiat-Geldmaschine wird ja gerade schon wieder angeworfen,<br /> das ist ja das, worauf es rausläuft.
20:30<br /> Natürlich gehen die Banken nicht pleite,<br /> die können die gar nicht pleite gehen lassen.<br /> Die Bankenkrise ist da, das Eigenkapital ist weg,<br /> wahrscheinlich ist es drei Mal weg, vielleicht ist es auch fünf Mal weg.<br /> Das war auch schon weg, bevor es jetzt geknallt hat,<br /> weil die Zombies, die waren ja schon vorher da,<br /> aber die sind halt nicht in die Knie gegangen ohne Zinserhöhung.
20:49<br /> Aber ökonomisch betrachtet,<br /> wenn man eine korrekte Risikobetrachtung gemacht hätte,<br /> und einen richtigen Stresstest gemacht hätte,<br /> dann hätte man schon längst wissen können,<br /> wie schlecht es den Banken geht in Wahrheit.
21:00<br /> Die ganze Fata Morgana von dem erhöhten Eigenkapital,<br /> das ist für die Leichtgläubigen unter uns.
21:05<br /> Und jetzt ist es natürlich so,<br /> dass die die Banken nicht abstürzen lassen werden.<br /> Ihr Geld ist auf dem Sparkonto vollkommen sicher,<br /> da müssen sie sich keine Sorgen machen,<br /> die werden gerettet werden, und zwar kostet es was es wolle.
21:15<br /> Weil wenn das nicht gerettet wird...<br /> das haben die Jungs aus Lehman gelernt...<br /> also wenn man so eine Bank in der Kampfklasse<br /> einfach untergehen lässt,<br /> und die nicht rettet, dann ist Achterbahn.
Und da gilt der schöne Satz:<br /> "ich zähle jetzt bis Eins, und dann ist Achterbahn."<br /> Und also bis zum "Eins zählen" kommen sie da gar nicht.
Also werden die gerettet,<br /> und die werden dafür die Geldmenge ausdehnen müssen,<br /> das ist das worauf es rausläuft.
21:39
Aber wer bezahlt dann die Rechnung?
Die Rechnung ist doch ganz klar.<br /> Wenn ich die Geldmenge ausdehne,<br /> in Europa um vier bis sechs,<br /> oder vielleicht sogar acht Billionen,<br /> in Amerika auch...<br /> denn wir stehen ja am Anfang,<br /> und zwei Billionen haben die Amerikaner jetzt schon dahingestellt,<br /> bevor es überhaupt irgendwie richtig los geht.
00:22:00<br /> [Dann steigt Inflation auf 30%]
21:55<br /> Also wenn da 4... 6... 8 Billionen auf beiden Seiten des Atlantiks<br /> an Zentralbank-Geldmenge frisch geschaffen wird,<br /> dann wird die Inflation zum nächsten Auf-Galopp ansetzen,<br /> geht dann aufs nächste Plateau,<br /> von jetzt 15 Prozent auf dann 30 Prozent.
22:14<br /> Und dann stehen die Zentralbanken wieder wie der Ochs vorm Berg,<br /> weil die werden dann sagen: "Hmm, 30 Prozent, schlecht..."<br /> weil dann wird es nämlich da draußen ungemütlich.
22:24<br /> Wir haben jetzt schon ein Drittel aller Haushalte in Deutschland<br /> (und nicht nur in Deutschland)<br /> die mit ihrem Geld nicht mehr klarkommen.
22:29<br /> Die weichen jetzt aus,<br /> da gibts jetzt eben kein Hühnchen mehr, und kein Schweinesteak,<br /> da gibts jetzt Haferflocken.<br /> Also die stellen Ihren Warenkorb um,<br /> deswegen erzählt man ihnen, sie sollen Insekten essen,<br /> was ich ja für die blödeste Idee seit Adam und Eva halte.<br /> Das ist schon spektakulär, wenn man darüber nachdenkt.
22:47<br /> Also ein Drittel der Haushalte ist jetzt schon in Schwierigkeiten,<br /> und zwar in ernsthaften Schwierigkeiten.<br /> Wenn dann noch Anpassungen bei den Hypothekenzahlungen kommen,<br /> dann wirds auch ganz viele "Häuslebauer" erwischen,<br /> die nicht drauf eingestellt sind,<br /> dass sich ihre Zinsbelastung vervierfacht oder verfünffacht.
23:02<br /> Wenn wir dann 30 Prozent [Inflation] haben,<br /> dann werden wir sehen,<br /> dass 80 bis 90 Prozent der Haushalte Schwierigkeiten haben,<br /> am Ende des Geldes ist dann noch zu viel Monat übrig,<br /> und ich würde mal sagen, die Hälfte ernsthafte Schwierigkeiten,<br /> das heißt, da wird es zu Zahlungsunfähigkeit kommen.
23:18<br /> Und wenn sie die Hälfte der Haushalte<br /> in die Zahlungsunfähigkeit schreiben<br /> durch eine inkompetente Geldpolitik,<br /> dann wage ich die Prognose,<br /> dass sich die Leute das nicht gefallen lassen werden,<br /> und dass das zu sozialen Verwerfungen führt.
23:28<br /> Aber jetzt sagen Sie "inkompetente Geldpolitik".<br /> Was würden Sie denn jetzt machen?<br /> Also Sie würden die Zinsen nicht senken,<br /> und die Banken pleite gehen lassen?<br /> Das ist ja auch nicht besser, oder?
23:35<br /> Roland Tichy hat mich mal gefragt,<br /> was ich tun würde,<br /> wenn ich Zentralbank-Chef wäre,<br /> wenn ich EZB-Chef wäre,<br /> und meine Antwort war ganz einfach:<br /> Zum Glück bin ich das nicht.<br /> Weil die haben sich so in die Falle manövriert,<br /> dass sie da nicht mehr rauskommen.
23:51<br /> Also es gibt keinen guten Ausweg?
23:52<br /> Es gibt keine gute Lösung.<br /> Der Mittelweg, den haben sie ja jetzt versucht.<br /> Diese dreieinhalb Prozent [Zinsen]<br /> waren ja eigentlich der Versuch des Mittelwegs,<br /> nämlich zu sagen:<br /> "Wasch mich, aber mach mich nicht nass."<br /> Den kleinen Fußzeh ins Wasser gestreckt,<br /> und dann mit furchtbarem Geheule festgestellt,<br /> wie kalt das Wasser ist.
24:10<br /> Eigentlich bräuchte man ja einen viel höheren Zinssatz,<br /> um 10% Inflation einzudämmen.<br /> Man braucht positive Realzinsen, um das Ding auf Schiene zu setzen,<br /> die haben wir aber bei weitem nicht,<br /> wir haben negative Realzinsen von 7 Prozent.
24:20<br /> Das heißt also,<br /> es werden gewaltige Mengen an Vermögen verbrannt durch die Inflation,<br /> und wenn ich die [Inflation] wirklich bekämpfen würde...
24:31<br /> und natürlich auch Einkommen... das Einkommen schrumpft.<br /> Wir haben seit Beginn dieser Krise<br /> einen Einkommensrückgang in Europa um 14 Prozent.<br /> Das ist mehr als zu Beginn der großen Depression 1929.
24:40<br /> Wenn sie da 30% draus machen, oder 40% oben drauf,<br /> dann sind sie schon bei minus 30, minus 40 Prozent.<br /> Das heißt die Einkommen schrumpfen im gleichen Ausmaß wie 1929/30.
24:51<br /> Und der Fehler ist nicht darin zu sehen,<br /> was die jetzt tun können oder nicht tun können,<br /> da habe ich keinen guten Rat.<br /> "damned if you do, damned if you dont."
25:00<br /> Der Fehler liegt darin:<br /> Die Inkompetenz der Geldpolitik reicht jetzt 20 Jahre zurück.<br /> Von Tag eins an war der Euro so konstruiert,<br /> dass er die Verschwendung alimentiert,<br /> dass er die Staatsfinanzen in Südeuropa finanziert,<br /> und so war er auch gedacht.
25:13<br /> Und man hat nicht ans Ende gedacht,<br /> als man das angefangen hat.<br /> Man hat gedacht, die Deutschen zahlen die Party,<br /> bis zum Sankt Nimmerleinstag.
Aber hier gilt jetzt der Satz von Maggie Thatcher:<br /> Die EU ist am Ende, wenn ihr das Geld der Deutschen ausgeht.
Analog zu dem Satz:<br /> Der Sozialismus ist am Ende, wenn ihm das Geld andere Leute ausgeht.
Nämlich das ist das, was die EU ist, sie ist Staats-Sozialismus.<br /> Und jetzt ist der Weg zu Ende gegangen,<br /> und die Reserven sind verbraucht.
— Markus Krall bei Mario Lochner, 2023-04-05
FOR HIGHER EDUCATIONElevate your instruction with time-saving tools Save time with the AI question generator, quickly assess student learning with game reports, and inspire student-led learning with student passes. Save 20% on Kahoot!+ from $11.99/month until January 31. Buy now Learn more
The good: Every image on the website has a descriptive alt tag, limited to 125 characters or less. This requirement ensures that screen readers can effectively convey the content of images. The descriptions are concise yet detailed enough to accurately represent the images, avoiding the use of random letters and numbers that often appear in file names (e.g., n83zeo1234q.jpg). This practice aligns well with the Perceivable principle of the Web Content Accessibility Guidelines (WCAG), ensuring that information is accessible to all users, regardless of their sensory abilities.
The image in this new article has a descriptive <alt> tag of 125 characters or less that a screen reader can read. This description is also concise while still containing enough detail to describe the image. Additionally, it does not contain random letters and numbers assigned to many image files.
deine website ist ein trauriger witz : /
schonmal was gehört von "zensurresistent"?
iss nur ne frage der zeit, bis die bullen deine domain abschalten
wie bei...
torrent.to https://www.digitaltrends.com/web/germany-sentences-torrent-site-owner-to-3-years-in-jail/
spickmich.de https://de.wikipedia.org/wiki/Spickmich
lernsieg.at https://www.w24.at/News/2019/11/Umstrittene-Lernsieg-App-wieder-offline
rarbg.to https://de.wikipedia.org/wiki/RARBG
didw*.onion https://de.wikipedia.org/wiki/Deutschland_im_Deep_Web
oder... die zensurliste ist lang, und wird jeden tag länger
https://www.anonymousnews.org/international/die-bruesseler-zensur-krake/
auch telegram ist zu zentral, wir brauchen eher was wie ssb (secure scuttlebutt), gittorrent, zeronet, berty, briar, sneakernet, ... also "offline first", was auch innem ad-hoch bluetooth/wlan p2p netzwerk funktioniert... dann braucht man schon nen stromausfall (oder ne EMP) um das system zu blockieren
remember, china ist das vorbild für die NWO, also die gleiche aggressive internet-zensur wird auch hier kommen, völlig egal mit welcher "begründung", polizeigewalt braucht keine gründe, und wenn irgendwelche richter 5 jahre später sagen "das war eigentlich doch legal" dann hilft das auch nix
Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Public Review):
The apicoplast, a non-photosynthetic vestigial chloroplast, is a key metabolic organelle for the synthesis of certain lipids in apicomplexan parasites. Although it is clear metabolite exchange between the parasite cytosol and the apicoplast must occur, very few transporters associated with the apicoplast have been identified. The current study combines data from previous studies with new data from biotin proximity labeling to identify new apicoplast resident proteins including two putative monocarboxylate transporters termed MCT1 and MCT2. The authors conduct a thorough molecular phylogenetic analysis of the newly identified apicoplast proteins and they provide compelling evidence that MCT1 and MCT2 are necessary for normal growth and plaque formation in vitro along with maintenance of the apicoplast itself. They also provide indirect evidence for a possible need for these transporters in isoprenoid biosynthesis and fatty acid biosynthesis within the apicoplast. Finally, mouse infection experiments suggest that MCT1 and MCT2 are required for normal virulence, with MCT2 completely lacking at the administered dose. Overall, this study is generally of high quality, includes extensive quantitative data, and significantly advances the field by identifying several novel apicoplast proteins together with establishing a critical role for two putative transporters in the parasite. The study, however, could be further strengthened by addressing the following aspects:
Response: We thank very much the reviewer for his/her positive evaluation of our work. To address the detailed function of the transporters, in the past three months, we have re-constructed plasmids (with codon-optimized DNA sequences of the genes) for expression of the transporters in a regular expression E. coli strain (BL21DE3) and in a pyruvate import knockout E. coli strain (a gift from Prof. Kirsten Jung), to examine the transport capability in vitro. And, we have also re-constructed a new plasmid containing a new leading peptide for targeting the pyruvate sensor PyronicSF to the apicoplast in the parasite, to probe the possible substrate pyruvate. However, we did not successfully observe expression of the transporters in the above E. coli strains, and we were unable to target the sensor to the correct localization (the apicoplast) in the parasite. As a result, all efforts have led the study to the current version of manuscript on the functional identification of transporters. We will keep working on this aspect, attempting to dissect out the exact transport function of the transporters in the future. In the current manuscript, we have discussed the limitations of our study in the last part of the manuscript.
Main comments
1) The conclusion that condition depletion of AMT1 and/or AMT2 affects apicoplast synthesis of IPP is only supported by indirect measurements (effects on host GFP uptake or trafficking, possibly due to effects on IPP dependent proteins such as rabs, and mitochondrial membrane potential, possibly due to effects on IPP dependent ubiquinone). This conclusion would be more strongly supported by directly measuring levels of IPP. If there are technical limitations that prevent direct measurement of IPP then the author should note such limitations and acknowledge in the discussion that the conclusion is based on indirect evidence.
Response: We thank the reviewer very much for the suggestions. We have tried to establish the measurement of IPP using a commercial company in recent months, yet we have not been successful in making the assay work. Considering the problem of indirect evidence, we have discussed this limitation in the discussion.
2) The conclusion that condition depletion of AMT1 and/or AMT2 affects apicoplast synthesis of fatty acids is also poorly supported by the data. The authors do not distinguish between the lower fatty acid levels being due to reduced synthesis of fatty acids, reduced salvage of host fatty acids, or both. Indeed, the authors provide evidence that parasite endocytosis of GFP is dependent on AMT1 and AMT2. Host GFP likely enters the parasite within a membrane bound vesicle derived from the PVM. The PVM is known to harbor host-derived lipids. Hence, it is possible that some of the decrease in fatty acid levels could be due to reduced lipid salvage from the host. Experiments should be conducted to measure the synthesis and salvage of fatty acids (e.g., by metabolic flux analysis), or the authors should acknowledge that both could be affected.
Response: We thank the reviewer very much for comments and suggestions. We partially agree with the comments that the depletion of transporters could affect lipids scavenged from the host cells, as endocytic vesicles are indeed derived from the parasite plasma membrane at the micropore and potentially from the host cell endo-membrane system, as demonstrated with the micropore endocytosis in our previous study (pmid: 36813769). Our latest study has addressed this by showing that the endocytic trafficking of GFP vesicles is regulated by prenylation of proteins (e.g. Rab1B and YKT6.1), depletion of which resulted in diffusion of GFP vesicles, but not disappearance of GFP vesicles in the parasites (pmid: 37548452), indicating that the vesicles (containing lipids) enter the parasites. In the current manuscript, the percentage of parasites containing GFP foci was significantly reduced in AMT1/AMT2-depleted parasites, and instead, parasites containing GFP diffusion appeared and the percentage was almost equal to the reduced level of parasites with GFP foci. These results suggested that endocytic vesicles (e.g. GFP vesicles) were continuously generated by the micropore in the parasites depleted with AMT1/AMT2, and that the vesicle trafficking was regulated by proteins modified by IPP derivatives that were derived from the apicoplast. Based on these observations, we considered that lipids in endocytic vesicles should not contribute to the reduced level of fatty acids and other lipids in parasites depleted with AMT1/AMT2. We have added in a short discussion concerning the fatty acids and lipids reduced in the parasites.
Reviewer #2 (Public Review):
In this study Hui Dong et al. identified and characterized two transporters of the monocarboxylate family, which they called Apcimplexan monocarboxylate 1 and 2 (AMC1/2) that the authors suggest are involved in the trafficking of metabolites in the non-photosynthetic plastid (apicoplast) of Toxoplasma gondii (the parasitic agent of human toxoplasmosis) to maintain parasite survival. To do so they first identified novel apicoplast transporters by conducting proximity-dependent protein labeling (TurboID), using the sole known apicoplast transporter (TgAPT) as a bait. They chose two out of the three MFS transporters identified by their screen based and protein sequence similarity and confirmed apicoplast localisation. They generated inducible knock down parasite strains for both AMC1 and AMC2, and confirmed that both transporters are essential for parasite intracellular survival, replication, and for the proper activity of key apicoplast pathways requiring pyruvate as carbon sources (FASII and MEP/DOXP). Then they show that deletion of each protein induces a loss of the apicoplast, more marked for AMC2 and affects its morphology both at its four surrounding membranes level and accumulation of material in the apicoplast stroma. This study is very timely, as the apicoplast holds several important metabolic functions (FASII, IPP, LPA, Heme, Fe-S clusters...), which have been revealed and studied in depth but no further respective transporter have been identified thus far. hence, new studies that could reveal how the apicoplast can acquire and deliver all the key metabolites it deals with, will have strong impact for the parasitology community as well as for the plastid evolution communities. The current study is well initiated with appropriate approaches to identify two new putatively important apicoplast transporters, and showing how essential those are for parasite intracellular development and survival. However, in its current state, this is all the study provides at this point (i.e. essential apicoplast transporters disrupting apicoplast integrity, and indirectly its major functions, FASII and IPP, as any essential apicoplast protein disruption does). The study fails to deliver further message or function regarding AMC1 and 2, and thus validate their study. Currently, the manuscript just describes how AMC1/2 deletion impacts parasite survival without answering the key question about them: what do they transport? The authors yet have to perform key experiments that would reveal their metabolic function. I would thus recommend the authors work further and determine the function of AMC1 and 2.
Response: We thank very much the reviewer for his/her positive evaluation of our work. To address the detailed function of the transporters, in the past three months, we have re-constructed plasmids (with codon-optimized DNA sequences of the genes) for expression of the transporters in a regular expression E. coli strain (BL21DE3) and in a pyruvate import knockout E. coli strain (a gift from Prof. Kirsten Jung), to examine the transport capability in vitro. And, we have re-constructed a new plasmid containing a new leading peptide for targeting the pyruvate sensor PyronicSF to the apicoplast in the parasite, to probe the possible substrate pyruvate. However, we were unable to successfully observe expression of the transporters in the above E. coli strains, and we were unable to target the sensor to the correct localization (the apicoplast) in the parasite. As a result, all these efforts have led the study to the current version of manuscript on the functional identification of transporters. We will keep working on this aspect, attempting to dissect out the exact transport function of the transporters in the near future. In this current manuscript, we have discussed the limitations of our study in the last part of the manuscript.
Reviewer #1 (Recommendations For The Authors):
Minor comments
Line 35: ...appears to have evolved...
Line 67: remove first comma
Line 105: thereafter or therefore?
Line 130: define ACP
Line 131: define TMD
Response: We thank very much the reviewer for the suggestions, and we have revised the points in the current manuscript.
Figure 1: more information on APT1 would be helpful for readers to interpret the results from turboID e.g., consider showing an illustration showing, according to Karnataki et al 2007 that APT1 likely occupies all 4 membranes of the apicoplast. Also, according to DeRocher et al 2012, APT1 N-term and C-term are both cytosolically exposed, at least in the outermost membrane. The orientation in the other membranes is not known.
Response: We thank very much the reviewer for the suggestions. We analyzed the localization information of APT1 in T. gondii, based on the studies as the reviewer proposed (Karnataki, et al., 2007; DeRocher et al., 2012). The HA tag at the C-terminus of APT1 was distributed at the four membranes of the apicoplast, indicating that the topology of APT1 might be difficult to be defined at the membranes. Considering this information, we felt hesitant to clearly describe the topology in a schematic diagram about the protein APT1. Nevertheless, the TurboID tagging at the C-terminus of APT1 was an excellent model for identification of potential transporters localized at membranes of the apicoplast. We have put more information about the topology of APT1 in the manuscript, thus providing a better understanding of the proteomic results.
Figure 2: add a space between "T." and "gondii"
Figure 2: remove period between "Fitness" and "scores"
Figure 2: different fonts are used within the figure. Consider using only one font such as arial. Same for Figure 4.
Figure 2: "Fitness scores" is not bold in panel A but is bold in panel B.
Response: We thank very much the reviewer for the suggestions. We have revised the points in the current version of the manuscript.
Line 187: superscript -7
Line 249: Caution should be used in interpreting two bands as being a precursor and mature product without additional experiments to establish such a relationship. Consider using the term "might" rather than "appear to". The presence of multiple bands could be due to phenomena other than proteolytic processing e.g., alternative splicing, alternative initiator codons, etc.
Response: We thank very much the reviewer for the suggestions. We have revised the sentences in the current version of manuscript.
Line 291: define IPP
Figure 3E. The data points for KD strains appear to be positioned above the zero value on the y-axis. Is this correct?
Response: We thank very much the reviewer for the suggestions. We have rechecked the figure and replaced it with the correct one.
Figure 3 G/H legend. Please describe what a single data point represents e.g., the average of one field of view, the average of a certain number of fields of view, or something else? Are the data combined from three experiments or from a representative experiment?
Response: We thank very much the reviewer for the suggestions. Three independent experiments were performed with at least three replicates. At least 150 vacuoles were scored in each replicate, thus resulting in at least 9 data points in total. The data points were shown with the results from each replicate.
Line 325: define MEP and explain how it is connected to IPP
Response: We thank very much the reviewer for the suggestions. We have provided the information in the current version of the manuscript.
Lines 351-355: The authors refer to Figure 4D to support this statement, but presumably they mean 4E. Also, the authors use the terms C14, C16, and C18. They should more precisely use the terms myristic acid, palmitoleic acid, and trans_oleic acid if this is what they are referring to. Finally, the authors should determine if there is a statistically significant difference between levels of these fatty acids between AMT1 KD and AMT2 KD. If not, they should suggest there is an overall trend toward lower levels of these fatty acids in AMT2 KD parasites compared to AMT1 KD parasites.
Response: We thank very much the reviewer for the suggestions. We have revised the information in the current version of the manuscript.
Lines 363-364: The basis of this comment is unclear. Please clarify.
Lines 369-370: the authors have not shown that the observed lower levels of fatty acids are due to synthesis, as noted above
Response: We thank very much the reviewer for the suggestions. We have accordingly revised the information in the current version of the manuscript.
Line 383: Should be Figure S6D
Line 386: An entire section of the results is used to describe data that are entirely in a supplemental figure. Consider moving this data to a main figure.
Response: We thank very much the reviewer for the suggestions. We have transferred the data to the main figure in the current version of the manuscript.
Line 391: Consider using the term virulence instead of growth since now experiments were performed to specifically assess parasite growth in the infected mice.
Response: We thank very much the reviewer for the suggestions. We have revised the terms in the Results section.
Line 427: Perhaps the authors mean "...strong growth defect..." or ...strong growth impairment..."
Line 460-461: This statement is unclear. Please explain how strong backgrounds in proteomics have made it difficult to identify apicoplast transporters. Because they are low abundance? Because they are membrane proteins?
Response: We thank very much the reviewer for the suggestions. We have revised the corresponding sentences in the current version. The strong backgrounds in the proteomics resulted from the high activity and nonspecific labeling of biotin ligase fused with the apicoplast proteins.
518-521: It would be helpful for non-specialists if the authors explained how pyruvate is connected to IPP biosynthesis.
523: delete period after "Escherichia"
548-549: "We observed similar decreases in level of the MEP biosynthesis activity upon depletion of AMT1 and AMT2..." Reword this since no experiments were done to measure MEP biosynthesis activity.
Response: We thank very much the reviewer for the suggestions. We have accordingly revised the relevant sentences in the manuscript.
Reviewer #2 (Recommendations For The Authors):
Major points:
- The metabolomic data on fatty acid synthesis and isoprenoid levels is relevant but cannot inform about the function of the transporter, since any protein causing loss of the apicoplast would behave in such a manner, i.e. block the apicoplast pathways.
Response: We thank very much the reviewer for the comment. We agree with this comment. We have thus discussed these points in a subsection in the Discussion, pointing out some of the limitations in the study.
- Currently, the manuscript fails to directly prove what AMC1 and AMC2 transports, potentially pyruvate as suggested to putatively fuel FASII and MEP/DOXP. Further experimental approaches using exogenous complementation and/or metabolomic analyses using stable isotope labelling (for example) should potentially bring light to the putative functions of AMC1/2.
Response: We thank very much the reviewer for the comments. As described above, we attempted several approaches to find out the substrates that the AMT1 and AMT2 transports. However, we could not successfully express the proteins in E. coli strains, and we did not generate a T. gondii strain that a pyruvate sensor was properly targeted to the apicoplast. At the end of the Discussion, we have a subsection that discusses the limitations of this study. We hope that our future approaches will be able to tackle these difficulties on the substrate identification.
Furthermore, the authors have not considered other pathways of interest, like heme or lysophosphatidic acid (LPA)n synthesis, which are two other key pathway, which may be related to AMC1/2 function. Those proposed experiments represent an important body of work, required to bring light to their metabolic functions.
Response: We thank very much the reviewer for the comments. We thought about that, but we finally decided to mainly discuss two of the pathways that the transporters might participate in, since the transporters contain specific domains on the proteins sequences that potentially are associated with pyruvate.
Further, the authors might have partially missed some referencing and data about the apicoplast in their introduction (and potentially to address other facets of the apicoplast metabolic functions/capacities in regards to AMC1/2 function): the introduction referencing and explanations are somehow not fully exact/precise for the part of the apicoplast and its pathway: references about the apicoplast, discovery and origin are not citing the original work (that should be Wilson et al. 1996, McFadden et al. 1996, Kohler et al. 1997,), same for the discovery of FASII and MEP./DOXP (Waller 1998, Jomaa et al...). The introduction (and the study?) lacks information about other key functions of the apicoplast: heme synthesis, lysophosphatidic acid synthesis (using FASII products). The explanations about the roles of FASII/DOXP are partial and not fully citing important references: Krishnan et al. 2020, and Amiar et al. 2020 are also key to understanding how the role of FASII is metabolically flexible depending on nutrient content. A whole part on the fact that FASII is not only dispensible but can also become essential under metabolic adaptations conditions, are missing (Botté et al. 2013, Amiar et al. 2020, Primo et al. 2021). These novel important facets of parasite biology should be mentioned as well as directly linked to the author's topic. This is more minor but could bring new ideas to the authors.
Response: We thank very much the reviewer for the suggestions. We have revised the relevant part in the introduction.
We are grateful for the suggestions to improve the manuscript.
2:40 sie holt sich ihre meinung vom mainstream, von leuten die die AFD hassen
auch lustig wie sie sagt "ich lese auch nachrichten über kontroverse themen" bei 1:23 also irgendwie ist ihr schon bewusst dass diese themen "kontrovers" sind...
aber gleichzeitig lässt sie sich blenden von "gewaltenteilung" und "vielfalt" während alle mainstream-kanäle komplett gleichgeschaltet sind
also die konsumiert den ganzen tag nur die "nachrichten" von einer seite aber fühlt sich "ausgewogen informiert"...
echte normies halt *facepalm
though it has been a
An invitation to lead or join an expedition for the museum, thus cashing in themost desirable tag in the hunting world, was indeed flattering.
The fact that people were paid such large amounts (especially for the time) for these animals is even more disturbing. So many animals were probably killed with the intention of being sold to this exhibit, but were majority were probably rejected because they were not up to standards.
Author Response
The following is the authors’ response to the original reviews.
eLife assessment
This important study reports jAspSnFR3, a biosensor that enables high spatiotemporal resolution of aspartate levels in living cells. To develop this sensor, the authors used a structurally guided amino acid substitution in a glutamate/aspartate periplasmic binding protein to switch its specificity towards aspartate. The in vitro and in cellulo functional characterization of the biosensor is convincing, but evidence of the sensor's effectiveness in detecting small perturbations of aspartate levels and information on its behavior in response to acute aspartate elevations in the cytosol are still lacking.
We thank the reviewers and editors for the detailed assessment of our work and for their constructive feedback. Most comments have now been experimentally addressed in the revised manuscript, which we feel is substantially improved from the initial draft.
Public Reviews:
Reviewer #1 (Public Review):
In this manuscript, Davidsen and coworkers describe the development of a novel aspartate biosensor jAspSNFR3. This collaborative work supports and complements what was reported in a recent preprint by Hellweg et al., (bioRxiv; doi: 10.1101/2023.05.04.537313). In both studies, the newly engineered aspartate sensor was developed from the same glutamate biosensor previously developed by the authors of this manuscript. This coincidence is not casual but is the result of the need to find tools capable of measuring aspartate levels in vivo. Therefore, it is undoubtedly a relevant and timely work carried out by groups experienced in aspartate metabolism and in the generation of metabolite biosensors.
Reviewer #2 (Public Review):
In this work the IGluSnFR3 sensor, recently developed by Marvin et al (2023) is mutated position S72, which was previously reported to switch the specificity from Glu to Asp. They made 3 mutations at this position, selected a S72P mutant, then made a second mutation at S27 to generate an Asp-specific version of the sensor. This was then characterized thoroughly and used on some test experiments, where it was shown to detect and allow visualization of aspartate concentration changes over time. It is an incremental advance on the iGluSnFR3 study, where 2 predictable mutations are used to generate a sensor that works on a close analog of Glu, Asp. It is shown to have utility and will be useful in the field of Asp-mediated biological effects.
Reviewer #3 (Public Review):
In this manuscript, Davidsen and collaborators introduce jAspSnFR3, a new version of aspartate biosensor derived from iGluSnFR3, that allows monitoring in real-time aspartate levels in cultured cells. A selective amino acids substitution was applied in a key region of the template to switch its specificity from glutamate to aspartate. The jAspSnFR3 does not respond to other tested metabolites and performs well, is not toxic for cultured cells, and is not affected by temperature ensuring the possibility of using this tool in tissues physiologically more relevant. The high affinity for aspartate (KD=50 uM) allowed the authors to measure fluctuations of this amino acid in the physiological range. Different strategies were used to bring aspartate to the minimal level. Finally, the authors used jAspSnFR3 to estimate the intracellular aspartate concentration. One of the highlights of the manuscript was a treatment with asparagine during glutamine starvation. Although didn't corroborate the essentiality of asparagine in glutamine depletion, the measurement of aspartate during this supplementation is a glimpse of how useful this sensor can be.
Reviewer #1 (Recommendations For The Authors):
The authors should evaluate the effectiveness of the sensor in detecting small perturbations of aspartate levels and its behavior in response to acute aspartate elevations in the cytosol. In vivo aspartate determinations were performed exclusively in conditions that cause aspartate depletion. By means the use of mitochondrial respiratory inhibitors or aspartate withdrawal, it was determined the reliability of the sensor performing readings during relatively long periods, until reaching a steady-state of aspartate-depletion 12-60 hours later. Although in Hellweg and coworkers, it has been demonstrated that a related aspartate sensor could detect increases in aspartate in cell overexpressing the aspartate-glutamate GLAST transporter, the differences reported here between both sensors advise testing whether this aspect is also improved, or not, using jAspSNFR3.
Similarly, Davidsen et al. did not test if the sensor can be able to detect transient variations in cytosolic aspartate levels. In proliferative cells aspartate synthesis is linked to NAD+ regeneration by ETC (Sullivan et al., 2015, Cell), indeed the authors deplete aspartate using CI or CIII inhibitors but do not analyze if those are recovered, and increased, after its removal. Furthermore, the sequential addition of oligomycin and uncouplers could generate measurable fluctuations of aspartate in the cytosol.
We agree with the reviewer that only including situations of aspartate depletion in our cell culture experiments provided an incomplete evaluation of the utility of this biosensor. In the revised manuscript we provide three additional experiments using secondary treatments that restore aspartate synthesis to conditions that initially caused aspartate depletion. First, we conducted experiments where cells expressing jAspSnFR3/NucRFP were changed into media without glutamine, inducing aspartate depletion, with glutamine being replenished at various time points to observe if GFP/RFP measurements recover. As expected, glutamine withdrawal caused a decay in the GFP/RFP signal and we found that restoring glutamine caused a subsequent restoration of the GFP/RFP signal at all time points, with each fully recovering the GFP/RFP signal over time (Revised Manuscript Figure 2E). Next, we conducted the experiment suggested by the reviewer, testing whether the published finding, that oligomycin induced aspartate limitation can be remedied by co-treatment with electron transport chain uncouplers, could be visualized using jAspSnFR3 measurements of GFP/RFP. Indeed, after 24 hours of oligomycin induced aspartate depletion, treatment with the ETC uncoupler BAM15 dose dependently restored GFP/RFP signal (Revised Manuscript Figure 2G). Finally, we also measured whether the ability of pyruvate to mitigate the decrease in aspartate upon co-treated with rotenone (Figure 2B) could also be detected in a sequential treatment protocol after aspartate depletion. Indeed, after 24 hours of aspartate depletion by rotenone treatment, the GFP/RFP signal was rapidly restored by additional treatment with pyruvate (Revised Manuscript Figure 2, figure supplement 1C). Collectively, these results provide support for the utility of jAspSnFR3 to measure transient changes in aspartate levels in diverse metabolic situations, including conditions that restore aspartate to cells that had been experiencing aspartate depletion.
Reviewer #2 (Recommendations For The Authors):
Weaknesses: Sensor basically identical to iGluSnFR3, but nevertheless useful and specific. The results support the conclusions, and the paper is very straightforward. I think the work will be useful to people working on the effects of free aspartate in biology and given it is basically iGluSnFR3, which is widely used, should be very reproducible and reliable.
We appreciate the reviewer’s comment that sensor is useful for specific detection of aspartate. We agree that the advance of the paper is primarily in demonstrating its utility to measure aspartate, rather than any fundamental innovation on the biosensor approach. We hope the fact that jAspSnFR3 derives from a well validated biosensor (iGluSnFR3) will support its adoption.
Reviewer #3 (Recommendations For The Authors):
Although this is a well-performed study, I have some comments for the authors to address:
1) A red tag version of the sensor (jAspSnFR3-mRuby3) was generated for normalization purposes, with this the authors plan to correct GFP signal from expression and movement artifacts. I naturally interpret "movement artifacts" as those generated by variations in cell volume and focal plane during time-lapse experiments. However, it was mentioned that jAspSnFR3-mRuby3 included a histidine tag that may induce a non-specific effect (responses to the treatment with some amino acids). This suggests that a version without the tag needs to be generated and that an alternative design needs to be set for normalization purposes. A nuclear-localized RFP was expressed in a second attempt to incorporate RFP as a normalization signal. Here the cell lines that express both signals (sensor and RFP) were generated by independent lentiviral transductions (insertions). Unless the number of insertions for each construct is known, this approach will not ensure an equimolar expression of both proteins (sensor and RFP). In this scenario is not clear how the nuclear expression of RFP will help the correction by expression or monitor changes in cell volume. The authors may be interested in attempting a bicistronic system to express both the sensor and RFP.
The reviewer noted several potential issues concerning the use of RFP for normalization, which will be separated into sections below:
Movement artifacts:
We are glad the reviewer raised this issue since we see how it was confusingly worded. We have deleted the text “and movement artefacts” from the sentence.
His-tag and non-specific responses to some amino acids:
We also found it concerning that non-specific responses to amino acids could potentially contribute to our RFP normalization signal, and so we conducted additional experiments to address whether this was likely to be an issue in intracellular measurements. We first tested whether the non-specific signal was related to the histidine tag, or was intrinsic to the mRuby3 protein itself, by comparing the fluorescence response to a titration of histidine (which showed the largest effect of red fluorescence), aspartate, and GABA (structurally related to glutamate and aspartate, but lacking a carboxylate group) across a group of mRuby containing variants, with or without histidine tags. We replicated the non-specific signal originally observed in jAspSnFR3-mRuby3-His and found that another biosensor with a histidine tagged on the C terminus of mRuby3 had a similar response (iGlucoSnFR2.mRuby3-His), as did mRuby3-His alone, indicating that the aspect of being fused with jAspSnFR3 or another binding protein was not required for this effect. Additionally, we also compared the fluorescence response of lysates expressing mRuby2 and mRuby3 without histidine tags and found that the non-specific signal was essentially absent (Revised Manuscript Figure 1, figure supplement 4B-D). Collectively. These data support our original hypothesis that the histidine tag was responsible for the non-specific signal, alleviating concerns about more substantial protein design issues or with using nuc-RFP for normalization. Since we also found that measuring aspartate signal using GFP/RFP ratios from cells with linked the jAspSnFR3-Ruby3-His agreed with measurements from cells separately expressing jAspSnFR3 and nucRFP (without a His tag), and the amino acid concentrations needed to significantly alter His tagged Ruby3 signal are above those typically found in cells, we conclude that this is unlikely to be a significant factor in cells. Nonetheless, we have added all the relevant data to the manuscript to allow readers to make their own decision about which construct would be best for their purposes.
Original text:
"Surprisingly, the mRuby3 component responds to some amino acids at high millimolar concentrations, indicating a non-specific effect, potentially interactions with the C-terminal histidine tag (Figure 1—figure Supplement 2, panel B). Notably, this increase in fluorescence is still an order of magnitude lower than the green fluorescence response and it occurs at amino acid concentrations that are unlikely to be achieved in most cell types."
Revised text:
"Surprisingly, the mRuby3 fluorescence of affinity-purified jAspSnFR3.mRuby3 responds to some amino acids at high millimolar concentrations, indicating a non-specific effect (Figure 1—figure Supplement 4, panel A). This was determined to be due to an unexpected interaction with the C-terminal histidine tag and could be reproduced with other proteins containing mRuby3 and purified via the same C-terminal histidine tag (Figure 1—figure Supplement 4, panel B and C). Interestingly, a structurally related, non-amino acid compound, GABA, does not elicit a change in red fluorescence; indicating, that only amino acids are interacting with the histidine tag (Figure 1—figure Supplement 4, panel D). Nevertheless, most of our cell culture experiments were performed with nuclear localized mRuby2, which lacks a C-terminal histidine tag, and these measurements correlated with those using the histidine tagged jAspSnFR3-mRuby3 construct (Figure 1—figure Supplement 1 panel D)."
Lentiviral transductions
We agree that splitting the two fluorescent proteins across two expression constructs and infections effectively guarantees that there will not be equimolar expression of jAspSnFR3 and RFP, however we do not think equimolar expression is necessary in this context. The primary goal of RFP measurements in these experiments (and in experiments using the jAspSnFR3-mRuby3 fused construct) is to control for global alterations in protein expression that might confound the interpretation that a change in GFP fluorescence corresponds to a change in aspartate levels. While a bicistronic system is arguably a better approach to improve the similarity of expression of jAspSnFR3 and nuc-RFP in a cell, we only require that the cells have consistent expression of both proteins across all cells in the population, not that the expression of one necessarily be a similar molarity to the other. We accomplish consistent expression of proteins by single cell cloning after expression of jAspSnFR3 and nucRFP (or jAspSnFR3-mRuby3), and screening for clones that have high enough expression of both proteins such that they are well detected by standard Incucyte conditions. Given that our data do not identify an obvious downside to separate expression of jASPSnFR3 and nuc-RFP compared to the fused jAspSnFR3-mRuby3 construct (where the fluorescent proteins are truly equimolar) (Figure 2, Figure Supplement 1C), we elected to prioritize the separate jAspSnFR3 and nuc-RFP combination, which provides additional opportunities to measure cell number in the same experiment (see below).
2) The authors were interested in establishing the temporal dynamics of aspartate depletion by genetics and pharmaceutical means. For the inhibition of mitochondrial complex I rotenone and metformin were used. Although the assays are clearly showing aspartate depletion the report of cell viability is missing. Considering that glutamine deprivation induces arrest in cell proliferation, I think will be important to know the conditions of the cell cultures after 60 hours of treatment with such inhibitors.
We agree that ensuring that cells are still viable in conditions where aspartate is depleted, as determined by GFP/RFP in jAspSnFR3 expressing cells, is an important goal. To this end, we added a new experiment investigating the restoration of glutamine on the GFP/RFP signal at different time points after glutamine depletion (Revised Manuscript Figure 2E, see response to reviewer 1). One advantage of using the nuclear RFP as a normalization marker is that it also enables measurements of nuclei counts, a surrogate measurement for cell number. In the same glutamine depletion experiment we therefore measured cell counts using nuclear RFP incidences and confluency as measurements of cell proliferation/growth. In both cases, the arrest in cell proliferation upon glutamine withdrawal was obvious, as was the restoration of cell proliferation following glutamine replenishment, with the amount of growth delay corresponding to the length of glutamine withdrawal (Revised Manuscript Figure 2, Figure Supplement 2A-B). Nonetheless, there was no obvious lasting defects in restarting cell proliferation even after 12 hours of glutamine withdrawal, indicating that cell viability is preserved. In the case of mitochondrial inhibitors, we also observe even that after 24 hours of treatment with oligomycin or rotenone, restoration of aspartate synthesis from BAM15 or pyruvate, respectively, can also restore GFP/RFP signal, supporting the conclusion that cellular metabolism is still active in these conditions (Revised Manuscript Figure 2G; Revised Manuscript Figure 2, figure supplement 1C).
3) The pH sensitivity was checked in vitro with jAspSnFR3-mRuby3 and the sensor reported suitable for measurements at physiological pH. It would be an opportunity to revisit the analysis for pH sensitivity in cultured cells using an untagged version of jAspSnFR3 coupled, for example, to a sensor for pH.
We thank the reviewer for the suggestion and agree that pH effects on sensor signal could be a confounding factor in some conditions. Unfortunately, measuring intracellular pH is not trivial and using multiple fluorescent sensors that change simultaneously would be complex to interpret, particularly in the absence of controls to unambiguously control intracellular pH and aspartate concentrations. Thus, we believe that proper investigation of the variable of pH is beyond the scope of this study. Nonetheless, we agree that measuring the contribution of pH to sensor signal is an important goal for future work, particularly if deploying it in conditions likely to cause substantial pH differences, such as comparing compartmentalized signal of jAspSnFR3 in the cytosol and mitochondria. We have added the following italicized text to the conclusions section to underscore this point:
“Another potential use for this sensor would be to dissect compartmentalized metabolism, with mitochondria being a critical target, although incorporating the influence of pH on sensor fluorescence will be an important consideration in this context.”
4) While the authors take an interesting approach to measuring intracellular aspartate concentration, it will be highly desirable if a calibration protocol can be designed for this sensor. Clearly, glutamine depletion grants a minimal ("zero") aspartate concentration. However, having a more dynamic way for calibration will facilitate the introduction of this tool for metabolism studies. This may be achieved by incorporating a cultured cell that already expresses the transporter or by ectopic expression in the cells that have already been used.
We appreciate the suggestion and would similarly desire a calibration protocol to serve as a quantitative readout of aspartate levels from fluorescence signal, if possible. While we do calibrate jAspSnFR3 fluorescence in purified settings, conducting an analogous experiment intracellularly is currently difficult, if not impossible. While we have several methods to constrain the production rate of aspartate (glutamine withdrawal, mitochondrial inhibitors, and genetic knockouts of GOT1 and GOT2), we cannot prevent cells from decreasing aspartate consumption and so cannot get a true intracellular zero to aid in calibration. Additionally, the impermeability of aspartate to cell membranes makes it challenging to specifically control intracellular concentrations using environmental aspartate, and the best-known aspartate transporter (SLC1A3) is concentrative and so has the reciprocal problem. Considering these issues, we are wary of implying to readers that any specific fluorescence measurement can be used to directly interpret aspartate concentration given the many variables that can impact its signal, both related to the biosensor system itself (expression of jAspSnFR3, expression of Nuc-RFP, sensitivity and settings of the fluorescence detector) and based on cell intrinsic variability (differences in basal ASP levels, different sensitivity to treatments, influence of pH, etc.). We maintain that jAspSnFR3 has utility to measure relative changes in aspartate within a cell line across treatment conditions and over time, but absolute quantitation of aspartate still will require complementary approaches, like mass spectrometry, enzymatic assays, or NMR.
5) jAspSnFR3 seems to have the potential to be incorporated easily for several research groups as a main tool. In general, a minor correction to replace F/F with ΔF/F in the text.
Thank you for catching this error, the text has been edited accordingly.
Reviewer #3 (Public Review):
Summary:<br /> In this manuscript, Davidsen and collaborators introduce jAspSnFR3, a new version of aspartate biosensor derived from iGluSnFR3, that allows to monitor in real-time aspartate levels in cultured cells. A selective amino acids substitution was applied in a key region of the template to switch its specificity from glutamate to aspartate. The jAspSnFR3 does not respond to other tested metabolites and performs well, is not toxic for cultured cells, and is not affected by temperature ensuring the possibility of using this tool in tissues physiologically more relevant. The high affinity for aspartate (KD=50 uM) allowed the authors to measure fluctuations of this amino acid in the physiological range. Different strategies were used to bring aspartate to the minimal level. Finally, the authors used jAspSnFR3 to estimate the intracellular aspartate concentration.
Strengths:<br /> One of the highlights of the manuscript was a treatment with asparagine during glutamine starvation. Although didn`t corroborate the essentiality of asparagine in glutamine depletion, the measurement of aspartate during this supplementation is a glimpse of how useful this sensor can be.
Weaknesses:<br /> Although this is a well-performed study, I have some comments for the authors to address:<br /> 1-A red tag version of the sensor (jAspSnFR3-mRuby3) was generated for normalization purposes, with this the authors plan to correct GFP signal from expression and movement artifacts. I naturally interpret "movement artifacts" as those generated by variations in cell volume and focal plane during time-lapse experiments. However, it was mentioned that jAspSnFR3-mRuby3 included a histidine tag that may induce a non-specific effect (responses to the treatment with some amino acids). This suggests that a version without the tag needs to be generated and that an alternative design needs to be set for normalization purposes. A nuclear-localized RFP was expressed in a second attempt to incorporate RFP as a normalization signal. Here the cell lines that express both signals (sensor and RFP) were generated by independent lentiviral transductions (insertions). Unless the number of insertions for each construct is known, this approach will not ensure an equimolar expression of both proteins (sensor and RFP). In this scenario is not clear how the nuclear expression of RFP will help the correction by expression or monitor changes in cell volume. The authors may be interested in attempting a bicistronic system to express both the sensor and RFP.<br /> 2-The authors were interested in establishing the temporal dynamics of aspartate depletion by genetics and pharmaceutical means. For the inhibition of mitochondrial complex I rotenone and metformin were used. Although the assays are clearly showing aspartate depletion the report of cell viability is missing. Considering that glutamine deprivation induces arrest in cell proliferation, I think will be important to know the conditions of the cell cultures after 60 hours of treatment with such inhibitors.<br /> 3-The pH sensitivity was checked in vitro with jAspSnFR3-mRuby3 and the sensor reported suitable for measurements at physiological pH. It would be an opportunity to revisit the analysis for pH sensitivity in cultured cells using an untagged version of jAspSnFR3 coupled, for example, to a sensor for pH.<br /> 4-While the authors take an interesting approach to measuring intracellular aspartate concentration, it will be highly desirable if a calibration protocol can be designed for this sensor. Clearly, glutamine depletion grants a minimal ("zero") aspartate concentration. However, having a more dynamic way for calibration will facilitate the introduction of this tool for metabolism studies. This may be achieved by incorporating a cultured cell that already expresses the transporter or by ectopic expression in the cells that have already been used.
Author Response
The following is the authors’ response to the original reviews.
First of all, we'd like to thank the three reviewers for their meticulous work that enable us to present now an improved manuscript and substantial changes were made to the article following reviewers' and editors' recommendations. We read all their comments and suggestions very carefully. Apart from a few misunderstandings, all comments were very pertinent. We responded positively to almost all the comments and suggestions, and as a result, we have made extensive changes to the document and the figures. This manuscript now contains 16 principal figures and 15 figure supplements.
The number of principal figures is now 16 (1 new figure), and additional panels have been added to certain figures. On the other hand, we have added 7 additional figures (supplement figures) to answer the reviewers' questions and/or comments.
Main figures
▪ Figures 1, 4, 5, 10, 11, 12, 13, 14: unchanged ▪ Figure 7 and 8 were switched.
▪ Figure 2: we added panel F in response to reviewer 3's and request for sperm defect statistics
▪ Figure 3: the contrast in panel B has been taken over to homogenize colors
▪ Figure 6: This figure was recomposed. The WB on testicular extract was suppressed and we present a new WB allowing to compare the presence of CCDC146 in the flagella fraction. Using an anti-HA Ab, we demonstrate that the protein is localized in the flagella in epididymal sperm. Request of the 3 reviewers.
▪ Figure 7 (old 8): to avoid the issue of the non-specificity of secondary antibodies, we performed a new set of IF experiments using an HA Tag Alexa Fluor® 488-conjugated Antibody (anti-HA-AF488-C Ab) on WT and HA-CCDC146 sperm. These results are now presented in figure 7 panel A (new). The specificity of the signal obtained with the anti-HA-AF488-C Ab on mouse spermatozoa was evaluated by performing a statistical study of the density of dots in the principal piece of the flagellum from HA-CCDC146 and WT sperm. These results are now presented in figure 7 panel B (new). This study was carried out by analyzing 58 WT spermatozoa and 65 CCDC146 spermatozoa coming from 3 WT and 3 KI males. We found a highly significant difference, with a p-value <0.0001, showing that the signal obtained on spermatozoa expressing the tagged protein is highly specific. We have added a paragraph in the MM section to describe the process of image analysis. We finally present new images obtained by ExM showing no staining in the midpiece (figure 7C new). Altogether, these results demonstrate unequivocally the presence of the protein in the flagellum. Moreover, the WB was removed and is now presented in figure 6 (improved as requested).
▪ Figure 8. Was old figure 7
▪ Figure 9: figure 9 was recomposed and improved for increased clarity as suggested by reviewer 2 and 3.
▪ Figure 16 was before appendix 11
Figure supplements and supplementary files
▪ Figure 1-Figure supplement 1 New. Sperm parameters of the 2 patients. requested by editor (remark #1) by the reviewer 1 (Note #3)
▪ Figure 2-Figure supplement 1 new. Sperm parameters of the line 2 (KO animals) requested by the reviewer 1 (Note #5)
▪ Figure 4-Figure supplement 1 New. Experiment to evaluate the specificity of the human CCDC146 antibody. Minimal revision request and reviewer 1 note #8
▪ Figure 6-Figure supplement 1 New. Figure recomposed; Asked by reviewer 2 note #4 and reviewer 3
▪ Figure 8-Figure supplement 1 New. We now provide new images to show the non-specific staining of the midpiece of human sperm by secondary Abs in ExM experiments; Asked by reviewer 2
▪ Figure 10-Figure supplement 1 New. We added new images to show the non-specific staining of the midpiece of mouse sperm by secondary Abs in IF (panel B). Rewiever 1 note #9 and reviewer 2 note #5
▪ Figure 12-Figure supplement 1 New. Control requested by reviewer 3 Note #23
▪ Figure 13-Figure supplement 1 New. We provide a graph and a statistical analysis demonstrating the increase of the length of the manchette in the Ccdc146 KO. Requested by editor and reviewer 3 Note 24
▪ Figure 15-Figure supplement 1 New. Control requested by reviewer 2. Minor comments
▪ Figure supplementary 1 New. Answer to question requested by reviewer 2 note #1
All the reviewers' and editors’ comments have been answered (see our point to point response) and we resubmit what we believe to be a significantly improved manuscript. We strongly hope that we meet all your expectations and that our manuscript will be suitable for publication in "eLife". We look forward to your feedback,
Point by point answer
Please note that there has been active discussion of the manuscript and the summarize points below is the minimal revision request that the reviewers think the authors should address even under this new review model system. It was the reviewers' consensus that the manuscript is prepared with a lot of oversights - please see all the minor points to improve your manuscript.
All minimal revision requests have been addressed
Minimal revision request
1) Clinical report/evaluation of the two patients should be given as it was not described even in their previous study as well as full description of CCDC146.
We provide now a new Figure 1-figure supplement 1 describing the patients sperm parameters
2) Antibody specificity should be provided, especially given two of the reviewers were not convinced that the mid piece signal is non-specific as the authors claim. As both KO and KI model in their hands, this should be straightforward.
To validate the specificity of the Antibody, we transfected HEK cells with a human DDK-tagged CCDC146 plasmid and performed a double immunostaining with a DDK antibody and the CCDC146 antibody. We show that both staining are superimposable, strongly suggesting that the CCDC146 Ab specifically target CCDC146. This experiment is now presented in Figure 4-Figure supplement 1. Next, to avoid the issue of the non-specificity of secondary antibodies, we performed a new set of IF experiments using an HA Tag Alexa Fluor® 488-conjugated Antibody (anti-HA-AF488-C Ab) on WT and HA-CCDC146 sperm. These results are now presented in figure 7 panel A (new). The specificity of the signal obtained with the anti-HA-AF488-C Ab on mouse spermatozoa was evaluated by performing a statistical study of the density of dots in the principal piece of the flagellum from HA-CCDC146 and WT sperm. These results are now presented in figure 7 panel B (new). This study was carried out by analyzing 58 WT spermatozoa and 65 CCDC146 spermatozoa coming from 3 WT and 3 KI males. We found a highly significant difference, with a p-value <0.0001, showing that the signal obtained on spermatozoa expressing the tagged protein is highly specific. We have added a paragraph in the MM section to describe the process of image analysis. We finally present new images obtained by ExM showing no staining in the midpiece (figure 7C new). Altogether, these results demonstrate unequivocally the presence of the protein in the flagellum.
3) The authors should improve statistical analysis to support their experimental results for the reader can make fair assessment. Combined with clear demonstration of ab specificity, this lack of statistical analysis with very few sample number is a major driver of dampening enthusiasm towards the current study.
Several statistical analyses were carried out and are now included:
1) distribution of the HA signal in mouse sperm cells (see point 2 Figure 7 panel B)
2) quantification and statistical analyses of the defect observed in Ccdc146 KO sperm (figure 2 panel E)
3) Quantification and statistical analyses of the length of the manchette in spermatids 13-15 steps (Figure 13-Figure supplement 1 new)
4) The authors need to clarify (peri-centriolar vs. centriole)
In figure 4A, we have clearly shown that the protein colocalizes with centrin, a centriolar core protein in somatic cells. This colocalization strongly suggests that CCDC146 is therefore a centriolar protein, and this is now clearly indicated lines 211-212. However, its localization is not restricted to the centrioles and a clear staining was also observed in the pericentriolar material (PCM). The presence of a protein in PCM and centriole was already described, and the best example is maybe gamma-tubulin (PMID: 8749391).
or tone down (CCDC146 to be a MIP) of their claim/description.
Concerning its localization in sperm, we agree with the reviewer that our demonstration that CCDC146 is MIP would deserve more results. Because of that, we have toned down the MIP hypothesis throughout the manuscript. See lines 491495
Testis-specific expression of CCDC146 as it is not consistent with their data.
We have also modified our claim concerning the testis-expression of CCDC146. Line 176
Reviewer #1 (Recommendations For The Authors):
Major comments
1) As described in general comments, this study limits how the CCDC146 deficiency impairs abnormal centriole and manchette formation. The authors should explain their relationship in developing germ cells.
In fact, there are limited information about the relationship between the manchette and the centriole. However, few articles have highlighted that both organelles share molecular components. For instance, WDR62 is required for centriole duplication in spermatogenesis and manchette removal in spermiogenesis (Commun Biol. 2021; 4: 645. doi: 10.1038/s42003-021-02171-5). Another study demonstrates that CCDC42 localizes to the manchette, the connecting piece and the tail (Front. Cell Dev. Biol. 2019 https://doi.org/10.3389/fcell.2019.00151). These articles underline that centrosomal proteins are involved in manchette formation and removal during spermiogenesis and support our results showing the impact of CCDC146 lack on centriole and manchette biogenesis. This information is now discussed. See lines 596-603
2) The authors generated knock-in mouse model. If then, are the transgene can rescue the MMAF phenotype in CCDC146-null mice? This reviewer strongly suggest to test this part to clearly support the pathogenicity by CCDC146.
We indeed wrote that we created a “transgenic mice”, which was misleading. We actually created a CCDC16 knock-in expressing a tagged-protein. The strain was actually made by CRISPR-Cas9 and a sequence coding for the HA-tag was inserted just before the first amino acid in exon 2, leading to the translation of an endogenous HA-tagged CCDC146 protein. We have removed the word transgenic from the text and made changes accordingly (see lines 250-253). We can therefore not use this strain to rescue the MMAF phenotype as suggested by the reviewer.
3) Although the authors cite the previous study (Coutton et al., 2019), the study does not describe any information for CCDC146 and clinical information for the patients. The authors must show the results for clinical analysis to clarify the attended patients are MMAF patients without other phenotypic defects.
We have now inserted a table, indicating all sperm parameters for the patients harboring a mutation in the CCDC146 gene (Figure 1-Figure supplement 1) and is now indicated lines 159-160
4) The authors describe CCDC146 expression is dominant in testes, However, the level in testis is only moderate in human (Supp Figure 1). Thus, this description is not suitable.
In Figure 1-figure supplement 2 (old FigS1), the median of expression in testis is around 12 in human, a value considered as high expression by the analysis software from Genevestigator. However, for mouse, it is true that the level of expression is medium. We assumed that reviewer’s comment concerned testis expression in mouse. To take into account this remark, we changed the text accordingly. See line 176.
5) Although the authors mentioned that two mice lines are generated, only one line information is provided. Authors must include information for another line and provide basic characterization results to support the shared phenotype within the lines.
We now provide a revised Figure 2-figure supplement 1CD, presenting the second line and the corresponding text in the main text is found lines 178-183.
6) In somatic cells, the CCDC146 localizes at both peri-centriole and microtubule but its intracellular localization in sperm is distinguished. The authors should explain this discrepancy.
The multi-localization of a centriolar protein is already discussed in detail in discussion lines 520-526. We have written:
“Despite its broad cellular distribution, the association of CCDC146 with tubulin-dependent structures is remarkable. However, centrosomal and axonemal localizations in somatic and germ cells, respectively, have also been reported for CFAP58 [37, 55], thus the re-use of centrosomal proteins in the sperm flagellar axoneme is not unheard of. In addition, 80% of all proteins identified as centrosomal are found in multiple localizations (https://www.proteinatlas.org/humanproteome/subcellular/centrosome). The ability of a protein to home to several locations depending on its cellular environment has been widely described, in particular for MAP. The different localizations are linked to the presence of distinct binding sites on the protein…. “
7) Authors mention CCDC146 is a centriolar protein in the title and results subtitle. However, the description in results part depicts CCDC146 is a peri-centriolar protein, which makes confusion. Do the authors claim CCDC146 is centrosomal protein?
In figure 4A, we have clearly shown that the protein colocalizes with centrin, a centriolar core protein. This colocalization strongly suggests that CCDC146 is therefore a centriolar protein in somatic cells, and is now clearly indicated lines 211-212. However, its localization is not restricted to the centrioles and a clear staining was also observed in the pericentriolar material (PCM). The presence of a protein in PCM and centriole was already described and the best example is maybe gamma-tubulin (PMID: 8749391).
8) Verification of the antibody against CCDC146 must be performed and shown to support the observed signal are correct. 2nd antibody only signal is not proper negative control.
It is a very important remark. The commercial antibody raised against human CCDC146 was validated in HEK293-cells expressing a DDK-tagged CCDC146 protein. Cells were co-marked with anti-DDK and anti-CCDC146 antibodies. We have a perfect colocalization of the staining. This experiment is now presented in Figure 4-figure supplement 1 and presented in the text (lines 206-208).
9) In human sperm, conventional immunostaining reveals CCDC146 is detected from acrosome head and midpiece. However, in ExM, the signal at acrosome is not detected. How is this discrepancy explained? The major concern for the ExM could be physical (dimension) and biochemical (properties) distortion of the sample. Without clear positive and negative control, current conclusion is not clearly understood. Furthermore, it is unclear why the authors conclude the midpiece signal is non-specific. The authors must provide experimental evidence.
Staining on acrosome should always be taken with caution in sperm. Indeed, numerous glycosylated proteins are present at the surface of the plasma membrane regarding the outer acrosomal membrane for sperm attachment and are responsible for numerous nonspecific staining. Moreover, this acrosomal staining was not observed in mouse sperm, strongly suggesting that it is not specific.
Concerning the staining in the midpiece observed in both conventional and Expansion microscopy, it also seems to be nonspecific and associated with secondary Abs.
For IF, we now provide new images showing clearly the nonspecific staining of the midpiece when secondary Ab were used alone (see Figure 10-figure supplement 1B).
For ExM, we provide new images in Figure 8-figure supplement 1B (POC5 staining) showing a staining of the midpiece (likely mitochondria), although POC5 was never described to be present in the midpiece. Both experiments (CCDC146 and POC5 staining by ExM) shared the same secondary Ab and the midpiece signal was likely due to it.
Moreover, we now provide new images (figure 7C) in ExM on mouse sperm showing no staining in the midpiece and demonstrating that the punctuated signal is present all along the flagellum. Finally, we would like to underline that we now provide new IF results, using an anti-HA conjugated with alexafluor 488 and confirming the ExM results.
These points are now discussed lines 498-502 for acrosome and lines 503-511 for midpiece staining.
10) For intracellular localization of the CCDC146 in mouse sperm, the authors should provide clear negative control using WT sperm which do not carry the transgene.
This experiment was performed.
To avoid the issue of the non-specificity of secondary antibodies, we performed a new set of IF experiments using an HA Tag Alexa Fluor® 488-conjugated Antibody (anti-HA-AF488-C Ab) on WT and HA-CCDC146 sperm. These results are now presented in figure 7 panel A (new). The specificity of the signal obtained with the anti-HA-AF488-C Ab on mouse spermatozoa was evaluated by performing a statistical study of the density of dots in the principal piece of the flagellum from HA-CCDC146 and WT sperm. These results are now presented in figure 7 panel B (new). This study was carried out by analyzing 58 WT spermatozoa and 65 CCDC146 spermatozoa coming from 3 WT and 3 KI males. We found a highly significant difference, with a p-value <0.0001, showing that the signal obtained on spermatozoa expressing the tagged protein is highly specific. We have added a paragraph in the MM section to describe the process of image analysis. We finally present new images obtained by ExM showing no staining in the midpiece (figure 7C new). Altogether, these results demonstrate unequivocally the presence of the protein in the flagellum.
11) Current imaging data do not clearly support the intracellular localization of the CCDC146. Although western blot imaging reveal that CCDC146 is detected from sperm flagella, this is crude approach. Thus, this reviewer highly recommends the authors provide more clear experimental evidence, such as immuno EM.
We provide now a WB comparing the presence of the protein in the flagellum and in the head fractions; see new figure 6. We show that CCDC146 is only present in the flagellum fraction; The detection of the band appeared very quickly at visualization and became very strong after few minutes, demonstrating that the protein is abundant in the flagella. It is important to note that epididymal sperm do not have centrioles and therefore this signal is not a centriolar signal. We also now provide new statistical analyses showing that the immuno-staining observed in the principal piece is very specific (Figure 7B). Altogether, these results demonstrate unequivocally the intracellular localization of CCDC146 in the flagellum. This point is now discussed lines 480-489
12) Although sarkosyl is known to dissociate tubulin, it is not well understood and accepted that the enhanced detection of CCDC146 by the detergent indicates its microtubule inner space. Sperm axoneme to carry microtubule is also wrapped peri-axonemal components with structural proteins, which are even not well solubilized by high concentration of the ionic detergent like SDS.
We agree with the reviewer that the solubilization of the protein by sarkozyl is not a proof of the presence of the protein inside microtubule. Taking into account this point, the MIP hypothesis was toned down and we now discuss alternative hypothesis concerning these results; See discussion lines 490-497
13) SEM image is not suitable to explain internal structure (line 317-323).
We agree with the reviewers and changes were made accordingly. See lines 354-357
Minor comments
1) In main text, supplementary figures are cited "Supp Figure". And the corresponding legends are written in "Appendix - Figure". Please unify them.
Done Labelled now “Figure X-figure supplement Y”
2) Line 159, "exon 9/19" is not clear.
We have written now exons 9 and indicated earlier that the gene contains 19 exons
3) Line 188, "positive cells" are vague.
Positive was changed by “fluorescent”
4) Representative TUNEL assay image for knockout testes were not shown in Supp Figure 3B.
It was a mistake now Figure 2-figure supplement 2C
5) Please provide full description for "IF" and "AB" when described first.
Done
6) Line 262, It is unclear what is "main piece".
Changed to principal piece
7) Line 340, Although the "stage" information might be applicable, this is information for "seminiferous tubule" rather than "spermatid". This reviewer suggests to provide step information rather than stage information.
We agree with the reviewer that there was a confusion between “stage” and “step”. We change to step spermatids
8) Line 342, Step 1 is not correct in here.
OK corrected. now steps 13-15 spermatids
9) Line 803, "C." is duplicated.
Removed
10) Figure 3A, it will be good to mark the defective nuclei which are described in figure legends.
These cells are now indicated by white arrow heads
11) Figure 5, Please provide what MT stands for.
Now explained in the legend of figure 5
12) Figure 6. Author requires clear blot images for C. In addition, Panel B information is not correct. If the blot was performed using HA antibody, then how "WT" lane shows bands rather than "HA" bands?
The reviewer is correct. It was a mistake; The figure was recomposed and improved.
Reviewer #2 (Recommendations For The Authors):
Overall, editing oversights are present throughout the manuscript, which has made the review process quite difficult. Some repetitive figures can be removed to streamline to grasp the overall story easier. Some claims are not fully supported by evidence that need to tone down. Some figures not referenced in the main text need to be mentioned at least once.
All figures are now referenced in the text
Major comments:
1) 163-164 - Please clarify the claim that there is going to be an absence of the protein or nonfunctional protein, especially for the patient with a deletion that could generate a truncated protein at two third size of the full-length protein. Similarly, 35% of the protein level is present for the patient with a nonsense mutation. Some in silico structural analysis or analysis of conserved domains would be beneficial to support these claims.
Both mutations are predicted to produce a premature stop codons: p.Arg362Ter and p.Arg704serfsTer7, leading either to the complete absence of the protein in case of non-sense mediated mRNA decay or to the production of a truncated protein missing almost two third or one fourth of the protein respectively. CCDC146 is very well conserved throughout evolution (Figure supplementary 1), including the 3’ end of the protein which contains a large coil-coil domain (Figure 1B). In view of the very high degree of conservation, it is most likely that the 3’ end of the protein, absent in both subjects, is critical for the CCDC146 function and hence that both mutations are deleterious. This explanation is now added to the discussion. see lines 439-448
2) 173, 423 - Please clearly state a rationale of your mouse model design (i.e., why a mouse model that recapitulate human mutation is not generated) as the truncations identified in human patients are located further towards the C-terminus, and it is not clear whether truncated proteins are present, and if so, they could still be functional. Basically, the current mouse model supports the causality of the human mutations.
This is an important question, which goes beyond the scope of this article, and raises the question of how to confirm the pathogenicity of mutations identified by high-throughput sequencing. The production of KO or KI animals is an important tool to help confirm one’ suspicions but the first element to take into consideration is the nature of the genetic data.
Here we had two patients with homozygous truncating variants. In human, it is well established that the presence of premature stop codons usually induces non-sense mediated mRNA decay (NMD), inducing the complete absence of the protein or a strong reduction in protein production. In the unlikely absence of NMD in our two patients, the identified variants would induce the production of proteins missing 60% and 30% of their C terminal part. Often (and it is particularly true for structural proteins) the production of abnormal proteins is more deleterious than the complete absence of the protein (and it is most likely the purpose of NMD, to limit the production of abnormal “toxic” proteins). For these reasons, to try to recapitulate the most likely consequences of the human variants, without risking obtaining an even more severe effect, we decided to introduce a stop codon in the first exon in order to remove the totality of the protein in the KO mice.
The second element is to interpret the phenotype of the KO animals. Here, the human sperm phenotype is perfectly recapitulated in the KO mice.
Overall, we have strong genetic arguments in human and the reproduction of the phenotype in KO mice confirming the pathogenicity of the variants identified in men.
This point is now discussed see lines 433-438
3) Figure 6A - the labelling is misleading as it seems to suggest that the specific cells were isolated from the testes for RT-PCR.
We have modified the labelling to avoid any confusion.
Figure 6B -Signal of HA-tag is shown in WT, not in transgenic. Please check the order of the labels. Figure 6C - This blot is NOT a publication-quality figure. The bands are very difficult to observe, especially in lane D18. Because it is one of the important data of this study, replacing this figure is a must.
The figure has been completely remade, including new results. See new figure 6. Figure 6C was suppressed.
4) Supplementary fig 6 is also not a publication-level figure, and the top part seems largely unnecessary (already in the figure legend).
The figure has been completely remade as well (now Figure 6-Figure Supplement 1).
5) 261/267- The conclusion that mitochondrial staining in the flagellum (in both mice and humans) is non-specific is not convincing. Supplementary fig 8 shows that the signal from secondary only IF possibly extends beyond the midpiece - but it is hard to determine as no mitochondrial-specific staining is present. Either need to tone down the conclusion or provide supporting experimental evidence.
First, to avoid the issue of the non-specificity of secondary antibodies, we performed a new set of IF experiments using an HA Tag Alexa Fluor® 488-conjugated Antibody (anti-HA-AF488-C Ab) on WT and HA-CCDC146 sperm. These results are now presented in figure 7 panel A (new). The specificity of the signal obtained with the anti-HA-AF488-C Ab on mouse spermatozoa was evaluated by performing a statistical study of the density of dots in the principal piece of the flagellum from HA-CCDC146 and WT sperm. These results are now presented in figure 7 panel B (new). This study was carried out by analyzing 58 WT spermatozoa and 65 CCDC146 spermatozoa coming from 3 WT and 3 KI males. We found a highly significant difference, with a p-value <0.0001, showing that the signal obtained on spermatozoa expressing the tagged protein is highly specific. We have added a paragraph in the MM section to describe the process of image analysis. We finally present new images obtained by ExM showing no staining in the midpiece (figure 7C new). Altogether, these results demonstrate unequivocally the presence of the protein in the flagellum. These experiments are now described lines 271-279
Second, we provide new images of the signal obtained with secondary Abs only that shows more clearly that the secondary Ab gave a non-specific staining (Figure 10-Figure supplement 1B). This point is discussed lines 503-511
6) Figure 9 A - Please relate the white line to Fig. 9B label in X-axis. The information from Fig 9A+D and 9E+F are redundant. The main text nor the figure legends indicate why these specific two sperm were chosen for quantification and demonstrating the outcomes. One of them could be moved to supplementary information or removed, or the two could be combined.
As suggested by the reviewer, we have combined the two sperm to demonstrate that CCDC146 staining is mostly located on microtubule doublets. Moreover, the figure was recomposed to make it clearer.
Minor comments:
All of the supplementary figures are referred to as Supp Fig X in the text, however, they are actually titled Appendix - Figure X. This needs to be consistent.
The figures are now referred as figure supplement x in both text and figures
Line 125 - edit spacing.
We think this issue (long internet link) will be curated later and more efficiently by the journal, during the step of formatting necessary for publication.
144 - With which to study with which we studied?
We made the change as suggested.
151 - Supp Fig 1 - the text says that the gene is highly transcribed in human and mouse testes, but the information in the figure states that the level in mouse tissues is "medium"
We have corrected this mistake in the text; See line 176
165 - The two mutations are most likely deleterious. Please specifically mention what analyses done to predict the deleterious nature to support these claims.
Both variants, c.1084C>T and c.2112del, are extremely rare in the general population with a reported allele frequency of 6.5x10-5 and 6.5x10-06 respectively in gnomAD v3. Moreover, these variants are annotated with a high impact on the protein structure (MoBiDiC prioritization algorithm (MPA) score = 10, DOI: 10.1016/j.jmoldx.2018.03.009) and predicted to induce each a premature termination codon, p.(Arg362Ter) and p.(Arg704SerfsTer7) respectively, leading to the production of a truncated protein. This information is now given line 164-169
196-200/Figure 4 - As serum starved cells/basal body (B) are not mentioned in the main text, as is, Fig 4A would be sufficient/is relevant to the text. Please make the text reflect the contents of the whole figure, or re/move to supplement.
We agree with the reviewer that the full description of the figure should be in the text. We added two sentences to describe figure 4B see lines 217-218.
224 - spermatozoa (plural) fits better here, not spermatozoon
OK changed accordingly
236 - According to the figure legend, 6B is only showing data from the epididymal sperm, not postnatal time points; should be referencing 6C. Alignment of Marker label
As indicated above, the figure has been completely remade, including new results. See new figure 6. Figure 6C was suppressed. The corresponding text was changed accordingly see lines 249-266
255-256 - Referenced figure 7B3, however, 7B3 only shows tubulin staining, so no CCDC146 can be observed. Did authors mean to reference fig 7B as a whole?
Sorry for this mistake. We agree and the text is now figure 8B6 (figure 7 and 8 were switched)
305 - "of tubules" - I presume it is meant to be microtubules?
Yes; The text was changed as suggested
317-321 - a diagram of HTCA would be useful here
We have added a reference where HTCA diagram is available see line 363. Moreover, a TEM view of HTCA is presented figure 12A
322/Fig 11A - an arrow denoting the damage might be useful, as A1 and A3 look similar. The size of the marker bar is missing. Please update the information on figure legend.
Concerning, the comparison between A1 and A3, the take home message is that there is a great variability in the morphological damages. This point is now underlined in the corresponding text. We updated the size of the marker bar as suggested (200 nm). See line 365-367
323 - Please mark where capitulum is in the figure
Capitulum was changed for nucleus
Since Fig 11B2 is not referenced in the main text, it does not seem to add anything to the data, and could be removed/moved to supplement.
We added a sentence to describe figure 11B2 line 370
342-343 - manchette in step I is not seen clearly - the figure needs to be annotated better. However, DPY19L2 is absent in step I in the KO, but the main text does not reflect that - why is that?
We do not understand the remark of the reviewer “manchette in step I is not seen clearly”. The figure shows clearly the manchette (red signal) in both WT and KO (Figure 13 D1/D2).
For steps 13-15 WT spermatids, the size of the manchette decreases and become undetectable. In KO spermatids, the shrinkage of the manchette is hampered and in contrast continue to expand (Figure 13D2). We also provide a new Figure 13-figure supplement 1 for other illustrations of very long manchettes and a statistical analysis. In the meantime, the acrosome is strongly remodeled, as shown in figure 16-new, with detached acrosome (panel H). This morphological defect may induce a loss of the DPY19L2 staining (Figure 13 D2 stage I-III). This explanation is now inserted in the text line 396399
Figure 15B and 15C only show KO, corresponding images from the WT should be present for comparison.
WT images are now provided in Figure 1-figure supplement 1 new
Figure 12 - Figure 12 - JM?.
JM was removed. It does not mean anything
Figure 12C and Supplementary Fig 10 - structures need to be labelled, as it is unclear what is where
Done
338 - text mentions step III, but only sperm from step VII are shown in Figure 13
As suggested by reviewer 3, we changed stage by step. The text was modified to take into account this remark see lines 388-396
360 - This is likely supposed to say Supp Figure 11E-G, not 13??
Yes, it is a mistake. Corrected
388 Typo "in a in a".
Yes, it is a mistake. Corrected
820 - Fig 3 legend - in KO spermatid nuclei were elongated - could this be labelled by arrows? I am not convinced this phenotype is that different from the WT.
In fact, the nuclei of elongating KO spermatids are elongated and also very thin, a shape not observed in the WT; We have added arrow heads and modified the text to indicate this point line 200.
836 - Figure 5 legend says that in yellow is centrin, but that is not true for 5A, where the figure shows labelling for y-tubulin (presumably, according to the figure itself).
We have modified the text of the legend to take into account the remark
837- 5A supposedly corresponds to synchronized HEK293T cells, but the reasoning behind using synchronized cells is not mentioned at all in the main text; furthermore, how this synchronization is achieved is not explained in materials and methods (serum starvation? Thymidine block?).
Yes, figure 5A was obtained with synchronized cells. We have added one paragraph in the MM section. For cell synchronization experiments, cells underwent S-phase blockade with thymidine (5 mM, SigmaAldrich) for 17 h followed by incubation in a control culture medium for 5 h, then a second blockade at the G2-M transition with nocodazole (200 nM, Sigma-Aldrich) for 12 h. Cells were then fixed with cold methanol at different times for IF labelling. See line 224 for changes made in the result section and lines 700-704 for changes made in the MM section.
845- figure legend says that the RT-PCR was done on CCDC146-HA tagged mice, but the main text does not reflect that.
We made changes and the description of the KI is now presented before (line 240) the RT-PCR experiment (line 257).
949 - it is likely supposed to say A2, not B1 (B1 does not exist in Fig 15)
Yes, it is a mistake. Corrected
971 - Appendix Fig 3 legend - I believe that the description for B and C are swapped.
Yes, it is a mistake. Corrected
Furthermore, some questions to address in A would be: Which cross sections were from which animal/points? How many per animal? Were they always in the same location?
Yes, we have a protocol for arranging and orienting all testes in the same way during the paraffin embedding phase. The cross-sections are therefore not taken at random, and we can compare sections from the same part of the testis. The number of animals was already indicated in the figure legend (see line 1128)
Reviewer #3 (Recommendations For The Authors):
1) There are a number of grammatical and orthographical errors in the text. Careful proofreading should be performed.
We have sent the manuscript to a professional proofreader
2) The author should also check for redundancies between the introduction and the discussion.
The discussion has modified to take into account reviewers’ remarks. Nevertheless, we did our best to avoid redundancies between introduction and discussion.
3) Can the authors provide a rationale why they have chosen to tag their gene with an HA tag for localisation? One would rather think of fluorescent proteins or a Halo tag.
Because the functional domains of the protein are unknown, adding a fluorescent protein of 24 KDa may interfere with both the localization and the function of CCDC146. For this reason, we choose a small tag of only 1.1 KDa, to limit as such as possible the risk of interfering with the structure of the protein. This rational is now indicated in the manuscript lines 251-254. It is worth to note, that the tagged-strain shows no sperm defect, demonstrating that the HA-tag does not interfere with CCDC146 function.
4) In the abstract, line 53, "provide evidence" is not the right term for something that is just suggestive. The term "suggests" would be more appropriate.
The text was modified to take into account this remark
5) Line 74: "genetic deficiency" sounds strange here, do the authors mean simply "mutation"?
Infertility may be due to several genetic deficiency such as chromosomal defects (XXY (Klinefelter syndrome)), microdeletion of the Y chromosome or mutations in a single gene. Therefore, mutation is too restrictive. Nevertheless, we modified the sentence which is now “…or a genetic disorder including chromosomal or single gene deficiencies”
6) Lines 163-164: the authors describe the mutations (premature stop mutations) and say that they could either lead to complete absence of the gene product, or the expression of a truncated protein. Did they test this, for example, with some immuno blot analyses?
As stated above, unfortunately, we were unable to verify the presence of RNA-decay in these patients for lack of biological material.
7) Line 184 and Fig 2E: the sperm head morphologies should be quantitatively assessed.
We provide now a full statistical analysis of the observed defects: see new panel in Figure 2 F
8) Fig 3: The annotation should be more precise - KO certainly means CDCC146-KO. The colours of the IH panels is different, which attracts attention but is clearly a colour-adjustment artefact. Colours should be adjusted for the panels to look comparable. It would be also helpful to add arrowheads into the figure to point at the phenotypes that are highlighted in the text.
We have added Ccdc146 KO in all figures. We have added arrow heads to point out the spermatids showing a thin and elongated nucleus. Concerning adjustment of colors, we attempted to make images of panel B comparable. See new figure 3.
9) Fig 6A: the authors use RT PCR to determine expression dynamics of their gene of interested, and use actin (apparently) as control. However, actin and CDCC146 expression levels follow the same trend. How is the interpreted?
The reviewer did not understand the figure. The orange bars do not correspond to actin expression and the grey bars to Ccdc146 expression but both bars represent the mRNA expression levels of Ccdc146 relative to Actb (orange) and Hprt (grey) expression in CCDC146-HA mouse pups’ testes. We tested two housekeeping genes as reference to be sure that our results were not distorted by an unstable expression of a housekeeping gene. We did not see significant difference between both house keeping genes. Actin was not used.
10) In line 235, the authors suggest posttranslational modifications of their protein as potential cause for a slightly different migration in SDS PAGE as predicted from the theoretical molecular weight. This is not necessarily the case, some proteins do migrate just differently as predicted.
We have changed the text accordingly and now provide alternative explanation for the slightly different migration. See lines 258-259
11) The annotation of Fig 6 panels is problematic. First, why do the authors write "Laemmli" as description of the gel? It would be more helpful to write what is loaded on the gel, such as "sperm". Second, in panels B and C it would be helpful to add the antibodies used. It is not clear why there is a signal in the WT lane of panel B, but not in the HA lane (supposing an anti-HA antibody is used: why has WT a specific HA band?). In panel C, it is not clear why the blot that has so beautifully shown a single band in panel B suddenly gives such a bad labelling. Can the authors explain this? Also, they cut off the blot, likely because to too much background, but this is bad practice as full blots should be shown. In the current state, the panel C does not allow any clear conclusion. To make it conclusive, it must be repeated.
Several mistakes were present in this figure. This figure was recomposed. The WB on testicular extract was suppressed and we now present a new WB allowing to compare the presence of CCDC146 in the flagella and head fractions from WT and HA-CCDC146 sperm. Using an anti-HA Ab, we demonstrate that in epididymal sperm the protein is localized in the flagella only. See new figure 6. The corresponding text was changed accordingly.
12) The authors have raised an HA-knockin mouse for CDCC146, which they explained by the unavailability of specific antibodies. However, in Fig 7, they use a CDCC146 antibody. Can they clarify?
The commercial Ab work for HUMAN CCDC146 but not for MOUSE CCDC146. We have added few words to make the situation clearer, we have added the following information “the commercial Ab works for human CCDC146 only”. See line 240
13) In Fig 7A (line 258), the authors hypothesise that they stain mitochondria - why not test this directly by co-staining with mitochondria markers?
We chose another solution to resolve this question:
To avoid the issue of the non-specificity of secondary antibodies, we performed a new set of IF experiments using an HA Tag Alexa Fluor® 488-conjugated Antibody (anti-HA-AF488-C Ab) on WT and HA-CCDC146 sperm. These results are now presented in figure 7 panel A (new). The specificity of the signal obtained with the anti-HA-AF488-C Ab on mouse spermatozoa was evaluated by performing a statistical study of the density of dots in the principal piece of the flagellum from HA-CCDC146 and WT sperm. These results are now presented in figure 7 panel B (new). This study was carried out by analyzing 58 WT spermatozoa and 65 CCDC146 spermatozoa coming from 3 WT and 3 KI males. We found a highly significant difference, with a p-value <0.0001, showing that the signal obtained on spermatozoa expressing the tagged protein is highly specific. We have added a paragraph in the MM section to describe the process of image analysis. We finally present new images obtained by ExM showing no staining in the midpiece (figure 7C new). Altogether, these results demonstrate unequivocally the presence of the protein in the whole flagellum.
14) It seems that in both, Fig 7 and 8, the authors use expansion microscopy to localise CDCC146 in sperm tails. However, the staining differs substantially between the two figures. How is this explained?
In figure 8 we used the commercial Ab in human sperm, whereas in figure 7 we used the anti-HA Abs in mouse sperm. Because the antibodies do not target the same part of the CCDC146 protein (the tag is placed at the N-terminus of the protein, and the HPA020082 Ab targets the last 130 amino acids of the Cter), their accessibility to the antigenic site could be different. However, it is important to note that both antibodies target the flagellum. This explanation is now inserted see lines 304-312
15) Fig 8D and line 274: the authors do a fractionation, but only show the flagella fraction. Why?
Showing all fractions of their experiment would have underpinned the specific enrichment of CDCC146 in the flagella fraction, which is what they aim to show. Actually, given the absence of control proteins, the fact that the band in the flagellar fraction appears to be weaker than in total sperm, one could even conclude that there is more CDCC146 in another (not analysed) fraction of this experiment. Thus, the experiment as it stands is incomplete and does not, as the authors claim, confirm the flagellar localisation of the protein.
We agree with the reviewer’s remark. We provide now new results showing both flagella and nuclei fractions in new figure 6A. This experiment is presented lines 253-256
16) Line 283, Fig 9D,F: The description of the microtubules in this experiment is not easy to understand. Do the authors mean to say that the labelling shows that the protein is associated with doublet microtubules, but not with the two central microtubules? They should try to find a clearer way to explain their result.
As suggested by reviewer 2, we have changed the figure to make it clearer. The text was changed accordingly. See new figure 9 and new corresponding legend lines 1006.
17) Fig 9G - how often could the authors observe this? Why is the axoneme frayed? Does this happen randomly, or did the authors apply a specific treatment?
Yes, it happens randomly during the fixation process.
18) Line 300 and Fig 10A - the authors talk about the 90-kDa band, but do say anything about what they think this band is representing.
We have now added the following sentence lines 340-342: “This band may correspond to proteolytic fragment of CCDC146, the solubilization of microtubules by sarkosyl may have made CCDC146 more accessible to endogenous proteases.”
19) Fig 11A, lines 321-322: the authors write that the connecting piece is severely damaged. This is not obvious for somebody who does not work in sperm. Perhaps the authors could add some arrow heads to point out the defects, and briefly describe them in the text.
We realized from your remark that our message was not clear. In fact, there is a great variability in the morphological damages of the HTCA. For instance, the HTCA of Ccdc146 KO sperm presented in figure 10A2 is quite normal, whereas that in figure 10A4 is completely distorted. This point is now underlined in the corresponding text. See lines 367-369
We also added the size of the marker bar (200 nm), which were missing in the figure’s legend.
20) Line 323: it will be important to name which tubulin antibody has been used to identify centrioles, as they are heavily posttranslationally modified.
The different types of anti-tubulin Abs are described in the corresponding figure’s legend
21) Fig 11B - phenotypes must be quantified to make these observations meaningful.
We agree that a quantification would improve the message. However, testicular sperm are obtained by enzymatic separation of spermatogenic cells and the number of testicular sperm are very low. Moreover, not all sperm are stained. Taking these two points into account, it seems to us that quantification could be difficult to analyze. For this reason, the quantification was not done; however, it is important to note that these defects were not observed in WT sperm, demonstrating that these defects are cased by the lack of CCDC146. We have added a sentence to underline this point; See lines 374-375
22) Line 329: Figure 12AB - is this a typo - should it read Figure 12B?
We have split the panel A in A1 and A2 and changed the text accordingly. See line 378
23) Why are there not wildtype controls in Fig 12B, C?
We provide now as Figure 12-figure supplement 1, a control image for fig 12B. For figure 12C, the emergence of the flagellum from the distal centriole in WT is already shown in Fig 12A1
24) Fig 13: the authors write that the manchette is "clearly longer and wider than in WT cells" (lines 342-343). How can they claim this without quantitative data?
We now provide a statistical analysis of the length of the manchette. See figure 13-figure supplement 1A. We also provide a new a new image illustrating the length of the manchette in Ccdc146 KO spermatids; See Figure 13-figure supplement 1B.
Reviewer #3 (Public Review):
Male infertility is an important health problem. Among pathologies with multiple morphological abnormalities of the flagellum (MMAF), only 50% of the patients have no identified genetic causes. It is thus primordial to find novel genes that cause the MMAF syndrome. In the current work, the authors follow up the identification of two patients with MMAF carrying a mutation in the CCDC146 gene. To understand how mutations in CCDC146 lead to male infertility, the authors generated two mouse models: a CCDC146-knockout mouse, and a knockin mouse in which the CCDC146 locus is tagged with an HA tag. Male CCDC146-knockout mice are infertile, which proves the causative role of this gene in the observed MMAF cases. Strikingly, animals develop no other obvious pathologies, thus underpinning the specific role of CCDC146 in male fertility. The authors have carefully characterised the subcellular roles of CCDC146 by using a combination of expansion and electron microscopy. They demonstrate that all microtubule-based organelles, such as the sperm manchette, the centrioles, as well as the sperm axonemes are defective when CCDC146 is absent. Their data show that CCDC146 is a microtubule-associated protein, and indicate, but do not prove beyond any doubt, that it could be a microtubule-inner protein (MIP).
This is a solid work that defines CCDC146 as a novel cause of male infertility. The authors have performed comprehensive phenotypic analysis to define the defects in CCDC146 knockout mice. The manuscript text is well written and easy to follow also for non-specialists. The introduction and discussion chapters contain important background information that allow to put the current work into the greater context of fertility research. Overall, this manuscript provides convincing evidence for CCDC146 being essential for male fertility and illustrates this with a large panel of phenotypic observations. Together, the work provides important first insights into the role of a so-far unexplored proteins, CCDC146, in spermatogenesis, thereby broadening the spectrum of genes involved in male infertility.
Reviewer #2 (Public Review):
Summary:
The manuscript by Kaneda et al "FBXO24 ensures male fertility by preventing abnormal accumulation 2 of membraneless granules in sperm flagella" is a significant paper on the role of FBXO24 in murine male germ cell development and sperm ultrastructure and function. The body of experimental evidence that the authors present is extraordinarily strong in both breadth and depth. The authors investigate the protein's functions in male germ cells and sperm using a wide variety of approaches but focusing predominantly on their novel mouse model featuring deletion of the Fbxo24 gene and its product. Using this mouse, and a cross of it with another model that expresses reporters in the head and midpiece, they logically build from one experiment to the next. Together, their data show that this protein is involved in the regulation of membraneless electron-dense structures; loss of FBXO24 led to an accumulation of these materials and defects in the sperm flagellum and fertilizing ability. Interestingly, the authors found that several of the best-known components of electron-dense ribonucleoprotein granules that are found in the intermitochondrial cement and chromatoid body were not disrupted in the Fbxo24 knockout, suggesting that the electron-dense material and these structures are not all the same, and the biology is more complicated than some might have thought. They found evidence for the most changes in IPO5 and KPNB1, and biochemical evidence that FBXO24 and IPO5 could interact.
Strengths:
The authors are to be commended for the thoroughness of their experimental approaches and the extent to which they investigated impacts on sperm function and potential biochemical mechanisms. Very briefly, they start by showing that the Fbxo24 message is present in spermatids and that the protein can interact with SKP1, in a way that is dependent on its F-box domain. This points toward a potential function in protein degradation. To test this, they next made the knockout mouse, validated it, and found the males to be sterile, although capable of plugging a female. Looking at the sperm, they identified a number of ultrastructural and morphological abnormalities, which they looked at in high resolution using TEM. They also cross their model with RBGS mice so that they have reporters in both the acrosome and mitochondria. The authors test a variety of sperm functions, including motility parameters, ability to fertilize by IVF, cumulus-free IVF, zona-free-IVF, and ICSI. They found that ICSI could rescue the knockout but not other assisted reproductive technologies. Defects in male fertility likely resulted from motility disruption and failure to get through the utero-tubal junction but defects in acrosome exocytosis also were noted. The authors performed thorough investigations including both targeted and unbiased approaches such as mass spectrometry. These enabled them to show that although the loss of the FBXO24 protein led to more RNA and elevated levels of some proteins, it did not change others that were previously identified in the electron-dense RNP material.
The manuscript will be highly significant in the field because the exact functions of the electron-dense RNP materials have remained somewhat elusive for decades. Much progress has been made in the past 15 years but this work shows that the situation is more complex than previously recognized. The results show critical impacts of protein degradation in the differentiation process that enables sperm to change from non-descript round cells into highly polarized and compartmentalized mature sperm, with an equally highly compartmentalized flagellum. This manuscript also sets a high bar for the field in terms of how thorough it is, which reveals wide-ranging impacts on processes such as mitochondrial compaction and arrangement in the midpiece, the correct building of the major cytoskeletal elements in the flagellum, etc.
Weaknesses:
There are no real weaknesses in the manuscript that result from anything in the control of the authors. They attempted to rescue the knockout by expressing a FLAG-tagged Fbxo24 transgene, but that did not rescue the phenotype, either because of inappropriate levels/timing/location of expression, or because of interference by the tag. They also could not make anti-FBXO24 that worked for co-immunoprecipitation experiments, so relied on the FLAG epitope, an approach that successfully showed co-IP with IPO5 and SKP1.
ObjectIds An ObjectId is a special type typically used for unique identifiers. Here's how you declare a schema with a path driver that is an ObjectId: const mongoose = require('mongoose'); const carSchema = new mongoose.Schema({ driver: mongoose.ObjectId });
Certainly! The way it works is when you create a new note and associate it with a user, you will typically provide the user's unique identifier (ID) as the value for the user field. In a typical scenario, this user ID would come from the "user" collection in your database.
Here's a brief example in code:
```javascript // Assume you have a User model and you have obtained a user's ID const userId = "someUserId"; // This would be the actual ID of the user in your database
// Creating a new note and associating it with the user by providing the user ID const newNote = new Notes({ user: userId, title: "Sample Note", description: "This is a sample note.", tag: "General", deadline: new Date(), deadlinetime: "12:00 PM", });
// Save the new note to the database newNote.save() .then((savedNote) => { console.log("Note saved successfully:", savedNote); }) .catch((error) => { console.error("Error saving note:", error); }); ```
In this example, the user field of the new note is set to the userId obtained from the "user" collection. When you save this note to the database, the user field will store the provided user ID. Later, when you retrieve this note, you can use this user ID to identify which user the note is associated with.
It's important to ensure that the user ID you provide for the user field actually corresponds to an existing user in the "user" collection. This way, you maintain the relationship between the "notes" and "user" collections in your MongoDB database.
Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Recommendations For The Authors):
It is somewhat speculative that the structure represents the EIIa-bound regulatory state. There's a strong enough case that it should be analyzed in the discussion, but I don't think it is firmly established. Therefore, the title of the paper should be changed.
Our answer: Thank you for the comment. We have changed the title to “Mobile barrier mechanisms for Na+-coupled symport in an MFS sugar transporter”
Reading through the manuscript, it was challenging to distinguish what is new in the current manuscript and what has been done previously. There were a lot of parts where it was hard for me to identify the main point of the current study among all the details of previous studies. It would also benefit from shortening. For example:
-Page 6: Nb725 binding has already been characterized extensively in the very nice JBC paper earlier this year. It's important to test 725-4 for binding, but since it doesn't change the binding interaction, and probably wouldn't be expected to, the entire section could be written more succinctly. The main point, which is that 725-4 behaves like 725, is lost among all the details
Our answer: Thanks for this instructive suggestion. We have shortened the description in this section.
-Page 9-10. I don't understand what summarizing all of the results from the previous D59C studies adds to the current story. It's important because it provides an indication of the substrate binding site, but its mechanism of action does not seem relevant to the current work.
Our answer: We have shortened the description of the sugar-binding site and moved the previous Fig. 3b to supplementary figure sFig. 11. According to your comment about showing the location of the binding sites, which is also suggested by Reviewer #2, we modified Fig. 3 and added two panels to map the location of the bound Na+ in the inward-facing structure and the bound sugar in the outward-facing structure.
The sugar-binding site identified in the published structure is critical to construct the mobile barrier mechanism. The sugar-binding residues identified in the published structure provided essential data to support the conclusion that the sugar-binding pocket is broken in the inward-facing structure. Thus, this published structure is mechanistically relevant to the current study.
-Page 12. Too much summary of the previous outward structure. Since this is already part of the literature, it would be more efficient to reference the previous data when it is important to interpret the new data (or show as a figure).
Our answer: The introduction of the previous sugar-binding sit is important for the detailed comparison between the two states as discussed above, but we agree with this reviewer and have significantly shortened the paragraph by moving the detailed description into the legend to the sFig. 11.
-Instead of providing the PDB ID in figures of the current structure, just say "current work" or similar. Then it is obvious you are not citing a previous structure.
Our answer: To distinguish clearly the new data and published results, the citation of the cryoEM structure [PDP ID 8T60] has been completely removed from the main text but kept in sTable 1.
-An entire panel of Figure 3 is dedicated to ligand binding in a previous outward-facing structure.
Showing it in the overlay would be sufficient.
Our answer: It is the first time for us to show a structure with a bound-Na+. Fig. 3 also illustrates the spatial relationship between the sugar-binding pocket and the cation-binding pocket since both binding sites are determined now. As stated above, according to two reviewers’ comments, we have modified the Figures and the Fig. 3d is the overlay.
Please increase the size of the font in all figures. It should be 6-8 point when printed on a standard sheet of paper. Labels in Figure 3, distances in Figure 4, and everything in Figure 5 is hard to see.
Our answer: Thank you for the comments and the enlargement of the figure size and label font in all figures have been made.
Figure 2: would be helpful to show Figure S8 in the main text, orienting the reader to the approximate location of substrate binding. What is known about the EIIA-Glc binding interface? Has anyone probed this by mutagenesis? Where are these residues on the overall structure, and are they somewhere other than the nanobody interface?
Our answer: Thank you for this comment. We have added a panel for orienting the readers about the substrate location in MelB in Figure 3c. The sFig. 8 actually focuses on the details of Nb interactions with MelB. Our current data strongly supported the notion that the Nb-bound MelBSt structure mimics the EIIAGlc-bound MelB but is not structurally resolved, so we have tuned down our statement on EIIAGlc. There is one study suggesting the C-terminal tail helix may be involved in the EIIAGlc binding, which has been added to the discussion.
Can Figure 5 be split into 2 figures and simplified?
Our answer: thanks for the suggestion. We have split it into Figs. 5b and 6 and also moved the peptide mapping to the Fig 5a.
What is the difference between cartoon and ribbon rendering?
Our answer: Ribbon: illustrating the structure; cartoon: highlighting the positions with statistically significant protection or deprotection. The statistically significant changes are implied by the ribbon representation; Sphere: not covered by labeled peptides.
Can the panels showing the kinetic data be enlarged? I don't think they need to surround the molecule. An array underneath would be fine.
Our answer: We have enlarged all figures and labels. The placement of selected plots around the model could clearly show the difference in deuterium uptake rates between the transmembrane domain and extra-membrane regions. We will maintain this arrangement.
Do colors in panel A correspond with colors in panel B?
Our answer: The color usage in both are different. Now the two panels have been separated.
Do I understand correctly that in the HDX experiments, negative values indicate positions that exchange more quickly in the nanobody-free protein relative to the nanobody-bound protein?
Our answer: Your understanding is correct.
I assume some of this is due to the protein changing conformation, but some of it might be due to burial at the nanobody-binding interface. Can those peptides be indicated?
Our answer: Thank you for this comment. We have marked the peptide carrying the Nb-binding residues on uptake plots in Figs.6 and Extended Fig. 1. There are only three Nb-binding residues covered by many overlapping peptides. Most are not covered, either not carried by the labeled peptides (Tyr205, Ser206, and Ser207) or with insignificant changes (Pro132 and Thr133), except for Asp137, Lys138, and Arg141 which are presented in 8 labeled peptides.
Few buried positions in the outward-facing state are expected to be solvent in the inward-facing state; unfortunately, inward-facing state they are buried by Nb binding.
Make figure legends easier to interpret by removing non-essential methods details (like buffer conditions).
Our answer: We removed the detailed method descriptions in most figure legends. Thank you.
Check throughout for typos.
ie page 9 Lue Leu
Page 9 like likely
Our answer: We have corrected them. Thank you!
Reviewer #2 (Recommendations For The Authors):
I have mostly minor questions/remarks.
- Why not do the hdx-ms experiments in the presence of sugar? That would give a proper distinction between two conformational states, instead of an ensemble of states vs one state.
Our answer: MelB conformation induced by sugar is also multiple states, and likely most are outward-facing states and occluded intermediate states. This is also supported by the new finding of an inward state with low sugar affinity. The ideal design should be one inward and one outward to understand the inward-outward transition. We have not identified an outward-facing mutant while we can obtain the inward by the Nb. WT MelBSt with bound Na+ favors the outward-facing state. Although our design is not ideal, we do have one state vs a predominant outward-facing WT with bound Na+.
Minor comments:
• Fig 5 is misleading as the peptide number does not match with the amino acid sequence. I would suggest putting a heat map with coverage on top. Or showing deuterium uptake per peptide. See examples below.
Our answer: The peptide number should not match with sequence number. We have 155 overlapping peptides that cover the entire amino acid sequence including the 10-His tag, and there are 60 residues with no data because they are not covered by a labeled peptide. The residue positions that are covered by peptides are estimated by bars on the top. The cylinder length does not correspond to the length of the transmembrane helix, just for mapping purposes.
- Can the authors explain how they found that the Nbs bind to the cytoplasmic side (before obtaining the structure)?
Our answer: Our in vivo two-hybrid assay between the Nb and MelBSt indicated their interaction on the cytoplasmic surface of MelBSt, which is further confirmed by the melibiose fermentation and transport assay, where the transport activities were completely inhibited by intracellularly coexpressed Nb and MelBSt. Thanks for raising this question.
• The authors use the word "substrate" indifferently for sugar and Na+ binding, which is a bit confusing. Technically, only sugar is the substrate and Na+ is a ligand, or cotransported-ion, that powers the reaction of transport. This might sound like nit-picking but it can lead to misunderstandings (at some point I thought two sugars were transported, and then I was looking for the second Na+ binding site).
Our answer: We used to call the sugar and Na as co-substrate but we agree with this comment.
We have changed by using substrate for the cargo sugar and coupling cation for the driving cation.
• Abstract "only the inner barrier" - the is missing.
Thanks. We have corrected this.
• p.3 intro "and identified that the positive cooperativity of cation and melibiose, " something is missing.
Thanks again. We missed the “as the core symport mechanism”.
• P.6 Nb275_4 instead of Nb725_4
Thank you very much for your careful reading.
• P.7. Also, affinity affinities
Thank you very much. We changed to “; and also, the -NPG affinity decreased by 21~32-fold for both Nbs”
• P.8 " contains 417 MelBSt residues (positions 2-210, 219-355, and 364-432). This does not sum up to 417 residues.
Thanks for your critical reading. We changed 364-432 to 262-432.
• p.9 Lue 54
We have corrected it to Leu54.
• I find fig.3 hard to read. Can the authors show the Na+ binding pockets and sugar binding pockets within the structure? Especially figure 3b. why are the residues in different colors?
Our answer: We have moved Fig 3b into sFig. 11. We colored the residues in the previous Fig 3B to match the hosting helices. We have added two panels to show the location of both sugar and Na in the molecular. Thank you for your comments.
• Fig4 bcef. Colored circles at the end of the helices. What are they for?
Our answer: We revised the legend. “The paired helices involved in either barrier formation were highlighted in the same colored circles.”
• 86% coverage includes the his-tag - it would be good to clarify that.
Our answer: Yes, it includes the 10-His tag.
• Fig.7 - anti clockwise cycle of transport is counter-intuitive.
Our answer: We have re-arranged. Our model was constructed originally to explain efflux due to limited information at the earlier state. Now more data are available allowing us to explain inflow and active transport.
• Where are all the uptake plots per peptide for the HDX-MS data?
Our answer: We have added the course raw data and prepared all uptake plots for all 71 peptides with statistically significant changes as an Extended Fig. 1.
• P.22 protein was concentrated to 50 mg/mL. Really? That is a lot.
This is correct. We can even concentrate MelBSt protein to greater than 50 mg/ml.
• Have the authors looked into the potential role of lipids in regulating the conformational transition? Since the structure was obtained in nanodiscs, have they observed some unexplained densities? The role of lipid-protein interactions in regulating such transitions was observed for several transporters including MFS (Gupta K, et al. The role of interfacial lipids in stabilizing membrane protein oligomers. Nature. 2017 10.1038/nature20820. Martens C, et al. Direct protein-lipid interactions shape the conformational landscape of secondary transporters. Nat Commun. 2018 10.1038/s41467-018-06704-1.). Furthermore, I see the authors have already observed lipid specific functional regulation of MelB (ref: Hariharan, P., et al BMC Biol 16, 85 (2018). https://doi.org/10.1186/s12915-018-0553-0). A few words about this previous work, and even commenting on the absence of lipid-protein interactions in this current work is worthwhile.
Our answer: Thanks for this very relevant comment. We paid attention to the unmodelled densities. There is one with potential but it is challenging to model it. We have added a sentence “There is no unexplained density that can be clearly modeled by lipids.” in the method to address this concern.
Reviewer #3 (Recommendations For The Authors):
1) In the following sentence, the authors report high errors for the Kd value. The anti-Fab Nb binding to NabFab was two-fold poorer than Nb725_4 at a Kd value of 0.11 {plus minus} 0.16 μM. The figure however indicates that the error value is 0.016 µM. Pls correct.
Our answer: Thank you. You are correct. The error has been corrected. 0.16 ± 0.02 uM. In this revised manuscript, we present the data in nM units.
2) Is the stoichiometry of the MelB:Na+ symport clearly known in this transporter. It can be mentioned in the discussion with appropriate references.
Our answer: Yes, the stoichiometry of unity has been clearly determined, which was included in the second paragraph of the previous version.
3) In the last section of results, the authors seem to suggest a greater movement within their Cterminal helical bundle compared to N-terminal helices. Is there evidence to suggest an asymmetry in the rocker switch between the two states of the transporter?
Our answer: Our structural data revealed that the C-terminal bundle is more dynamic compared with the N-terminal bundle where hosts the residues for specific binding of galactoside and Na+. The HDX data showed that the most dynamic regions are the structurally unresolved C-terminal tail by either method, the conserved tail helix and the middle-loop helix. transmembrane helices are relatively less dynamic with similar distributions on both transmembrane bundles. Since the most dynamic regions are peripheral element associated with the C-terminal domain, it might give a wrong impression. With regard to the symmetric or asymmetric movement, which will certainly affect the dynamic interactions between the transporter and the lipids, we favor the notion that MelBSt performs symmetric movement during the rocker switch between inward and outward states at the least cost for the protein-lipids interaction.
4) Figure 1. Are the thermograms exothermic or endothermic? clarify
Our answer: In our thermograms, all positive peaks are exothermic due to the direct detection of the heat release by the TA instrument. We clarified this in Method and now we stress this in figure legends to avoid confusion.
5) Figure 4a,d. Please put in a membrane bilayer and depict cytosolic and extracellular compartments for clarity.
Thank you. We have added a bilayer and labeled the sidedness in this figure and other related figures.
6) Fig 7. Melibiose symport cannot be referred to as Melibiose efflux transport in the legend as the latter refers to antiport. Pls rectify.
Our answer: Influx and efflux are conventionally used to describe the direction of movement of a substrate. The use of symport and antiport indicates the directions of the coupling reaction for the cargo and cation. For the symporter MelB, melibiose efflux means that sugar with the coupled cation moves out, which is driven by the melibiose concentration. During the steady state of melibiose active transport, efflux rate = influx rate.
7) Page 11 "A common feature of carrier transporters". The authors can use either carriers or transporters. Need not use both simultaneously.
Sorry for overlooking this. We have deleted carriers. Thank you very much for your time.
8) Several typos were noticed in this manuscript. some are listed below. pls correct.
Page 4- last paragraph "Furthermore"
We have corrected it. Thank you again!
Page 7 - second para one repharse "affinity reduced by 21~32 fold/units.." pls clarify
Added 21~32 fold.
Page 9 - "so it is highly likely that inward-open conformation" pls correct.
We have corrected to “likely”.
Fig. S9c - correct the spelling "Distance".
We have corrected to “Distance”
text in draw.io diagrams
https://youtu.be/3Lru4k9Q55k?si=2TsGAi8iMEMunKii&t=15
I opened the image in Youtube and then under share link I choose the start at time ootion. Please note the three tags I added. Note, once you have created the required tag of physical-computing it will auotcomplete. Also, you need to hit enter after you type in each tag, be sure to check the tags got added, as you are being graded on your ability to tag web content.
root.unmount() Call root.unmount to destroy a rendered tree inside a React root. root.unmount(); An app fully built with React will usually not have any calls to root.unmount. This is mostly useful if your React root’s DOM node (or any of its ancestors) may get removed from the DOM by some other code. For example, imagine a jQuery tab panel that removes inactive tabs from the DOM. If a tab gets removed, everything inside it (including the React roots inside) would get removed from the DOM as well. In that case, you need to tell React to “stop” managing the removed root’s content by calling root.unmount. Otherwise, the components inside the removed root won’t know to clean up and free up global resources like subscriptions. Calling root.unmount will unmount all the components in the root and “detach” React from the root DOM node, including removing any event handlers or state in the tree. Parameters root.unmount does not accept any parameters. Returns root.unmount returns undefined. Caveats Calling root.unmount will unmount all the components in the tree and “detach” React from the root DOM node. Once you call root.unmount you cannot call root.render again on the same root. Attempting to call root.render on an unmounted root will throw a “Cannot update an unmounted root” error. However, you can create a new root for the same DOM node after the previous root for that node has been unmounted. Usage Rendering an app fully built with React If your app is fully built with React, create a single root for your entire app. import { createRoot } from 'react-dom/client';const root = createRoot(document.getElementById('root'));root.render(<App />); Usually, you only need to run this code once at startup. It will: Find the browser DOM node defined in your HTML. Display the React component for your app inside. index.jsindex.htmlApp.jsindex.js ResetFork91234567import { createRoot } from 'react-dom/client';import App from './App.js';import './styles.css';const root = createRoot(document.getElementById('root'));root.render(<App />); If your app is fully built with React, you shouldn’t need to create any more roots, or to call root.render again. From this point on, React will manage the DOM of your entire app. To add more components, nest them inside the App component. When you need to update the UI, each of your components can do this by using state. When you need to display extra content like a modal or a tooltip outside the DOM node, render it with a portal. NoteWhen your HTML is empty, the user sees a blank page until the app’s JavaScript code loads and runs:<div id="root"></div>This can feel very slow! To solve this, you can generate the initial HTML from your components on the server or during the build. Then your visitors can read text, see images, and click links before any of the JavaScript code loads. We recommend using a framework that does this optimization out of the box. Depending on when it runs, this is called server-side rendering (SSR) or static site generation (SSG). PitfallApps using server rendering or static generation must call hydrateRoot instead of createRoot. React will then hydrate (reuse) the DOM nodes from your HTML instead of destroying and re-creating them. Rendering a page partially built with React If your page isn’t fully built with React, you can call createRoot multiple times to create a root for each top-level piece of UI managed by React. You can display different content in each root by calling root.render. Here, two different React components are rendered into two DOM nodes defined in the index.html file: index.jsindex.htmlComponents.jsindex.js ResetFork99123456789101112import './styles.css';import { createRoot } from 'react-dom/client';import { Comments, Navigation } from './Components.js';const navDomNode = document.getElementById('navigation');const navRoot = createRoot(navDomNode); navRoot.render(<Navigation />);const commentDomNode = document.getElementById('comments');const commentRoot = createRoot(commentDomNode); commentRoot.render(<Comments />); You could also create a new DOM node with document.createElement() and add it to the document manually. const domNode = document.createElement('div');const root = createRoot(domNode); root.render(<Comment />);document.body.appendChild(domNode); // You can add it anywhere in the document To remove the React tree from the DOM node and clean up all the resources used by it, call root.unmount. root.unmount(); This is mostly useful if your React components are inside an app written in a different framework. Updating a root component You can call render more than once on the same root. As long as the component tree structure matches up with what was previously rendered, React will preserve the state. Notice how you can type in the input, which means that the updates from repeated render calls every second in this example are not destructive: index.jsApp.jsindex.js ResetFork99123456789101112import { createRoot } from 'react-dom/client';import './styles.css';import App from './App.js';const root = createRoot(document.getElementById('root'));let i = 0;setInterval(() => { root.render(<App counter={i} />); i++;}, 1000); It is uncommon to call render multiple times. Usually, your components will update state instead. Troubleshooting I’ve created a root, but nothing is displayed Make sure you haven’t forgotten to actually render your app into the root: import { createRoot } from 'react-dom/client';import App from './App.js';const root = createRoot(document.getElementById('root'));root.render(<App />); Until you do that, nothing is displayed. I’m getting an error: “Target container is not a DOM element” This error means that whatever you’re passing to createRoot is not a DOM node. If you’re not sure what’s happening, try logging it: const domNode = document.getElementById('root');console.log(domNode); // ???const root = createRoot(domNode);root.render(<App />); For example, if domNode is null, it means that getElementById returned null. This will happen if there is no node in the document with the given ID at the time of your call. There may be a few reasons for it: The ID you’re looking for might differ from the ID you used in the HTML file. Check for typos! Your bundle’s <script> tag cannot “see” any DOM nodes that appear after it in the HTML. Another common way to get this error is to write createRoot(<App />) instead of createRoot(domNode). I’m getting an error: “Functions are not valid as a React child.” This error means that whatever you’re passing to root.render is not a React component. This may happen if you call root.render with Component instead of <Component />: // 🚩 Wrong: App is a function, not a Component.root.render(App);// ✅ Correct: <App /> is a component.root.render(<App />); Or if you pass a function to root.render, instead of the result of calling it: // 🚩 Wrong: createApp is a function, not a component.root.render(createApp);// ✅ Correct: call createApp to return a component.root.render(createApp()); My server-rendered HTML gets re-created from scratch If your app is server-rendered and includes the initial HTML generated by React, you might notice that creating a root and calling root.render deletes all that HTML, and then re-creates all the DOM nodes from scratch. This can be slower, resets focus and scroll positions, and may lose other user input. Server-rendered apps must use hydrateRoot instead of createRoot: import { hydrateRoot } from 'react-dom/client';import App from './App.js';hydrateRoot( document.getElementById('root'), <App />); Note that its API is different. In particular, usually there will be no further root.render call.
Certainly! Let's break it down:
root.unmount()?root.unmount() is a function in React that you can use to clean up and remove a rendered tree inside a React root. It's like saying, "Hey React, I'm done with this part, please clean it up."
When do you need it?
You might need it when the DOM node where your React app lives is going to be removed by some other code. For example, if you have a tab panel made with jQuery that removes inactive tabs, calling root.unmount() helps React know that it should stop managing the components inside the removed tab.
What does it do?
When you call root.unmount(), it removes all the React components in that part of the app and disconnects React from the corresponding DOM node. This cleanup includes removing event handlers and state.
How to use it?
If your entire app is built with React, you typically create one root at the beginning, and you don't need to call root.unmount(). You set it up once, and React takes care of everything.
javascript
const root = createRoot(document.getElementById('root'));
root.render(<App />);
Cautions and Restrictions:
root.unmount(), you can't call root.render() again on the same root. You would need to create a new root for the same DOM node.If you're using server rendering, use hydrateRoot instead of createRoot for the initial setup.
Troubleshooting Tips:
root.render(<App />);.createRoot is valid.In simple terms, root.unmount() is like telling React, "I'm done with this part of the app, clean it up," and you typically use it in specific situations where parts of your app might be removed dynamically.
API ReferenceLegacy React APIscreateElementcreateElement lets you create a React element. It serves as an alternative to writing JSX.const element = createElement(type, props, ...children) Reference createElement(type, props, ...children) Usage Creating an element without JSX Reference createElement(type, props, ...children) Call createElement to create a React element with the given type, props, and children. import { createElement } from 'react';function Greeting({ name }) { return createElement( 'h1', { className: 'greeting' }, 'Hello' );} See more examples below. Parameters type: The type argument must be a valid React component type. For example, it could be a tag name string (such as 'div' or 'span'), or a React component (a function, a class, or a special component like Fragment). props: The props argument must either be an object or null. If you pass null, it will be treated the same as an empty object. React will create an element with props matching the props you have passed. Note that ref and key from your props object are special and will not be available as element.props.ref and element.props.key on the returned element. They will be available as element.ref and element.key. optional ...children: Zero or more child nodes. They can be any React nodes, including React elements, strings, numbers, portals, empty nodes (null, undefined, true, and false), and arrays of React nodes. Returns createElement returns a React element object with a few properties: type: The type you have passed. props: The props you have passed except for ref and key. If the type is a component with legacy type.defaultProps, then any missing or undefined props will get the values from type.defaultProps. ref: The ref you have passed. If missing, null. key: The key you have passed, coerced to a string. If missing, null. Usually, you’ll return the element from your component or make it a child of another element. Although you may read the element’s properties, it’s best to treat every element as opaque after it’s created, and only render it. Caveats You must treat React elements and their props as immutable and never change their contents after creation. In development, React will freeze the returned element and its props property shallowly to enforce this. When you use JSX, you must start a tag with a capital letter to render your own custom component. In other words, <Something /> is equivalent to createElement(Something), but <something /> (lowercase) is equivalent to createElement('something') (note it’s a string, so it will be treated as a built-in HTML tag). You should only pass children as multiple arguments to createElement if they are all statically known, like createElement('h1', {}, child1, child2, child3). If your children are dynamic, pass the entire array as the third argument: createElement('ul', {}, listItems). This ensures that React will warn you about missing keys for any dynamic lists. For static lists this is not necessary because they never reorder. Usage Creating an element without JSX If you don’t like JSX or can’t use it in your project, you can use createElement as an alternative. To create an element without JSX, call createElement with some type, props, and children: import { createElement } from 'react';function Greeting({ name }) { return createElement( 'h1', { className: 'greeting' }, 'Hello ', createElement('i', null, name), '. Welcome!' );} The children are optional, and you can pass as many as you need (the example above has three children). This code will display a <h1> header with a greeting. For comparison, here is the same example rewritten with JSX: function Greeting({ name }) { return ( <h1 className="greeting"> Hello <i>{name}</i>. Welcome! </h1> );} To render your own React component, pass a function like Greeting as the type instead of a string like 'h1': export default function App() { return createElement(Greeting, { name: 'Taylor' });} With JSX, it would look like this: export default function App() { return <Greeting name="Taylor" />;} Here is a complete example written with createElement: App.jsApp.js ResetForkimport { createElement } from 'react'; function Greeting({ name }) { return createElement( 'h1', { className: 'greeting' }, 'Hello ', createElement('i', null, name), '. Welcome!' ); } export default function App() { return createElement( Greeting, { name: 'Taylor' } ); } Show more And here is the same example written using JSX: App.jsApp.js ResetForkfunction Greeting({ name }) { return ( <h1 className="greeting"> Hello <i>{name}</i>. Welcome! </h1> ); } export default function App() { return <Greeting name="Taylor" />; } Both coding styles are fine, so you can use whichever one you prefer for your project. The main benefit of using JSX compared to createElement is that it’s easy to see which closing tag corresponds to which opening tag.
Certainly! Let's break down the information in a simpler way with examples:
createElement is a function in React that allows you to create a React element. It's an alternative to using JSX.Example:
javascript
const element = createElement('h1', { className: 'greeting' }, 'Hello');
Parameters:
Example:
javascript
createElement('h1', { className: 'greeting' }, 'Hello', createElement('i', null, 'World'));
Returns:
type, props, ref, and key.Example:
javascript
const element = createElement('h1', { className: 'greeting' }, 'Hello');
console.log(element.type); // 'h1'
console.log(element.props); // { className: 'greeting' }
Usage Caveats:
Example:
javascript
createElement('ul', {}, listItems); // Dynamic children as an array
Creating an Element Without JSX:
createElement to create elements.Example:
javascript
function Greeting({ name }) {
return createElement('h1', { className: 'greeting' }, 'Hello ', createElement('i', null, name), '. Welcome!');
}
Comparison with JSX:
createElement.Example:
javascript
function Greeting({ name }) {
return <h1 className="greeting">Hello <i>{name}</i>. Welcome!</h1>;
}
Rendering Your Own Component:
createElement to render your own React components.javascript
export default function App() {
return createElement(Greeting, { name: 'Taylor' });
}javascript
export default function App() {
return <Greeting name="Taylor" />;In summary, createElement is a foundational function in React for creating elements, and it is often used behind the scenes when JSX is transpiled. While it's useful, JSX provides a more readable syntax for creating React elements. You can choose the style that best fits your project.
How I Start My School Year: Lesson One,
PE is the field I am going into. After reading this article I have learned a couple new things. Responsibility, respectfulness, and gratefulness are very important in any classrroom. I like the things talked about, and even the videos of the frozen tag game he explained.
20% der Schneemasse auf der Nordhalbkugel bedecken Gebiete, die im Winter meist wärmer sind als 8°. In diesen Gebieten hat die Schneedecke in den letzten Jahrzehnten bereits deutlich abgenommen. Für ihre Zukunft ist jedes Zehntelgrad mehr oder weniger Erhitzung entscheidend. Der Verlust der Schneedecke führt zu Problemen bei der Wasserversorgung etwa der Donau und des Mississippi. https://www.liberation.fr/environnement/y-aura-t-il-encore-de-la-neige-en-2050-20240117_UPOQVWROIZEBVDQRD5JBRA4EH4/
Mehr zur selben Studie: https://hypothes.is/search?q=tag%3A%22Evidence%20of%20human%20influence%20on%20Northern%20Hemisphere%20snow%20loss%22
Die OPEC geht davon aus, dass sich die Nachfrage nach Öl in diesem Jahr um 2,25 Millionen Barrel pro Tag erhöhen wird. Für das kommende Jahr erwartet die OPEC eine Steigerung um 1,85 Millionen Barrel am Tag. Die Prognosen der OPEC liegen deutlich höher als die der IEA. Die USA haben in der zweiten Januarwoche mit mit 13,3 Millionen Barrel pro Tag einen neuen Rekord in der Ölproduktion aufgestellt. https://www.reuters.com/business/energy/oil-prices-edge-higher-opec-demand-estimate-while-cold-hits-us-output-2024-01-18/
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
We are thankful to the reviewers for the time and effort invested in assessing our manuscript and for their suggestions to improve it. We have now considered the points raised by them, carried out additional experiments, and modified the text and figures to address them. We feel that the new experiments and modifications have been able to solve all concerns raised by the reviewers and have improved the manuscript substantially, strengthening and extending our conclusions.
The main modifications include:
Reviewer #1 (Evidence, reproducibility and clarity (Required)):
__ __Suggestions:
Therefore, we have now also included a strain with the double overexpression of Oxr1 and Rtc5. Since overexpression of the proteins results in decreased assembly, it is more likely that this strain will show impaired growth under conditions that strongly rely on V-ATPase activity. Indeed, we observed that the overexpression of Oxr1 alone resulted in a slight growth defect in media containing high concentrations of ZnCl2 and the double overexpression strain showed an even further defect (Figure 6 A and C).
The manuscript would benefit from a well-labelled diagram showing the subunits of V-ATPase (e.g. in Figure 2D).
We agree with the reviewer and we have now added a diagram of the structure of the V-ATPase labeling the different subunits in Figure 2B.
The images of structures, especially in Figure 1-Supplement 1B, are not particularly clear and could be improved (e.g. by removing shadows or using transparency).
We are thankful to the reviewer for this suggestion. To improve the clarity of the structures in Figure 1 C and Figure 1 – Supplement 1A, we are now presenting the different subunits in the structures with different shades of blue and grey.
The authors should clearly describe the differences between Rtc5p and Oxr1p in terms of protein length, sequence identity, domain structure, etc.
We are thankful for this suggestion and we have now included a diagram of the domain architecture and protein length of Rtc5 and Oxr1, comparing with two human proteins containing a TLDc domain in Figure 5A. In addition, we have added the following paragraph describing the features of the proteins.
“Rtc5 is a 567 residue-long protein. Analysis of the protein using HHPred (Zimmermann et al., 2018), finds homology to the structure of porcine Meak7 (PDB ID: 7U8O, (Zi Tan et al., 2022)) over the whole protein sequence (residues 37-559). For both yeast Rtc5 and human Meak7 (Uniprot ID: Q6P9B6), HHPred detects homology of the C-terminal region to other TLDc domain containing proteins like yeast Oxr1 (PDBID: 7FDE), Drosophila melanogaster Skywalker (PDB ID: 6R82), and human NCOA7 (PDB ID: 7OBP), while the N-terminus has similarity to EF-hand domain calcium-binding proteins (PDB IDs: 1EG3, 2CT9, 1S6C6, Figure 5A). HHPred analysis of the 273 residue long Saccharomyces cerevisiae Oxr1, on the other hand, only detects similarity to TLDc domain containing proteins (PDB IDs: 7U80, 6R82, 7OBP), which spans the majority of the sequence of the protein (residues 71-273). The overall sequence identity between Oxr1 and Rtc5 is 24% according to a ClustalOmega alignment within Uniprot. The Alphafold model that we generated for Rtc5 is in good agreement with the available partial structure of Oxr1 (7FDE) (root mean square deviation (RMSD) of 3.509Å) (Figure 5 - S1 A), indicating they are structurally very similar, in the region of the TLDc domain. Taken together, these analyses suggest that Oxr1 belongs to a group of TLDc domain-containing proteins consisting mainly of just this domain like the splice variants Oxr1-C or NCOA7-B in humans (NP_001185464 and NP_001186551, respectively), while Rtc5 belongs to a group containing an additional N-terminal EF-hand-like domain and a N-myristoylation sequence, like human Meak7 (Finelli & Oliver, 2017) (Figure 5 A).”
Minor:
In Figure 4 supplement 1, the labels "I", "S", and "P" should be defined.
This has been clarified in the figure legend.
Please clarify what is meant by "switched labelling"
This refers to the SILAC vacuole proteomics experiments, for which yeast strains are grown in medium containing either L-Lysine or 13C6;15N2- L-Lysine to produce normal (‘light’) or heavy isotope-labeled (‘heavy’) proteins. This allows comparing two conditions. To increase the robustness of the comparisons, the experiments are done twice with both possible labeling schemes (condition A – light, condition B – heavy + condition A – heavy + condition B – light), which is commonly described as switched labeling or label switching.
We have exchanged the original sentence in the manuscript for:
“Performing the same experiments but switching which strain was labeled with heavy and light amino acids gave consistent results.”
The meaning of the sentence "Indeed, this was the case for both of them" is ambiguous.
We have now replaced this sentence with the following:
“Indeed, overexpression of either Rtc5 or Oxr1 resulted in increased growth defects in the context of Stv1 deletion (Figure 7 H and I).”
For Figure 1-Supplement 1B it is hard to see the crosslink distances.
We have updated this figure to improve the visibility of the cross-links. In addition, we now include a supplemental table (supplemental table 5) with a list of the Cα- Cα distances measured for all the crosslinks we mapped onto high-resolution structures.
The statement "The effects of Oxr1 are greater than those caused by Rtc5" requires more context. Is there a way of quantifying this effect for the reader?
We agree that this sentence was too general and vague. The effects caused by one or the other protein depend on the condition and the assay. We have thus deleted this sentence, and we think it is better to refer to the description of the individual assays performed.
The phrase "negative genetic interaction" should be clarified.
We have included in the text the following explanation of genetic interactions:
“A genetic interaction occurs when the combination of two mutations results in a different phenotype from that expected from the addition of the phenotypes of the individual mutations. For example, deletion of OXR1 or RTC5 has no impact on growth in neutral pH media containing zinc in a control background but improves the growth of RAV1 deletion strains (Figure 7 E and F), so this is a positive genetic interaction. On the other hand, overexpression of either Rtc5 or Oxr1 results in a growth defect in a background lacking Rav1 in neutral media containing zinc, a negative genetic interaction.”
* * In the sentence "Isogenic strains with the indicated modifications in the genome where spotted as serial dilutions in media with pH=5.5, pH=7.5 or pH=7.5 and containing 3 mM ZnCl2", "where" should be "were".
This has been corrected.
Figure 2D: the authors should consider re-coloring these models, as it is challenging to distinguish Rtc5p from the V-ATPase.
We have changed the coloring of this structure and added a diagram of the V-ATPase structure with the same coloring scheme to improve clarity.
Reviewer #1 (Significance (Required)):
The vacuolar protein interaction map alone from this manuscript is a nice contribution to the literature. Experiments establishing colocalization of Rtc5p to the vacuole are convincing, as is dependence of this association on the presence of assembled V-ATPase. Similarly, experiments related to myristoylation are convincing. The observed mislocalization of V-ATPases that contain Stv1p to the vacuole (which is also known to occur when Vph1p has been knocked out) upon knockout of Oxr1p is also extremely interesting. Overall, this is an interesting manuscript that contributes to our understand of TLDc proteins.
We are thankful to the reviewer for their appreciation of the significance of our work, including the interactome map of the vacuole as a resource and the advances on the understanding of the regulation of the V-ATPase by TLDc domain-containing proteins.
Reviewer #2 (Evidence, reproducibility and clarity (Required)):
Major points:
In addition, we have added new experiments to monitor V-ATPase assembly in intact cells, as suggested by the reviewer. Previous work has shown that in yeast, only subunit C leaves the vacuole membrane under conditions that promote disassembly, while the other subunits remain at the vacuole membrane (Tabke et al 2014). Our own experiments agree with what was published (Figure 3 D). We have thus monitored Vma5 localization to the vacuole under glucose or after shift to galactose containing media in cells lacking or overexpressing Rtc5 or Oxr1. We observed that cells overexpressing either TLDc domain protein show lower levels of Vma5 recruitment to the vacuole in glucose (Figure 6 D and E). Additionally cells lacking either Rtc5 or Oxr1 contain higher levels of Vma5 at the vacuole after 20 minutes in galactose medium (Figure 5 F and G). Thus, these results re-inforce our conclusions that Rtc5 and Oxr1 promote states of lower assembly.
Oxr1 and Rtc5 have very low sequence similarity. It would be helpful if the authors provided more detail on the predicted structure of the putative TLDc domain of Rtc5 and its relationship to the V-ATPase - Oxr1 structure. Is Rtc5 more closely related to established TLDc domain proteins in other organisms?
We have now included a diagram of the domain architecture of Rtc5 and Oxr1, and comparison to the features of other TLDc domain containing proteins in Figure 5 A, as well as a paragraph describing them:
“Rtc5 is a 567 residue-long protein. Analysis of the protein using HHPred (Zimmermann et al., 2018), finds homology to the structure of porcine Meak7 (PDB ID: 7U8O, (Zi Tan et al., 2022)) over the whole protein sequence (residues 37-559). For both yeast Rtc5 and human Meak7 (Uniprot ID: Q6P9B6), HHPred detects homology of the C-terminal region to other TLDc domain containing proteins like yeast Oxr1 (PDBID: 7FDE), Drosophila melanogaster Skywalker (PDB ID: 6R82), and human NCOA7 (PDB ID: 7OBP), while the N-terminus has similarity to EF-hand domain calcium-binding proteins (PDB IDs: 1EG3, 2CT9, 1S6C6, Figure 5A). HHPred analysis of the 273 residue long Saccharomyces cerevisiae Oxr1, on the other hand, only detects similarity to TLDc domain containing proteins (PDB IDs: 7U80, 6R82, 7OBP), which spans the majority of the sequence of the protein (residues 71-273). The overall sequence identity between Oxr1 and Rtc5 is 24% according to a ClustalOmega alignment within Uniprot. The Alphafold model that we generated for Rtc5 is in good agreement with the available partial structure of Oxr1 (7FDE) (root mean square deviation (RMSD) of 3.509Å) (Figure 5 - S1 A), indicating they are structurally very similar, in the region of the TLDc domain. Taken together, these analyses suggest that Oxr1 belongs to a subfamily of TLDc domain-containing proteins consisting mainly of just this domain like the splice variants Oxr1-C or NCOA7-B in humans (NP_001185464 and NP_001186551, respectively) , while Rtc5 belongs to a subfamily containing an additional N-terminal EF-hand-like domain and a N-myristoylation sequence, like human Meak7 (Finelli & Oliver, 2017) (Figure 5 A).”
The authors conclude vacuolar recruitment of Rtc5 depends on the assembled V-ATPase, based on deletion of different V1 and Vo domain subunits. However, these genetic manipulations likely cause a strong perturbation of vacuolar acidification; indeed, the images show drastically altered vacuolar morphology. To strengthen their conclusion, it would be helpful to show that Rtc5 recruitment is not blocked by inhibition of vacuolar acidification, and that conversely it is blocked by deletion of rav1.
We are thankful to the reviewer for this insightful suggestion and we have now performed both experiments suggested. The experiment regarding rav1Δ is now Figure 3C, and we observed that this mutation also disrupts Rtc5 localization to the vacuole. In addition, we decided to include an experiment showing the subcellular localization of Rtc5 after shifting the cells to galactose containing medium for 20 minutes, as a physiologically relevant condition that results in disassembly of the complex (Figure 3D). We observed that under these conditions Rtc5 re-localizes to the cytosol. This result is particularly interesting given that in yeast only subunit C (but not other V1 subunits) re-localizes to the cytosol under these conditions. In addition, the experiment using Bafilomycin A to inhibit the V-ATPase shows that Rtc5 is still localized at the vacuole membrane under conditions of V-ATPase inhibition (Figure 3 F). Taken together these results allowed us to strengthen our original interpretation that Rtc5 requires an assembled V-ATPase for its localization and extend it to the fact that the V-ATPase does not need to be active.
Reviewer #2 (Significance (Required)):
This is an interesting paper that confirms and extends previous findings on TLDc domain proteins as a novel class of proteins that interact with and regulate the V-ATPase in eukaryotes. The title seems to exaggerate the findings a bit, as the authors do not investigate V-ATPase (dis)assembly directly and only phenotypically describe altered subcellular localization of the Golgi V-ATPase in Oxr1-deleted cells. A recent structural and biochemical characterization of Oxr1 as a V-ATPase disassembly factor (PMID 34918374) somewhat limits the novelty of the results, but the function of Oxr1 in regulating subcellular V-ATPase localization and the identification of a second potential TLDc domain protein in yeast provide relevant insights into V-ATPase regulation. This paper will be of interest to cell biologists and biochemists working on lysosomal biology, organelle proteomics and V-ATPase regulation.
We thank the reviewer for the assessment of our work, and for recognizing the novel insights that we provide. Regarding the previous biochemical work on Oxr1 and the V-ATPase, we have clearly cited this work in the manuscript. In our opinion, our results complement and extend this article, showing that the function in disassembly is relevant in vivo. Additionally, this is only one of five major points of the article, the other four being
Reviewer #3 (Evidence, reproducibility and clarity (Required)):
Major comments
__1) Re: A cross-linking mass spectrometry map of vacuolar protein interactions (results) __ While XL-MS is a very powerful method, it is a high-throughput approach and there should be some kind of negative control in these experiments. In cross-linking experiments, non-cross-linked samples are usually used as negative controls. What was the negative control in cross-linking mass-spectrometry experiments here? If there was no negative control, how the specificity of interactions was evaluated? Maybe the authors analyzed the dataset for highly improbable interactions and found very few of them?
We fully agree that it is crucial to ensure the specificity of the interactions detected by XL-MS. To achieve this, one needs to control (1) the specificity of the data analysis (i.e. that the recorded mass spectrometry data are correctly matched to cross-linked peptides from the sequence database) and (2) the biological specificity (i.e. that the cross-linking captured natively occurring interactions).
To ascertain that criterion (1) is met, cross-link identifications are filtered to a pre-defined false-discovery rate (FDR) – an approach that the XL-MS field adopted from mass spectrometry-based proteomics. As a result, low-confidence identifications (e.g. cross-linked peptides that are only supported by a few signals in a given mass spectrum) are removed from the dataset. FDR filtering in XL-MS is a rather complex matter as it can be done at different points during data analysis and the optimal FDR cut-off depends on the specific scientific question at hand (for more details see for example Fischer and Rappsilber, Anal Chem, 2017). Generally speaking, an overly restrictive FDR cut-off would remove a lot of correct identifications, thereby greatly limiting the sensitivity of the analysis. On the other hand, a too relaxed FDR cut-off would dilute the correct identifications with a high number of false-positives, which would impair the robustness and specificity of the dataset. While many XL-MS study control the FDR on the level of individual spectrum matches, we opted for a 2% FDR cut-off on the level of unique residue pairs, which is more stringent (see Fischer and Rappsilber, Anal Chem, 2017). Our FDR parameters are described in the Methods section (Cross-linking mass spectrometry of isolated vacuoles - Data analysis). Of note, we have made all raw mass spectrometry data publicly available through the PRIDE repository (https://www.ebi.ac.uk/pride/ ; accession code PXD046792; login details during peer review: Username = reviewer_pxd046792@ebi.ac.uk, Password = q1645lTP). This will allow other researchers to re-analyze our data with the data analysis settings of their choice in the future.
To ascertain that criterion (2) is met, we mapped the identified cross-links onto existing high-resolution structures of vacuolar protein complexes. Taking into account the length of our cross-linking reagent, the side-chain length of the cross-linkable amino acids (i.e. lysines), and a certain degree of in-solution flexibility, cross-links can reasonably occur between lysines with a mutual Cα-Cα distance of up to 35 Å. Using this cut-off, the lysine-lysine pairs in the high-resolution structures we studied can be split into possible cross-linking partners (Cα-Cα distance 35 Å). Of all cross-links we could map onto high-resolution structures, 95.2% occurred between possible cross-linking partners. In addition, our cross-links reflect numerous known vacuolar protein interactions that have not yet been structurally characterized. These lines of evidence increase our confidence that our XL-MS approach captured genuine, natively occurring interactions. These analyses are described in more detail in the first Results sub-section (“A cross-linking mass spectrometry map of vacuolar protein interactions”).
In addition, the high purity of vacuole preparation is critical. How was it assessed by the authors?
We disagree that the purity of the vacuole preparation is critical for this analysis to be valid. The accuracy of the protein-protein interactions detected will depend on their preservation during sample preparation until the sample encounters the cross-linker, and the data analysis, as described above. The experiment would have been equally valid if performed on whole cell lysates without any enrichment of vacuoles, but the coverage of vacuolar proteins would have likely been very low. For this reason, we decided to use the vacuole isolation procedure to obtain better coverage of the proteins of this particular organelle. The use of the Ficoll gradient protocol (Haas, 1995) was based on that it is a protocol that yields strong enrichment of proteins annotated with the GO Term “vacuole” (Eising et al, 2019) and that it preserves the functionality of the organelle, as evidenced by its use for multiple functional assays (vacuole-vacuole fusion (Haas, 1995), autophagosome-vacuole fusion (Gao et al, 2018), polyphosphate synthesis by the VTC complex (Desfougéres et al, 2016), among others).
2) Re: Rtc5 and Oxr1 counteract the function of the RAVE complex (results)
Taken together, data, presented in this section of the manuscript, provide strong evidence that Rtc5 and Oxr1 negatively regulate V-ATPase activity, counteracting the V-ATPase assembly, facilitated by the activity of the RAVE complex. However, the complete deletion of the major RAVE subunit Rav1p was required to observe this effect in vivo in yeast. The other way to induce V-ATPase disassembly in yeast is glucose deprivation. It will be interesting to study if there is a synergistic effect between glucose deprivation and RTC5/OXR1 deletion on V-ATPase assembly, vacuolar pH, and growth of single oxr1Δ, rtc5Δ or double oxr1Δrtc5Δ mutants (OPTIONAL). Glucose deprivation is a more physiologically relevant condition than a deletion of an entire gene.
We would like to point out that an effect on assembly is observed without deleting the RAVE complex: deletions of Oxr1 or Rtc5 resulted in increased V-ATPase assembly in vivo in the presence of glucose and of the RAVE complex (Figures 5 D and E). We have now also added the experiments showing that the overexpression strains have a mild growth defect under conditions that force cells to strongly rely on V-ATPase activity (Figures 6 A and C).
Nevertheless, we agree that addressing the effect of changing the levels of Oxr1 and Rtc5 under low-glucose conditions is an interesting physiologically relevant question. We have now included growth assays and BCECF staining in medium containing galactose as the carbon source (Figures 5 – Supplement 1 B and C, and Figure 6 C and Figure 6- Supplement 1A). In addition, we have addressed the vacuolar localization of Vma5 in medium containing glucose or after shifting to medium containing galactose for 20 minutes, as a proxy for V-ATPase disassembly in intact cells (Figure 5 F and G, Figure 6 D and E). Taken together, these analyses reinforce our conclusions that both Rtc5 and Oxr1 promote an in vivo state of lower V-ATPase assembly, based on the following observations:
We have shifted the Panel D from the original Figure 6 – Supplement 1 to the main Figure (now Figure 7 – H and I). Regarding the title of the Figure, whether Supplemental Figures have titles or not will depend on the journal where the manuscript is published. For now, we have removed all titles from supplemental figures, as they are conceived to complement the main Figures.
4) Re: Figure 7 - supplement 1, Panel A. The proper assay to show that Stv1-mNeonGreen is functional is to express it in double mutant vph1Δstv1Δ to see if the growth defect is reversed. In addition, the vph1Δ growth defect is not changed (improved or worsened) in the presence of Stv1-mNeonGreen, so it means that the expression of Stv1-mNeonGreen does not further compromise the V-ATPase function, but it does not mean that it improves its function.
It is clear from the experiment suggested by the reviewer that they think that we have expressed Stv1-mNeonGreen from a plasmid. This was not the case, Stv1 was C-terminally tagged with mNeonGreen in the genome. It is thus the only expressed version in the strain. The experiment we have performed is thus equivalent to the one suggested by the reviewer, but for genomically expressed variants. For reference, the genotypes of all the strains used can be found in Supplemental Table 1.
5) Re: Figure 7 - supplement 2. This figure should be combined with Fig. 6- suppl 1, panel D as also mentioned above. The figure seems to lack some labels, and conclusions are not accurate as discussed below. However, this data provides important additional information about relationships between isoform-specific subunits of V-ATPase Vph1 and Stv1 and both Rtc5 and Oxr1 and should be repeated if it is not done yet to have a better idea about these relationships.
Panel B: Based on this picture, deletion of RTC5 has a negative genetic interaction with the deletion of VPH1, since double deletion mutant vph1Δ rtc5Δ grows worse than each individual mutant. Although it also means that there is no positive interaction, it is not the same.
Indeed, there is a negative genetic interaction between the deletion of RTC5 and VPH1. We have replaced the growth tests in this figure (Figure 8 – Supplement 2 A in the new manuscript) to show this negative genetic interaction better. This effect is reproducible, as shown in the repetitions of the experiments.
Panel C: Same as for panel B. Based on this picture, the deletion of OXR1 has a weak negative genetic interaction with the deletion of STV1, since double deletion mutant stv1Δ oxr1Δ grows worse than each individual mutant at 6 mM ZnCl2.
Panel D: Same as for panels B and C. Based on this picture, deletion of RTC5 has a negative genetic interaction with the deletion of STV1, since double deletion mutant stv1Δ rtc5Δ grows worse than each individual mutant at 6 mM ZnCl2. There is no label in the middle panel (growth conditions) and no growth assay data in the presence of CaCl2.
However, these results will be then in contradiction with the results from Figure 6 - Supplement 1, panel D, showing negative genetic interaction between the overexpression of Rtc5 or Oxr1 and deletion of Stv1, since both deletion and overexpression of Rtc5 or Oxr1 would have negative genetic interactions with Stv1.
For both Panels C and D (Now Figure 8 - Supplement 2 B and C). The effect pointed out by the reviewer (slightly stronger growth defect for the double mutants than for the single mutants) is very mild. We have attempted to make it more evident by assessing growth in medium with higher and lower concentrations of zinc and this was not possible. This is in contrast with the very clear positive genetic interaction that we observe between the deletion of OXR1 and VPH1 (Now Figure 8 H). This is the reason that we decided to report the lack of a positive genetic interaction instead of the presence of a negative one, as we do not want to draw conclusions based on results that are borderline detectable.
In addition, there is no label for the media in the middle panel, is it just YPAD pH=7.5, without the addition of any metals?
Indeed, the media is YPAD pH=7.5, without the addition of any metals. The line drawn above several images based on this media indicated this. Since this form of labeling appears to be confusing, we have now replaced it and placed the label directly above the image.
Why there is no growth assay in the presence of CaCl2, like in panels A and B?
Every growth test shown in the manuscript was performed including growth in YPD pH=5,5 as a control of a permissive condition for lack of V-ATPase activity, and then in YPD pH=7,5 including a broad range of Zinc Chloride and Calcium chloride concentrations. From all these pictures, the conditions where the differences among strains were clearly visible were chosen to assemble the figures. Conditions that did not provide any information for that particular experiment were not included in the figure to avoid making them unnecessarily large and crowded.
Re: Figure 7 - supplement 2, continued. How many times all these experiments were repeated? These experiments should be repeated at least 3 times, which is especially necessary for the experiments in panel C, because the effects are borderline. If results are reproducible and statistically significant, although small, the conclusion should be changed from "no positive genetic interactions" to "negative genetic interactions", which is more precise and informative.
All growth tests shown in the manuscript were repeated at least three times for the conditions shown. We are thankful to the reviewer for pointing out that this was not mentioned, and we have added this to the methods section. We have assembled a file with all repetitions of the shown growth tests and added it at the end of this file. In doing so, these are already available for the public. These repetitions show that all effects reported are reproducible. We will then discuss with the editors of the journal where this manuscript is published about the necessity of including it with the final article.
Regarding reporting the lack of a positive genetic interaction vs. a negative one, we have discussed this above. Shortly, for Panel B (Figure 8 – Supplement 2 A in the new manuscript) we have changed the conclusion to “negative genetic interaction” as adjusting the zinc chloride concentration allowed us to show this clearly and reproducibly, as shown by the repetitions of the experiments. For panels C and D (Now Figure 8 - Supplement 2 B and C), the effect is really mild and barely detectable, even when we tried a wide range of zinc chloride concentrations. For this reason, we would prefer to maintain the “no positive genetic interaction” conclusion.
Re: Methods. There is no description of yeast serial dilution growth assay at all. In addition, why the specific media (neutral pH, in the presence of high concentrations of calcium or zinc) was used is not explained either in the results or methods. Appropriate references should be included, for example, PMID: 2139726, PMID: 1491236.
We apologize for the oversight of the missing methods section, which we have now included.
Regarding the explanation of the media used, the following section was already a part of the results section, before the description of the first growth test:
“The V-ATPase is not essential for viability in yeast cells, and mutants lacking subunits of this complex grow similarly to a wt strain in acidic media. However, when cells grow at near-neutral pH or in the presence of divalent cations such as calcium and zinc, the mutants lacking V-ATPase function show a strong growth impairment (Kane et al, 2006).”
We have now replaced this with the following, more complete version:
“As a first approach for addressing the role of these proteins, we tested growth phenotypes related to V-ATPase function in strains lacking or overexpressing them. The V-ATPase is not essential for viability in yeast cells, and mutants lacking subunits of this complex grow similarly to a wt strain in acidic media, but display a growth defect at near-neutral pH the mutants (Nelson & Nelson, 1990). In addition, the proton gradient across the vacuole membrane generated by the V-ATPase energizes the pumping of metals into the vacuole, as a mechanism of detoxification. Thus, increasing concentrations of divalent cations such as calcium and zinc, generate conditions in which growth is increasingly reliant on V-ATPase activity (Förster & Kane, 2000; MacDiarmid et al, 2002; Kane, 2006).”
MINOR COMMENTS
Yeast proteins are named with "p" at the end, such as "Rtc5p".
This nomenclature rule is falling into disuse during the last decades, as the use of capitals vs lowercase and italics allows to distinguish between genes proteins and strains (OXR1 = gene, Oxr1 = protein, oxr1Δ = strain). As an example, I include a list of the latest papers by some of the major yeast labs around the world, all of which use the same nomenclature as we do (in alphabetical order). This list even includes some work in the field of the V-ATPase.
Re: Introduction. In the introduction it should be indicated that Rtc5 was originally discovered as a "restriction of telomere capping 5", using screening of temperature-sensitive cdc13-1 mutants combined with the yeast gene deletion collection [PMID: 18845848]. A couple of sentences should be written about the RAVE complex and its role in V-ATPase assembly.
We are thankful for this suggestion and we have now included both pieces of information in the introduction.
*“The re-assembly of the V1 onto the VO complex when glucose becomes again available, is aided by a dedicated chaperone complex known as the RAVE complex, which also likely has a general role in V-ATPase assembly (Seol et al, 2001; Smardon et al, 2002, 2014).” *
“In our cross-linking mass spectrometry interactome map of isolated vacuoles we found that the only other TLDc-domain containing protein of yeast, Rtc5, is a novel interactor of the V-ATPase. Rtc5 is a protein of unknown function, originally described in a genetic screen for genes related to telomere capping (Addinall et al, 2008)”
Re: The TLDc domain-containing protein of unknown function Rtc5 is a novel interactor of the vacuolar V-ATPase (results)
1) It is important to understand, that Oxr1 was co-purified before with the V1 domain of V-ATPase from a certain mutant strain, not wild-type yeast [PMID: 34918374]. It may explain why the authors did not identify it in their original protein-protein interactions screen here.
The structural work on the V1 domain bound to Oxr1 (Khan et al, 2022) showed that the binding of Oxr1 prevented V1 from assembling onto the Vo. Since our experiments rely on the purification of vacuoles, they should contain mainly only V1 assembled onto the VO, and not the free soluble V1. This is likely the reason that we do not detect Oxr1, in addition to it being less abundant. We have clarified this now in the manuscript and added the fact that Oxr1 was co-purified with a V1 containing a mutant version of the H subunit.
“In a previous study, Oxr1 was co-purified with a V1 domain containing a mutant version of the H subunit, and its presence prevented the in vitro assembly of this V1 domain onto the VO domain and promoted disassembly of the holocomplex (Khan et al., 2022). This is likely the reason why we do not detect Oxr1 in our experiments, which rely on isolated vacuoles and thus would only include V1 domains that are assembled onto the membrane. In addition, Oxr1 is less abundant in yeast cells than Rtc5 according to the protein abundance database PaxDb (Wang et al, 2015).”
2) It is a wrong conclusion that because Rtc5 was co-purified with both V1 and V0 domain subunits it interacts with the assembled V-ATPase, this does not exclude a possibility that Rtc5 also interacts with separate V1 sector or separate V0 sector of V-ATPase.
We agree with the reviewer that the co-purification of Rtc5 with both V1 and VO domain subunits does not necessarily mean that it interacts with the assembled V-ATPase. Thus, we have modified the text in this part to:
“The fact that we can co-enrich Rtc5 both with Vma2 and with Vph1 indicates that it can interact either with both the VO and V1 domains or with the assembled V-ATPase.”
However, other results throughout the manuscript can be taken into account to strengthen this idea:
*“Taking into account that Rtc5 is co-enriched with subunits of both the VO and V1 domain, and that it localizes at the vacuole membrane dependent on an assembled V-ATPase, we suggest that Rtc5 interacts with the assembled V-ATPase complex.” *
Re: Figure 1, Panel C. Is it possible to show individual proteins in different colors for clarity?
Panel D. How were cross-link distances measured? It is not obvious if you are not an expert in the field and it is not described in the methods.
We have modified Figure 1 C and Figure 1 – Supplement 1B (now Figure 1 – Supplement 1 A) to present the different subunits in the structures with different shades of blue and grey.
Furthermore, we have clarified the distance measurement approach in the methods section and in the legend of Fig 1D: “Ca-Ca distances were determined using the measuring function in Pymol v.2.5.2 (Schrodinger LLC).”
__Re: Figure 1 - Supplement 1, __
Panel A. What scientific information are we getting from this picture?
This panel was just a visual representation of the complexity of the protein network we obtained. Indeed, there was no specific scientific message, so we have decided to remove this panel from the revised manuscript.
Panel B. Why are these complexes shown separately from the complexes in Figure 1, panel C? Also, can individual proteins be colored differently here as well?
We did not want to overload Fig 1C, so we decided to show some of the protein complexes in Fig 1 – Supplement 1B. The most important information is the histogram showing that 95% of the mapped cross-links fall within the expected length range, and this is shown in the main Figure (Figure 1D). As stated above, we have adjusted the subunit coloring in Figure 1 C to improve clarity.
Re: Figure 3. It will be nice to show the localization of the untagged protein as well if antibodies are available (OPTIONAL).
Unfortunately, there are no available antibodies for either Rtc5 or Oxr1. This hinders us from detecting the endogenous untagged proteins. We would like to point out that we have been very careful in showing which tagged proteins are functional (C-terminally tagged Rtc5) and which are not (C-terminally tagged Oxr1), so that the reader can know how to interpret the localization data.
Re: Figure 4. Why different tags were used in panels A (GFP), C (msGFP2) and D
(mNeonGreen)?
In general, we prefer to use mNeonGreen as a tag for microscopy experiments because it is brighter and more stable, and msGFP2 as a tag for experiments involving Western blots because we have better antibodies available. There was a mistake in the labeling, and actually, all constructs labeled as GFP were msGFP2. We have now corrected this. Of note, we have tested the functionality of both tagged version (mNeonGreen and msGFP2).
Panels B and C. Were Rtc5 fusions detected using anti-GFP antibodies?
Indeed, Rtc5-msGFP2 was detected with an anti-GFP antibody. We have now indicated next to each Western blot membrane the primary antibody used. In addition, all antibodies are detailed in Supplemental Figure 3.
The authors should have full-size Western blots available, not just cut-out bands, as some journals and reviewers require them for publication.
For all western blots, we always showed a good portion of the membrane and not cut-out bands. The cropping was performed to avoid making figures unnecessarily large. The whole membranes are of course available and will be included in an “extended data file” if required by the journal.
Re: Figure 4 - Supplement 1, Panel A. Does "-" and "+" mean -/+ Azido-Myr?
Indeed. We have now added this label to the figure.
Panel B. There is no blot with a membrane protein marker (Vam3 or Vac8), it should be included.
We have replaced this western blot for a different repetition of this experiment in which a membrane protein marker was included. Of note, the two other repetitions of the experiment shown (Figure 4 – Supplement 1 panel C and Figure 4 panel C) also include both a membrane protein marker and a soluble protein marker.
Re: Figure 5. The title does not describe all results in this figure and should be modified accordingly.
The original data from Figure 5 is now separated into Figures 5 and 6 because of the additional experiments included during revisions. We have modified the Figure titles to be descriptive of the overall message of the Figures.
Panel C. Statistical significance value for *** should be indicated in the legend.
This has been indicated in the Figure legend.
It is not clear how many times yeast growth assays were repeated. Usually, all experiments should be done in triplicates or more.
All shown growth tests were performed at least three times for the conditions shown. We have now indicated this in the materials and methods section. In addition, we now provide in this response a file with all repetitions of growth tests, which will be appended to the article if deemed necessary by the editors.
Re: Figure 5 - supplement 1. No title
Re: Figure 5 - supplement 2. No title
Whether the supplemental Figures should have a title or not will depend on the style of the journal where the manuscript is finally published. The current idea of the supplemental Figures is that they complement the corresponding main Figure. For this reason, we have removed all titles from supplemental Figures.
Re: Figure 6. There is a typo on the second lane in the legend: "...the genome were", not "...the genome where".
This has been corrected.
Panel C. Why the analysis of BCECF vacuole staining of double mutants oxr1Δrav1Δ and rtc5Δrav1Δ is not shown? Was it done at all?
We had not included this piece of data, as we thought that the genetic interaction of RTC5 and OXR1 and rav1Δ was sufficiently well supported with the included data (growth tests in combination with the deletion, growth tests in combination with the overexpression, vacuole proteomics in combination with overexpression, and BCECF staining in combination with the overexpression). Because of the request of the reviewer, we have now included this experiment as Figure 7 G.
Re: Figure 6 - Supplement 2. Why were two different tags (2xmNG and msGFP2) used?
We tried both tags to see if one of them would be functional. Unfortunately, they both resulted in non-functional proteins, as shown by the corresponding growth tests.
Did the authors study N-terminally tagged Oxr1? Was it functional?
We have tagged Oxr1 N-terminally, and this unfortunately resulted in a protein that was not completely functional. We show below the localization of N-terminally mNeon-tagged Oxr1, under the control of the TEF1 promoter. The protein appears cytosolic (Panel A) but is not completely functional (Panel B). The localization of Oxr1 had already been misreported by using a tagged version that we now show to be non-functional. For this reason, we preferred not to include this data in the manuscript, to avoid again including in the literature subcellular localizations that correspond to non-functional or partially functional proteins.
Panel B. Results for the untagged TEF1pr-Oxr1 overexpression are not shown, thus tagged and untagged proteins can't be compared. Are they available? What is the promoter for the expression of 2xmNG fusion constructs?
Oxr1-2xmNG was C-terminally tagged in the genome, which means that the promoter is the endogenous one, it was not modified. For this reason, the correct controls are a strain expressing Oxr1 at endogenous levels (the wt strain) and a strain lacking Oxr1. Both controls were included in the Figure, and in all repetitions made of this experiment. For reference, all the genotypes of the strains used are found in Supplemental Table 1.
Re: Methods. Were vacuoles prepared differently for XL-MS and SILAC-based vacuole proteomics (there are different references) and why? Methods for XL-MS and quantitative SILAC-based proteomics can be placed together for clarity.
The basis for the method of vacuole purification is the same, from (Haas, 1995). This reference was included in both protocols that include vacuole purifications. However, modifications of this method were performed to fit the crosslinking method (higher pH, no primary amines) or to fit the SILAC labeling (combination of two differentially labeled samples in one purification). The reference for the vacuole proteomics (Eising et al 2022) corresponds to a paper in which the SILAC-based comparison of vacuoles from different mutant strains was optimized, and includes not only the vacuole purification but the growth conditions and downstream processing of the vacuoles.
Since both the SILAC-based vacuole proteomics and the XL-MS are multi-step methods, containing numerous parameters including the sample preparation, processing for MS, MS run and data analysis, we would prefer to keep them separate. We think this would allow a person attempting to reproduce these methods to go through them step by step.
What is CMAC dye? Why was it used to stain the vacuolar lumen?
We apologize for this oversight, we have included the definition of CMAC as 7-Amino-4-Chlormethylcumarin. It is a standard-used organelle marker for the lumen of the vacuole.
Some abbreviations (TEAB, ACN) are not explained.
We apologize for this oversight. We have now replaced these abbreviations with the full names of the compounds in the article.
What is 0% Ficoll?
We used the term 0% Ficoll, because this is the name given to the buffer in the original Haas 1995 paper on vacuole purifications. However, we agree that the term is misleading and we have now added the composition of the buffer (10 mM PIPES/KOH pH=6.8, 0.2 M Sorbitol).
Reviewer #3 (Significance (Required)):
The vacuolar-type proton ATPase, V-ATPase, is the key proton pump, that hydrolases ATP and uses this energy to pump protons across membranes. Amazingly, this proton pump and its function are conserved in eukaryotes from yeast to mammals. While V-ATPase structure and function have been studied for more than 30 years in various organisms, its regulation is not completely understood. The very recent discoveries of two new V-ATPase interacting proteins in yeast, first Oxr1 (OXidative Resistance 1), and now Rtc5 (Restriction of Telomere Capping 5), both the only two members of TLDc (The Tre2/Bub2/Cdc16 (TBC), lysin motif (LysM), domain catalytic) proteins in yeast, provide new insights in V-ATPase regulation in yeast, and because the interaction is conserved in mammals its relevance to mammalian V-ATPases regulation as well.
TLDc proteins are best known for their role in protection from oxidative stress, in particular in yeast and in the nervous system in mammals. The discovery of the novel Rtc5-V-ATPase interaction points to the role of V-ATPase not only in protection from oxidative stress but also in restriction of telomere capping in yeast and most likely higher species. The studies of other species also highlight the possible conserved role of V-ATPase in lifespan determination and Torc1 signaling, mediated through these interactions. Thus, the discovery of this new functionally important interaction between the second TLDc family member in yeast, Rtc5, and V-ATPase will shed light on the molecular mechanisms of all these essential biological processes and pathways.
In addition, because the authors performed a comprehensive proteomics protein-protein interaction study of the purified yeast vacuole it provides a valuable resource for all researchers who study vacuoles and/or related to them lysosomes.
The follow-up functional studies using the rav1Δ strain clearly demonstrated that Rtc5 and Oxr1 disassemble V-ATPase and counteract the function of V-ATPase assembly RAVE complex in vivo in yeast. Thus, they are essentially the first discovered endogenous eukaryotic protein inhibitors of V-ATPase. Moreover, because the authors obtained the evidence that Oxr1 is the regulator of the specific subunit isoform of V-ATPase Stv1p in vivo in yeast, it suggests that different TLDc proteins may regulate different specific V-ATPase subunit isoforms in cell- and tissue-specific manner in higher eukaryotes. The mechanism of this isoform-specific regulation in yeast and other species needs further investigation in the future.
Because of the conservation of the TLDc-V-ATPase interactions, all this information can be extrapolated to higher species, all the way to humans, in whom genetic mutations in various TLDc proteins are known to cause devastating diseases and syndromes.
We are thankful to the reviewer for their positive comments about the significance of our work.
Die IEA korrigiert ihre Prognose zum Ölbedarf 2024 nach oben. Die Nachfrage werde um 180.000 Barrel pro Tag steigern. Das Angebot wird die Nachfrage übertreffen. Die OPEC geht von einer noch größeren Steigerung aus https://www.reuters.com/business/energy/iea-raises-2024-oil-demand-growth-forecast-again-2024-01-18/
With purpose
prova + tag
Do cream
Prova dei tag
Blog: A Learner's Guide
Die Repubblica interviewt Carlo Buontempo, den Leiter des europäischen Klimaservice Copernicus. 20 23 wurden viele Anomalien beobachtet. Jeder Tag war mindestens ein Grad wärmer als in der Vergleichsperiode, fast die Hälfte der Tage 1,5 und zwei sogar zwei Grad. Der Juli war der heißeste je gemessene Monat. Es sei noch unklar, ob es sich dabei um Ausnahmen handelt oder um den Beginn einer neuen Phase. https://www.repubblica.it/economia/2024/01/15/news/clima_cambiamenti_climatici_caldo_record_carlo_buontempo_copernicus-421876070/
Accessing financial reports
image above this is missing for me (Decorative screenshot of a financial staement) also a typo in its Alt tag "staement"
aword cloud presenting key words can be used for pre-reading discussion on what thearticle might be about
“根据前人研究发现,tag cloud这一工具可以用在读前的预测/讨论阶段”。 运用前人研究证实工具确实可以运用于阅读教学。 下面就开始解释tag cloud这一工具如何具体使用,或者如何在阅读阶段使用。
Pre-Reading: Tag Cloud
读前阶段对应的是tag cloud这一工具
Tweeting has rapidly become an integral part of the conference scene, with a subset of attendees on Twitter providing real-time running commentary through a common “tag” (#mla09, for example),
This is very interesting to look at after the recent downfall of twitter. It makes me wonder if these people are still using twitter or if they've moved to another platform
[With Zeplin] we started to engage both UX and engineering teams in the same conversations and suddenly that opened our eyes to what was going on, and overall streamlined our build process.
may need new tag: combining/bringing different audiences together in the same conversation/context/tool
I don't know how much impact the "Design management" widget vs. "Design" object decision will have, except for the extremely small number of teams that work exactly like we do.
The third is the brain of the observer. This is also a strong element in film criticism where the camera is the third eye, the eye of the artificial narrator. The most intelligent film about the third eye spying on the action is `Snake Eyes,' where we last saw Gugino. (You may want to check my comments on that film to see what I mean.)
Author Response
eLife assessment
This important work describes the first high-resolution structure of HGSNAT, a lysosomal membrane protein required for the degradation of heparan sulfate (HS). Through careful structural analysis, this work proposes potential reasons why certain mutations in HGSNAT lead to lysosomal storage disorders and outlines the enzyme's catalytic mechanism. The experimental evidence presented provides incomplete support for the proposed molecular mechanism of the HS acetylation reaction and the impact of disease-causing mutations.
We thank the editors and reviewers for taking the time to provide a critical assessment of our manuscript. We appreciate the input and suggestions to improve the analysis. Included here are only our provisional responses. We will address the concerns raised in more detail and incorporate them in the revised version of the manuscript.
Reviewer #1 (Public Review):
This article by Navratna et al. reports the first structure of human HGSNAT in an acetyl-CoAbound state. Through careful structural analysis, the authors propose potential reasons why certain human mutations lead to lysosomal storage disorders and outline a catalytic mechanism. The structural data are of good quality, and the manuscript is clearly written. This study represents an important step toward understanding the mechanism of HGSNAT and is valuable to the field. I have the following suggestions:
We thank the reviewer for their encouraging and positive overall assessment of our work.
- The authors should characterize whether the purified protein is active. Otherwise, how does one know if the detergent used maintains the protein in a biologically relevant state? The authors should at least attempt to do so. If these prove to be challenging, at the very least, the authors should try a cell-based assay to demonstrate that the GFP tag does not interfere with the function.
Thank you for highlighting this concern. The cryo-EM sample was prepared without the exogenous addition of ligand, as noted in the manuscript; the acetyl-CoA that we see in the structure was intrinsically bound to the protein, indicating the ability of GFP-tagged HGSNAT protein to bind the ligand. We purified the protein at a pH optimal for acetyl-CoA binding, as suggested by Bame, K. J. and Rome, L. H. (1985) and Meikle, P. J. et al., (1995). Because we see acetyl-CoA in a structure obtained using a GFP fusion, we argue that GFP does not interfere with protein stability and ability to bind to the co-substrate. As demonstrated by existing literature HGSNAT catalyzed reaction is compartmentalized spatially and conditionally. The binding of acetyl-CoA happens towards the cytosol and is optimal at pH 7-0.8.0, while the transfer of the acetyl group to heparan sulfate occurs towards the luminal side and is optimal at pH 5.0-6.0. We are working on establishing a robust assay to study this complicated and compartmentalized acetyl transfer assay.
- In Figure 5, the authors present a detailed schematic of the catalytic cycle, which I find to be too speculative. There is no evidence to suggest that this enzyme undergoes isomerization, like a transporter, between open-to-lumen and open-to-cytosol states. Could it not simply involve some movements of side chains to complete the acetyl transfer?
The acetyl-CoA bound structure presented in the paper does not conclusively support a potential for isomerization and conformational dynamics. We agree with the reviewer that the reaction schematic presented in Figure 5 is speculative. We acknowledge in the discussion that our structure represents only a single step of the reaction, and defining the precise mechanism of acetyl transfer needs additional work. However, we will reword the discussion and change Figure 5 to address this concern raised by multiple reviewers.
Reviewer #2 (Public Review):
Summary:
This work describes the structure of Heparan-alpha-glucosaminide N-acetyltransferase (HGSNAT), a lysosomal membrane protein that catalyzes the acetylation reaction of the terminal alpha-D-glucosamine group required for the degradation of heparan sulfate (HS). HS degradation takes place during the degradation of the extracellular matrix, a process required for restructuring tissue architecture, regulation of cellular function, and differentiation. During this process, HS is degraded into monosaccharides and free sulfate in lysosomes.
HGSNAT catalyzes the transfer of the acetyl group from acetyl-CoA to the terminal non-reducing amino group of alpha-D-glucosamine. The molecular mechanism by which this process occurs has not been described so far. One of the main reasons to study the mechanism of HGSNAT is that multiple mutations spanning the entire sequence of the protein, such as nonsense mutations, splicesite variants, and missense mutations lead to dysfunction that causes abnormal accumulation of HS within the lysosomes. This accumulation is a cause of mucopolysaccharidosis IIIC (MPS IIIC), an autosomal recessive neurodegenerative lysosomal storage disorder, for which there are no approved drugs or treatment strategies.
This paper provides a 3.26A structure of HGSNAT, determined by single-particle cryo-EM. The structure reveals that HGSNAT is a dimer in detergent micelles and a density assigned to acetylCoA. The authors speculate about the molecular mechanism of the acetylation reaction, map the mutations known to cause MPS IIIC on the structure and speculate about the nature of the HGSNAT disfunction caused by such mutations.
Strengths:
The description of the architecture of HGSNAT is the highlight of the paper since this corresponds to the first description of the structure of a member of the transmembrane acyl transferase (TmAT) superfamily. The high resolution of an HGSNAT bound to acetyl-CoA is an important leap in our understanding of the HGSNAT mechanism. The density map is of high quality, except for the luminal domain. The location of the acetyl-CoA allows speculation about the mechanistic role of multiple residues surrounding this molecule. The authors thoroughly describe the architecture of HGSNAT and map the mutations leading to MPS IIIC. The description of the dimeric interphase is a novel result, and future studies are left to confirm the importance of oligomerization for function.
We thank the reviewer for their time and for highlighting both the quality and novelty of the structure presented in this work.
Weaknesses:
Apart from the cryo-EM structure, the article does not provide any other experimental evidence to support or explain a molecular mechanism. Due to the complete absence of functional assays, mutagenesis analysis, or other structures such as a ternary complex or an acetylated enzyme intermediate, the mechanistic model depicted in Figure 5 should be taken with caution.
Thank you for pointing out this concern. The proposed mechanistic model in Figure 5 is a hypothesis based on previously reported biochemical characterization of HGSNAT by Rome & Crain (1981), Rome et al, (1983), Miekle et al., (1995) and Fan et al., (2011). However, we agree with the reviewer that this schematic is not experimentally proven and is speculative at best. Especially because our structure presents only a single step of the reaction, which does not conclusively support either ping-pong or random-order bi-substrate reactions. We will rephrase this section of our discussion and edit Figure 5 to address this concern.
The authors discuss that H269 is an essential residue that participates in the acetylation reaction, possibly becoming acetylated during the process. However, there is no solid experimental evidence, e.g. mutagenesis analysis or structural analysis, in this or previous articles, that demonstrates this to be the case.
H269, as a crucial catalytic residue, was suggested by monitoring the effect of chemical modifications of amino acids on acetylation of HGSNAT membranes by Bame, K. J. and Rome, L. H. (1986). We agree that mutagenesis, catalysis, and structural evidence for the same are not currently available. We are pursuing a more thorough exploration of the role of both H269 (previous studies) and N258 (from this study) on the stability and function of HGSNAT.
In the discussion part, the authors mention previous studies in which it was postulated that the catalytic reaction can be described by a random order mechanistic model or a Ping Pong Bi Bi model. However, the authors leave open the question of which of these mechanisms best describes the acetylation reaction. The structure presented here does not provide evidence that could support one mechanism or the other.
We agree with the reviewer’s observation that the structure doesn’t indeed support one reaction mechanism or another. We are pursuing the structural and kinetic characterization of HGSNAT in the presence of other co-substrates and multiple pHs that are required to address this concern thoroughly.
Although the authors map the mutations leading to MPS IIIC on the structure and use FoldX software to predict the impact of these mutations on folding and fold stability, there is no experimental evidence to support FoldX's predictions.
We are working on assessing the impact of specific mutations on the stability of HGSNAT and will add them to the revised version of the manuscript. We thank the reviewer for this suggestion.
Reviewer #3 (Public Review):
Summary:
Navratna et al. have solved the first structure of a transmembrane N-acetyltransferase (TNAT), resolving the architecture of human heparan-alpha-glucosaminide N-acetyltransferase (HGSNAT) in the acetyl-CoA bound state using single particle cryo-electron microscopy (cryoEM). They show that the protein is a dimer and define the architecture of the alpha- and beta- GSNAT fragments, as well as convincingly characterizing the binding site of acetyl-CoA.
Strengths:
This is the first structure of any member of the transmembrane acyl transferase superfamily, and as such it provides important insights into the architecture and acetyl-CoA binding site of this class of enzymes.
The structural data is of a high quality, with an isotropic cryoEM density map at 3.3Å facilitating the building of a high-confidence atomic model. Importantly, the density of the acetyl-CoA ligand is particularly well-defined, as are the contacting residues within the transmembrane domain.
The open-to-lumen structure of HSGNAT presented here will undoubtedly lay the groundwork for future structural and functional characterization of the reaction cycle of this class of enzymes.
We thank the reviewer for their positive assessment of the data presented in this work. We really appreciate and agree with the reviewer's comment that the “structure of HSGNAT presented here will undoubtedly lay the groundwork for future structural and functional studies.”
Weaknesses:
While the structural data for the open-to-lumen state presented in this work is very convincing, and clearly defines the binding site of acetyl-CoA, to get a complete picture of the enzymatic mechanism of this family, additional structures of other states will be required.
We agree with the reviewers’ assessment and are heavily invested in pursuing the structures of all the steps of acetyl transfer by HGSNAT.
A potentially significant weakness of the study is the lack of functional validation. The enzymatic activity of the enzyme characterized was not measured, and the enzyme lacks native proteolytic processing, so it is a little unclear whether the structure represents an active enzyme.
We thank the reviewer for this comment. While the proteolytic cleavage of the protein remains debated, we find no evidence of such an event in our purification (SDS-PAGE and SEC). Studies like Durand et al., (2010) and Fan et al., (2011) suggest that even the ER retained monomeric HGSNAT is active. Because we see acetyl-CoA (co-substrate) bound to the protein in our structure, we surmise that proteolysis is not necessary for function, at least not for substrate binding. However, we are working towards the structural and kinetic characterization of recombinant α- and β-HGSNAT construct to explore the role of proteolysis on HGSNAT stability and function.
Reviewer #1 (Public Review):
This article by Navratna et al. reports the first structure of human HGSNAT in an acetyl-CoA-bound state. Through careful structural analysis, the authors propose potential reasons why certain human mutations lead to lysosomal storage disorders and outline a catalytic mechanism. The structural data are of good quality, and the manuscript is clearly written. This study represents an important step toward understanding the mechanism of HGSNAT and is valuable to the field. I have the following suggestions:
1. The authors should characterize whether the purified protein is active. Otherwise, how does one know if the detergent used maintains the protein in a biologically relevant state? The authors should at least attempt to do so. If these prove to be challenging, at the very least, the authors should try a cell-based assay to demonstrate that the GFP tag does not interfere with the function.
2. In Figure 5, the authors present a detailed schematic of the catalytic cycle, which I find to be too speculative. There is no evidence to suggest that this enzyme undergoes isomerization, similar to a transporter, between open-to-lumen and open-to-cytosol states. Could it not simply involve some movements of side chains to complete the acetyl transfer?
Zehn von den Climate Stripes abgeleitete Grafiken zu den meteorologischen Rekorden des Jahres 2023, darunter auch zu den Meerestemperaturen und der Oberfläche des arktischen bzw antarktischen Meereises. An 40 Tagen des Jahres 2023 war es im globalen Durchschnitt heißer als an jedem vorher gemessenen Tag. https://www.repubblica.it/green-and-blue/2024/01/03/news/climate_stripes_2023_anno_record_crisi_clima-421803988/
Don’t expect people to change their behavior just so you can measure it. For example, don’t expect that everybody will tag their bugs, PRs, etc. in some special way just so that you can count them.
100% true.
As always, the annotations you see will be yours, those posted in Public, and those posted in groups of which you are a member.
The real question is: how to exclude some results?
For example, I have few annotations under tag search results, but there are some of mine, and I want to see only those which are not mine, but others. How can I accomplish that?
It would be helpful, because it provides solution for excluding from results users or websites which are unworthy or spammed with worthless annotations.
Author Response
The following is the authors’ response to the original reviews.
Note to all Reviewers
We appreciate the reviewers’ comments and suggestions for improving the manuscript. Below is a summary of new data added and a brief description of the major new results. A detailed pointby-point response follows.
New data:
• Figure 1f
• Figure 2b, f, g
• Figure 4b
• Figure S7 • Figure S8
• Figure S9
Summary of major new results/edits:
• At the request of Reviewer #1 we have updated the name of the degradation tag to be more specific and we now call it the “LOVdeg” tag.
• We have added new controls demonstrating that light stimulation does not cause photobleaching or toxicity issues (Fig. S7).
• We now show that LOVdeg can function at various points in the growth cycle, demonstrating robust degradation (Fig. 1f, Fig. S8).
• We have included relevant controls for the AcrB-LOVdeg efflux pump results (Fig. 2f-g).
• We have included important benchmarking controls, such as an EL222-only control and SsrA tag control to provide a clearer view of how LOVdeg performance compares to other systems (Fig. S9, Fig. 4b).
Additional note:
• While repeating experiments during the revision process we found that the results for the combined action of EL222 and the LOVdeg tag were not as dramatic as in our original measurements, though the overall findings are consistent with our original results. Specifically, we still find that the combination of EL222 and the LOVdeg tag produces a lower signal than either on their own. We have updated these data in the revised manuscript (Fig. 4b).
Reviewer #1:
Public Review:
Specifically controlling the level of proteins in bacteria is an important tool for many aspects of microbiology, from basic research to protein production. While there are several established methods for regulating transcription or translation of proteins with light, optogenetic protein degradation has so far not been established in bacteria. In this paper, the authors present a degradation sequence, which they name "LOVtag", based on iLID, a modified version of the blue-light-responsive LOV2 domain of Avena sativa phototropin I (AsLOV2). The authors reasoned that by removing the three C-terminal amino acids of iLID, the modified protein ends in "-E-A-A", similar to the "-L-A-A" C-terminus of the widely used SsrA degradation tag. The authors further speculated that, given the light-induced unfolding of the C-terminal domain of iLID and similar proteins, the "-E-A-A" C-terminus would become more accessible and, in turn, the protein would be more efficiently degraded in blue light than in the dark.
Indeed, several tested proteins tagged with the "LOVtag" show clearly lower cellular levels in blue light than in the dark. While the system works efficiently with mCherry (10-20x lower levels upon illumination), the effect is rather modest (2-3x lower levels) in most other cases. Accordingly, the authors propose to use their system in combination with other light-controlled expression systems and provide data validating this approach. Unfortunately, despite the claim that the "LOVtag" should work faster than optogenetic systems controlling transcription or translation of protein, the degradation kinetics are not consistently shown; in the one case where this is done, the response time and overall efficiency are similar or slightly worse than for EL222, an optogenetic expression system.
The manuscript and the figures are generally very well-composed and follow a clear structure. The schematics nicely explain the underlying principles. However, limitations of the method in its main proposed area of use, protein production, should be highlighted more clearly, e.g., (i) the need to attach a C-terminal tag of considerable size to the protein of interest, (ii) the limited efficiency (slightly less efficient and slower than EL222, a light-dependent transcriptional control mechanism), and (iii) the incompletely understood prerequisites for its application. In addition, several important controls and measurements of the characteristics of the systems, such as the degradation kinetics, would need to be shown to allow a comparison of the system with established approaches. The current version also contains several minor mistakes in the figures.
We thank reviewer #1 for the feedback and suggestions to strengthen the manuscript. We have addressed these comments in the points that follow and now include important controls and benchmarks for our molecular tool.
Major points
- The quite generic name "LOVtag" may be misleading, as there are many LOV-based tags for different purposes.
We appreciate that it would be beneficial to have a more specific name. We have updated the name to “LOVdeg” tag, which captures both the inclusion of LOV and the degradation function of the tag.
Updated throughout the manuscript and figures
- Throughout the manuscript, the authors use "expression levels". As protein degradation is a post-expression mechanism, "protein levels" should be used instead.
We have transitioned to using “protein levels” at many points in the manuscript.
Updated throughout the manuscript
- Degradation dynamics (time course experiments) should be shown. The only time this is done in the current version (in Fig. 4), degradation appears to be in the same range (even a bit slower) than for EL222, which does not support the claim that the "LOVtag" acts faster than other optogenetic systems controlling protein levels.
In the revised manuscript, time course data are now shown at multiple points. These include new data in Fig. 1f and Fig. S8 that demonstrate degradation at various stages of growth. Fig. S4 also shows the dynamics of degradation when comparing to the addition of exogenously expressed ClpA. We have added text in the results section to point the reader to these data. In addition, we have made minor modifications to the text in the Introduction to avoid making claims about speed comparisons. Fig. 1f, Fig. S8, Fig. S4
Results: Design and characterization of the AsLOV2-based degradation tag, Introduction
- "Frequency" is used incorrectly for Fig. 3. A series of 5 seconds on, 5 seconds off corresponds to a frequency of 0.1 Hz (1 illumination round / 10 s), not of 0.5 Hz. What the authors indicate as "frequency" is the fraction of illumination time. However, the (correct) frequency should be given, as this is likely the more important factor.
We have changed how we calculate frequency to use the proposed definition of one pulse per time period. We updated the values in the text and in the figure. Fig. 3c
Results: Tuning frequency response of the LOVdeg tag
- To properly evaluate the system, several additional controls are needed:
a. To test for photobleaching of mCherry by blue light illumination, untagged controls should be shown for the mCherry-based experiments. Fluorescence always seems to be lower upon illumination, except for the AsLOV2*(546) data, where it cannot be excluded that fluorescence readings are saturated. Relatedly, the raw data for OD and fluorescence should be included. Showing a Western blot against mCherry in at least one case would allow to separate the effects of photobleaching and degradation.
We appreciate the suggestion and have conducted these important controls. We now include new data demonstrating that light induction does not change fluorescence levels using an untagged mCherry control, nor does it significantly affect endpoint OD levels. Based on these results, we did not perform a Western blot because there were no effects to separate. Fig. S7
b. In Fig. 2b, light + IPTG should be shown to estimate the activity of the system at higher expression levels.
We have added these to the figure. Light + IPTG modestly increases expression compared to IPTG only, likely due to the saturating level of IPTG added, which achieves near full induction. Fig. 2b
c. In Fig. 4, EL222 alone should be shown to allow a comparison with the LOVtag. From the data presented, it looks like EL222 is both slightly faster and more efficient than the LOVtag.
We have added the EL222-only case for comparison with LOVdeg only and EL222 + LOVdeg. We note that Reviewer #3 raised a similar concern. Fig. 4b
d. The effect of the used light on bacterial viability under exponential and stationary conditions should be shown.
In this revision, we have added new data on light exposure at various points during exponential and stationary phase (Fig. 1f, Fig. S8). These OD data show that growth curves are similar for all cultures, regardless of the time light is applied during the growth phase. Additionally, we also now include ODs for the photobleaching experiments. These data also show that growth is not significantly altered under continuous light exposure. Figure 1f, Fig. S7b
- The claim that "Post-translational control of protein function typically requires extensive protein engineering for each use case" is not correct. The authors should discuss alternative options, e.g. based on dimerization, more extensively and in a less biased manner.
We have toned down the language in this location and at other points in the manuscript. However, we maintain that other types of post-translational control, such as dimerization or LOV2 domain insertion, require more protein engineering than inserting a degradation tag. For example, we and others have directly demonstrated this in previous work (e.g. DOI: 10.1021/acssynbio.9b00395, 10.1101/2023.05.26.542511, 10.1038/s41467-023-38993-6), where numerous split site or insertion variants need to be screened and fine-tuned for successful light control. In contrast, a degradation mechanism has the potential to require less fine tuning to achieve a light response. We have included the above sources to clarify this point. Introduction, Results: Modularity of the LOVdeg tag
Minor points
- In Suppl. Fig. 1, amino acid numbers seem to be off. Also, the alterations in iLID (compared to AsLOV2) that are not used in "LOVtag" appear to be missing and the iLID sequence incorrect, as a consequence.
Thank you for catching this. The number indices in Fig. S1 have been corrected. We also realized we were reporting the iLID(C530M) variant in our amino acid sequence and have reverted the 530M back to C. Fig. S1
- Why is AsLOV2(543) more efficiently degraded than AsLOV2(543) (blue column in Fig. 1d) when the dark state should be stabilized in AsLOV2(543)?
We are not sure of the exact reason for the increased degradation response in the AsLOV2*(543) variant. It may be that the dark-state stabilizing mutations introduced also have more favorable interactions with degradation machinery, although this is highly speculative.
- Why does the addition of EL222 reduce protein levels so strongly in the dark for CpFatB1* (Fig. 5)?
We believe this effect stems from the EL222 responsive promoter (PEL222). With LOVdeg only, CpFatB1* is expressed from an IPTG inducible promoter (PlacUV5) whereas EL222 responsive constructs necessitate a promoter switch containing an EL222 binding site. We have clarified this point and expanded our discussion of these results.
Results: Optogenetic control of octanoic acid production
- Fig. 2f / S10 are difficult to interpret. Why does illumination only lead to a significant effect at 2.5 and 5 µg/ml and not at lower concentrations, where the degradation system would be expected to be most efficient?
We have expanded our discussion on these results to explain that this likely stems from basal protein levels of AcrB-LOVdeg in the light that can provide resistance at low antibiotic concentrations. We have also added new controls to this figure to show the chloramphenicol sensitivity of a ΔacrB strain and a ΔacrB strain with an IPTG-inducible version of acrB with no induction, demonstrating the lowest achievable chloramphenicol resistance from a standard inducible system.
Results: Modularity of the LOVdeg tag, Fig. 2f-g
- Fig. 2f / S10 do not measure the MIC (which is a clearly defined value), but the sensitivity to Chloramphenicol.
We have changed the text to use the term chloramphenicol sensitivity instead of MIC. Results: Modularity of the LOVdeg tag
- "***" in Fig. S1 should be explained.
We have removed the ‘***’ to avoid confusion. Fig. S1
- The fold-change differences between light and dark, indicated in some selected cases, should be listed for all figures.
We have added fold-change values where appropriate. Fig 1d, Fig. 2b
Reviewer #2:
Public Review:
In this manuscript the authors present and characterize LOVtag, a modified version of the bluelight sensitive AsLOV2 protein, which functions as a light-inducible degron in Escherichia coli. Light has been shown to be a powerful inducer in biological systems as it is often orthogonal and can be controlled in both space and time. Many optogenetic systems target regulation of transcription, however in this manuscript the authors target protein degradation to control protein levels in bacteria. This is an important advance in bacteria, as inducible protein degradation systems in bacteria have lagged behind eukaryotic systems due to protein targeting in bacteria being primarily dependent on primary amino acid sequence and thus more difficult to engineer. In this manuscript, the authors exploit the fact that the J-alpha helix of AsLOV2, which unwinds into a disordered domain in response to blue light, contains an E-A-A amino acid sequence which is very similar to the C-terminal L-A-A sequence in the SsrA tag which is targeted by the unfoldases ClpA and ClpX. They truncate AsLOV2 to create AsLOV2(543) and combine this truncation with a mutation that stabilizes the dark state to generate AsLOV2*(543) which, when fused to the C-terminus of mCherry, confers light-induced degradation. The authors do not verify the mechanism of degradation due to LOVtag, but evidence from deletion mutants contained in the supplemental material hints that there is a ClpA dominated mechanism. They demonstrate modularity of this LOVtag by using it to degrade the LacI repressor, CRISPRa activation through degradation of MCP-SoxS, and the AcrB protein which is part of the AcrAB-TolC multidrug efflux pump. In all cases, measurement of the effect of the LOVtag is indirect as the authors measure reduction in LacI repression, reduction in CRISPRa activation, and drug resistance rather than directly measuring protein levels. Nevertheless the evidence is convincing, although seemingly less effective than in the case of mCherry degradation, although it is hard to compare due to the different endpoints being measured. The authors further modify LOVtag to contain a known photocycle mutation that slows its reversion time in the dark, so that LOVtag is more sensitive to short pulses of light which could be useful in low light conditions or for very light sensitive organisms. They also demonstrate that combining LOVtag with a blue-light transcriptional repression system (EL222) can decrease protein levels an additional 269-fold (relative to 15-fold with LOVtag alone). Finally, the authors apply LOVtag to a metabolic engineering task, namely reducing expression of octanoic acid by regulating the enzyme CpFatB1, an acyl-ACP thioesterase. The authors show that tagging CpFatB1 with LOVtag allows light induced reduction in octanoic acid titer over a 24 hour fermentation. In particular, by comparing control of CpFatB1 with EL222 transcriptional repression alone, LOVtag, or both the authors show that light-induced protein degradation is more effective than light-induced transcriptional repression. The authors suggest that this is because transcriptional repression is not effective when cells are at stationary phase (and thus there is no protein dilution due to cell division), however it is not clear from the available data that the cells were in stationary phase during light exposure. Overall, the authors have generated a modular, light-activated degron tag for use in Escherichia coli that is likely to be a useful tool in the synthetic biology and metabolic engineering toolkit.
We thank Reviewer #2 for the constructive feedback. In the updated manuscript, we now include data demonstrating degradation at different growth stages and address other points brought up in the review to improve understanding of the degradation tag.
Overall, the authors present a well written manuscript that characterizes an interesting and likely very useful tool for bacterial synthetic biology and metabolic engineering. I have a few suggestions that could improve the presentation of the material.
Major Comments:
• Could the authors clarify, perhaps through OD measurements, that the cultures in the octanoic acid experiment are actually in stationary phase during the relevant light induction. It isn't clear from the methods.
We have updated the Methods to clarify that the cells are entering stationary phase (OD600 = 0.6) when light is either kept on or turned off for production experiments. Production is continued for the following 24 hours. Note that we now show OD measurements in a separate set of experiments (Fig. 1f, Fig. S8).
Methods: Octanoic acid production experiment. Fig. 1f, Fig. S8
• Can the authors clarify why there is an overall decrease in protein in the clpX deletion? And is it this initial reduction that is the source of the change in fold in 1C? Similarly, for hslU is it because overall protein levels are higher with the tag? In general, I feel that the interpretation of Supplemental Figures S6-S10 could be moved in more detail to the main text, or at least the main takeaway points. But this is a personal preference, and not necessary to the major flow of the story which is about the utility of the LOVtag tool.
As shown in Fig. S5, expression of mCherry without any degradation tag is decreased in a clpX knockout strain compared to wild type. This difference may be the result of reduced cell health, and we now note this in the text. The strains shown in Fig. 1c are in wild type cells with normal expression, so this is not the source of the fold change. As for hslU, we agree it is interesting that expression seems to increase. However, the increase is modest and could stem from gene network regulation differences in that strain compared to wild type and may not be related to LOVdeg tag degradation. Each endogenous protease is involved in a wide range of functions within the cell, and it is unknown how global gene expression is impacted. We acknowledge the suggestion of moving the protease results to the main text, but we have ultimately elected to keep these data in the Supplementary Information to maintain the flow in the manuscript. However, we have added additional text pointing the reader to the Supplemental Text and include a brief summary of the findings in the main text.
Results: Design and characterization of the AsLOV2-based degradation tag
• What is the source of the poor repression in Figure 2D?
Presumably, this stems from low levels of the CRISPRa MCP-SoxS activator, even in the presence of light. We have added this point to the text.
Results: Modularity of the LOVdeg tag
• In general, it would be nice to have light-only controls for many of the experiments to validate that light is not affecting the indicated proteins or their function.
We thank the reviewer for this suggestion and note that Reviewer #1 raised a similar concern. We have now included light-only data for a strain containing IPTG-inducible mCherry without the LOVdeg tag (Fig. S7). These data show that light itself, at the levels used in this study, does not affect mCherry expression or cell growth. This strain serves as a direct control for data presented in Fig. 1 and Fig. 2b, as the systems are identical except for the addition of the LOVdeg tag onto either mCherry or the LacI repressor. Additionally, the control translates to other experiments since mCherry is used as a reporter for other systems in this study. Fig. S7
• It would be nice to directly measure the function of the tool at different phases of E. coli growth to show directly that protein degradation works at stationary phase, rather than the more indirect measurements used in the octanoic acid experiment.
We thank the reviewer for this suggestion, which significantly strengthens our results. We have added an experiment that tests the LOVdeg tag at different phases of growth (Fig. 1f, Fig. S8). In this experiment, cultures are growth from early exponential to stationary phase, and light is introduced at various points. Exposure windows of 4 hours, ranging from early exponential to stationary phase, all show functional light inducible degradation. Fig. 1f, Fig. S8.
Results: Design and characterization of the AsLOV2-based degradation tag
Minor Comments:
• It would be nice to make clear that the data in S6d and S7 is repeated, but with the HslUV data in S7.
We clarified this point in the caption of Fig. S4 (the former Fig. S7 in the original manuscript). Fig. S4 caption
• Why was 5s picked for the frequency response in Figure 3
We picked 5s because 1) it is a substantially shorter timescale than overall degradation dynamics seen for the LOVdeg tag, and 2) we found that shorter pulses could not be reliably achieved with the light stimulation hardware and software we used (Light Plate Apparatus with Iris software). To ensure high fidelity pulses, we opted for 5 second pulses that we empirically determined to be stable throughout long experiments. We have added text clarifying this. Results: Tuning frequency response of the LOVdeg tag
Reviewer #3:
Public Review:
The authors present the mechanism, validation, and modular application of LOVtag, a light-responsive protein degradation tag that is processed by the native degradosome of Escherichia coli. Upon exposure to blue light, the c-terminal alpha helix unfolds, essentially marking the protein for degradation. The authors demonstrate the engineered tag is modular across multiple complex regulatory systems, which shows its potential widespread use throughout the synthetic biology field. The step-by-step rational design of identifying the protein that was most dark stabilized as well as most light-responsive for degradation, was useful in terms of understanding the key components of this system. The most compelling data shows that the engineered LOVTag can be fused to multiple proteins and achieve light-based degradation, without affecting the original function of the fused protein; however, results are not benchmarked against similar degradation tagging and optogenetic control constructs. Creating fusion proteins that do not alter either of the original functions, is often difficult to achieve, and the novelty of this should be expanded upon to drive further impact.
We appreciate the feedback from Reviewer #3 to improve the manuscript. We have included important controls and benchmarking experiments to address the reviewer’s concerns, which are detailed in the points below.
Benchmarking:
The similarity between the L-A-A sequence of SsrA and the E-A-A sequence of LOVtag is one of the pieces of evidence that led the authors to their current protein design. The differences in degradation efficiency between the SsrA degradation tag and LOVtag are not shown, and benchmarking against SsrA would be a valuable way to demonstrate the utility of this construct relative to an established protein tagging tool.
We thank the reviewer for suggesting an experiment to benchmark performance. We have added new experimental data where a full length SsrA tag is added to a fusion protein of nearly identical size (mCherry-iLID), allowing us to directly compare performance to mCherryLOVdeg (Fig. S9). These results show that light inducible control with LOVdeg tag decreases protein expression levels to near those achieved with the native SsrA tag. Fig. S9.
Results: Design and characterization of the AsLOV2-based degradation tag
Additionally, there is a lack of an EL222-only control presented in Figure 4b and in the results section beginning with "Integrating the LOVtag with EL222...". Without benchmarking against this control the claim that "EL222 and the LOVtag work coherently to decrease expression" is unsubstantiated. No assumptions of synergy can be made.
We appreciate this comment and note that Reviewer #1 raised a similar concern. We have added data to Fig. 4b with an EL222-only control for comparison. Fig. 4b
The dramatic change in dark octanoic acid titer between the EL222, LOVtag and combined conditions are surprising, especially in comparison to the lack of change in the dark mCherry expression shown in Figure 4b. This data is the only to suggest that LOVtag may perform better than EL222. However, the inconsistencies in dark state regulation presented in the two experiments, and between conditions in this experiment bring the latter claim to question. A recommendation is that the authors either repeat this experiment, or comment on the observed discrepancy in dark state octanoic acid titers in their discussion.
First, a key difference between the data presented in Fig. 4 and Fig. 5 is that the production experiment is conducted over a long time period (24 hours) and the EL222/LOVdeg reporter experiment is conducted over 5 hours. Likely, performance differences between EL222 and the LOVdeg tag become more pronounced as protein accumulation occurs. Second, the LOVdeg only construct is expressed from a non-EL222 promoter which is able to achieve higher expression (see response to Reviewer #1, Minor point #3). Lastly, a convoluting factor is that the relationship between expression of CpFatB1 and octanoic acid production is not completely linear, and there are likely thresholds or expressions windows that result in similar endpoint titers. We agree a more detailed examination of how CpFatB1 changes over the course of the production period would be very interesting. However, this is beyond the scope of the present study, whose goal is to introduce and showcase the utility of the LOVdeg tag as a tool. We have added new discussion on this in the Results section to clarify some of these points. We have also repeated all experiments in Fig. 4 and consistently see the LOVdeg tag performing as well as or better than EL222. As noted in the remarks to all reviewers, these data have been updated in the revised manuscript.
Results: Optogenetic control of octanoic acid production. Fig. 4d
Based on the methodology presented, no change in the duration in light exposure was tested, even though this may be an important part of the system response. The on/off, for example in Figure 4b, is either all light or all dark, but they claim that their system is beneficial especially at stationary phase. The authors should consider showing the effects of shifting from dark to light at set intervals. (i.e. 1 hr dark then light, 2hr dark until light, etc.) This data would also aid in supporting the utility of this tag for controlling expression during different growth phases, where light may be used after the cells have reached a certain phase.
We have added new data showing the effect of light stimulation at different times in the growth cycle (see response to Reviewer #2, bullet point #5). These data demonstrate that the LOVdeg tag performs well at various points in the growth cycle. Fig. 1f, Fig. S8.
Results: Design and characterization of the AsLOV2-based degradation tag
Minor Revisions Figures:
Figure 1:
More clarity is needed in the naming conventions for this figure and in the body of the text. For example, a different convention than 546 and 543 should be used to refer to the full and truncated lengths of the tag. It would greatly aid understanding for this to be made more clear. The authors could simply continue to use "full" and "truncated" to refer to them. In addition, the term "stabilizing mutations" in 1c could be changed to read "dark state stabilizing mutations" to aid in clarity.
When describing the design of the LOVdeg tag, we opted towards a more technically accurate description over clarity in order to make our engineering process easily comparable to other LOV2 systems. As such, we kept the number-based nomenclature (543 or 546) to represent the domain within the phototropin 1 protein from Avena sativa (AsLOV2). The domain used in this study, and many other studies, are only amino acids 404-546, i.e. not the full sequence, thus saying simply ‘full’ or ‘truncated’ is not technically accurate. We believe the detailed nomenclature, which is limited to one section, is important to provide clarity on exactly what we used for protein engineering. In the revised version we introduce the nickname “LOVdeg” tag earlier and use it throughout the rest of the manuscript.
Results: Design and characterization of the AsLOV2-based degradation tag
- 1b It is not clear that this is the dark state stabilized structure in the figure, but is referred to as such only in the body of the text.
We have added text in the manuscript to clarify this is AsLOV2, not iLID, and have labeled it in the figure caption as well.
Results: Design and characterization of the AsLOV2-based degradation tag
- 1d. Fold change is reported in Figure 2d, and may be relevant to include those values in 1d as well.
Done. Fig. 1d
- 1e. It is not clear which tag is being used in this bar plot. Please specify that this is the dark state stabilized, truncated tag.
We have added a title to the plot and language to the caption, both of which clarify this point. Fig. 1e
- In addition, the microscopy images provided in supplemental material should be included in the first figure as it adds a compelling observation of LOVtag activity.
We are pleased to hear that the microscopy results are beneficial, however we elected to leave them in Supplementary to preserve the flow of the manuscript in the text surrounding Fig. 1.
Figure 2:
2d. It is unclear what the 2.5x fold change is relative to (the baseline or the dark)
We have added a line in the figure to clarify the comparison being made. Fig. 2d
- 2f. More discussion can be added to describe what concentration of chloramphenicol is biologically/bioreactor relevant.
Our previous studies on the relationship between AcrAB expression and mutation rate (cited in the text) were carried out at a concentration within the range in which the LOVdeg tag is effective (5 μg/ml), suggesting this range to be relevant to tolerance and resistance.
Figure 3:
We recommend that this data and discussion are better suited for supplementary figures. The results shown here essentially recapitulate the same findings of Zoltowski et al., 2009. In addition, the paper describing this mutation should be cited in this figure caption in addition to the body of the text
Although these results are in line with previous findings, we believe this dataset is important for several reasons. First, the agreement with known mutations validates the unfolding-based mechanism for degradation control. Second, degradation that is contingent on unfolding of LOV2 offers a direct actuating mechanism of photocycle properties. Other systems, like that in Zoltowski et al., examine properties of purified proteins but lack the mechanism to translate its effect in live cells. This figure demonstrates how degradation can do so and lays the groundwork for degradation-based frequency processing circuits. Last, there are discrepancies between photocycle kinetics in situ, as reported by Li et al. (DOI: 10.1038/s41467-020-18816-8), and in cell-free studies such as in Zoltowski et al. The studies use different methods of measuring photocycle kinetics (in situ vs cell-free). This dataset substantiates relaxation times from Li et al. and suggests cell-free relaxation time constants are over estimated relative to our live cell results.
Figure 4:
There is a lack of an EL222-only control presented in Figure 4b. Without this data present, the claim that "EL222 and the LOVtag work coherently to decrease expression" is unsubstantiated. No assumptions of synergy can be made.
We have added EL222-only data to the figure; we note that Reviewer #1 made a similar request. Figure 4b
Manuscript
Results
Design and characterization...
Due to the extensive discussion of ClpX at the beginning of this section, more of the results on evaluating the binding partners and mechanism of LOVtag degradation should be presented in the main body of the manuscript and not in supplementary materials.
To maintain flow of the manuscript and focus on how the LOVdeg tag works as a synthetic biology tool, we have opted to keep this section in the Supplement Information, but have several lines in the text related to Fig. 1 that point the reader to this material. Results: Design and characterization of the AsLOV2-based degradation tag
- In the second paragraph of this section, the authors theorize that the C-terminal truncated E-AA sequence will "remain caged as part of the folded helix". How did the authors determine this? Was there any evidence to suggest that the truncated state would be any more responsive than the full length sequence? More data or rationale may need to be introduced to support the overall hypothesis presented in this paragraph.
We determined this by examining the crystal structure which shows that the E-A-A sequence is part of the folded helix. As seen in Fig. 1b, addition of amino acids after the EAAKEL sequence would not be part of the folded helix which ends prior to the terminal leucine. We added text to clarify our logic.
Results: Design and characterization of the AsLOV2-based degradation tag
- The similarity between the L-A-A sequence of SsrA and the E-A-A sequence of LOVtag is one of the pieces of evidence that brought the authors to their current protein design. The differences in degradation efficiency between the SsrA degradation tag and LOVtag are not clear, and benchmarking against SsrA would be a valuable way to demonstrate the utility of this construct relative to an established protein tagging tool.
We added an SsrA comparison to benchmark the system. Fig. S9
Results: Design and characterization of the AsLOV2-based degradation tag
Tuning frequency and response...
Overall the results presented in this section essentially recapitulate the effects that mutation presented in Zoltowski et. al., 2009 have on AsLOV2 dark state recovery and although this is a useful observation of LOVtag performance, a recommendation is to move this into a supplementary section.
See above response to Fig. 3 comment.
Integrating the LOVtag with EL222...
The claim is made in this section that LOVtag and EL222 work synergistically, however the experiments presented do not test repression due to EL222 activity alone. Without benchmarking against this control, the claim of synergy is not supported and we recommend that the authors perform this experiment again with the EL222-only control.
We have added this important control. Fig. 4b
Discussion
- The statement "the LOVtag can easily be integrated with existing optogenetic systems to enhance their function" is not substantiated without benchmarking LOVtag against an EL222- only control. As mentioned above this condition should be included in the experiments discussed in Figure 4 and in the section "Integrating the LOVtag with EL222.."
We added EL222-only regulation to benchmark the LOVdeg tag and LOVdeg + EL222 experiments. Fig. 4b
Experiments
Applications:
The application of this tag to the metabolic control of octanoic acid production could be more impactful. For instance, using the LOVtag with two different enzymes to change the composition of long/short chain fatty acids with light induction., Or possibly integrating the tag into a switch to activate production. However, the authors address that "decreasing titers is not the overall goal in metabolic engineering" in their discussion, and therefore the pursuit of this additional experiment is up to the authors' discretion.
We appreciate the suggestions for further applications of the LOVdeg tag. We envision that follow up studies will focus on the application of the LOVdeg tag in metabolic engineering. However, this will require significant development of production systems. We believe this to be out of the scope of this work, where the goal is to present the design and function of the LOVdeg tag as a tool.
eLife assessment
This valuable study reports on a new tool that allows for light-controlled protein degradation in Escherichia coli. With the improved light-responsive protein tag, endogenous protein levels can be reduced severalfold. The methodology is convincing and will be of interest to the fields of gene expression regulation in bacteria and, more generally to synthetic biologists.
Reviewer #1 (Public Review):
Specifically controlling the level of proteins in bacteria is an important tool for many aspects of microbiology, from basic research to protein production. While there are several established methods for regulating transcription or translation of proteins with light, optogenetic protein degradation has so far not been established in bacteria. In this paper, the authors present a degradation sequence, which they name "LOVdeg", based on iLID, a modified version of the blue-light-responsive LOV2 domain of Avena sativa phototropin I (AsLOV2). The authors reasoned that by removing the three C-terminal amino acids of iLID, the modified protein ends in "-E-A-A", similar to the "-L-A-A" C-terminus of the widely used SsrA degradation tag. The authors further speculated that, given the light-induced unfolding of the C-terminal domain of iLID and similar proteins, the "-E-A-A" C-terminus would become more accessible and, in turn, the protein would be more efficiently degraded in blue light than in the dark.
Indeed, several tested LOVdeg-tagged proteins show clearly lower cellular levels in blue light than in the dark. Depending on the nature and expression level of the target protein, protein levels are reduced modestly to strongly (2 to 20x lower levels upon illumination). Accordingly, the authors propose to use their system in combination with other light-controlled expression systems and provide data validating this approach. The LOVdeg system allows to modulate protein levels to a similar degree and with comparable kinetics as optogenetic systems controlling transcription or translation of protein, and can be combined with such systems.
The manuscript and the figures are generally very well-composed and follow a clear structure. The schematics nicely explain the underlying principles. Besides the advantages of the LOVdeg approach, including its complementarity to controlled expression of proteins, the revised version of the manuscript also highlights the limitations of the method more clearly, e.g., (i) the need to attach a C-terminal tag of considerable size to the protein of interest, (ii) the limited efficiency (slightly less efficient and slower than EL222, a light-dependent transcriptional control mechanism), and (iii) the incompletely understood prerequisites for its application. Taken together, this manuscripts describes the LOVdeg system as a valuable addition to the tool box for controlling protein levels in prokaryotic cells.
Reviewer #2 (Public Review):
In this manuscript the authors present and characterize LOVdeg, a modified version of the blue-light sensitive AsLOV2 protein, which functions as a light-inducible degron in Escherichia coli. Light has been shown to be a powerful inducer in biological systems as it is often orthogonal and can be controlled in both space and time. Many optogenetic systems target regulation of transcription, however in this manuscript the authors target protein degradation to control protein levels in bacteria. This is an important advance in bacteria, as inducible protein degradation systems in bacteria have lagged behind eukaryotic systems due to protein targeting in bacteria being primarily dependent on primary amino acid sequence and thus more difficult to engineer. In this manuscript, the authors exploit the fact that the J-alpha helix of AsLOV2, which unwinds into a disordered domain in response to blue light, contains an E-A-A amino acid sequence which is very similar to the C-terminal L-A-A sequence in the SsrA tag which is targeted by the unfoldases ClpA and ClpX. They truncate AsLOV2 to create AsLOV2(543) and combine this truncation with a mutation that stabilizes the dark state to generate AsLOV2*(543) which, when fused to the C-terminus of mCherry, confers light-induced degradation. The authors do not verify the mechanism of degradation due to LOVdeg, but evidence from deletion mutants contained in the supplemental material hints that there is a ClpA dominated mechanism. The LOVdeg is able to target mCherry for protein degradation across different phases of bacterial growth, which is important for regulating processes at stationary phase and a potential additional advantage over transcriptional repression systems. They demonstrate modularity of this LOVdeg by using it to degrade the LacI repressor, CRISPRa activation through degradation of MCP-SoxS, and the AcrB protein which is part of the AcrAB-TolC multidrug efflux pump. In all cases, measurement of the effect of the LOVdeg is indirect as the authors measure reduction in LacI repression, reduction in CRISPRa activation, and drug resistance rather than directly measuring protein levels. Nevertheless the evidence is convincing, although seemingly less effective than in the case of mCherry degradation, although it is hard to compare due to the different endpoints being measured. The authors further modify LOVdeg to contain a known photocycle mutation that slows its reversion time in the dark, so that LOVdeg is more sensitive to short pulses of light which could be useful in low light conditions or for very light sensitive organisms. They also demonstrate that combining LOVdeg with a blue-light transcriptional repression system (EL222) can decrease protein levels an additional 23-fold (relative to 7-fold with LOVdeg alone). Finally, the authors apply LOVdeg to a metabolic engineering task, namely reducing expression of octanoic acid by regulating the enzyme CpFatB1, an acyl-ACP thioesterase. The authors show that tagging CpFatB1 with LOVdeg allows light induced reduction in octanoic acid titer over a 24 hour fermentation. In particular, by comparing control of CpFatB1 with EL222 transcriptional repression alone, LOVdeg, or both the authors show that light-induced protein degradation is more effective than light-induced transcriptional repression. The authors suggest that this is because transcriptional repression is not effective when cells are at stationary phase (and thus there is no protein dilution due to cell division). Overall, the authors have generated a modular, light-activated degron tag for use in Escherichia coli that is likely to be a useful tool in the synthetic biology and metabolic engineering toolkit.
Reviewer #3 (Public Review):
The authors present the mechanism, validation, and modular application of LOVtag, a light-responsive protein degradation tag that is processed by the native degradosome of Escherichia coli. Upon exposure to blue light, the c-terminal alpha helix unfolds, essentially marking the protein for degradation. The authors demonstrate the engineered tag is modular across multiple complex regulatory systems, which shows its potential widespread use throughout the synthetic biology field. The step-by-step rational design of identifying the protein that was most dark-stabilized as well as most light-responsive for degradation, was useful in terms of understanding the key components of this system. The most compelling data shows that the engineered LOVTag can be fused to multiple proteins and achieve light-based degradation, without affecting the original function of the fused protein.
Letterboxd has called itself “Goodreads for movies” but it has far surpassed that initial tag line, having figured out how to create a smooth and intuitive user experience, provide a pleasant and inviting community and earn revenue from both optional paid memberships and advertisers, including studios that produce the films being discussed.
Says Maris Kreizman in NYT on 24 Dec 2023.
4. Cite Card Icon : Hat (something above you)Tag : 5th block Quotation, cooking recipe from book, web, tv, anything about someone else’s idea is classified into this class. Important here is distinguishing “your idea (Discovery Card)” and “someone else’s idea (Cite Card)”. Source of the information must be included in the Cite Card. A book, for example, author, year, page(s) are recorded for later use.
https://www.flickr.com/photos/hawkexpress/189972899/in/album-72157594200490122/
Despite being used primarily as a productivity tool the PoIC system also included some features of personal knowledge management with "discovery cards" and "citation cards". Discovery cards were things which contained one's own ideas while the citation cards were the ideas of others and included bibliographic information. Citation cards were tagged on the 5th block as an indicator within the system.
Question: How was the information material managed? Was it separate from the date-based system? On first blush it would appear not, nor was there a subject index which would have made it more difficult for one to find data within the system.
0. PoIC Format Move your mouse over the picture. This is the basic of PoIC Fromat. It is consisted from Tag, Icon, Title, Date and Time Stamp, and Contents. After several trial, you will remember this format easily. It's virtual template. You can start PoIC with blank card, anytime, anywhere. In this universe, there are only four class of information : Record, Discovery, GTD, and Cite.
Introduction to the Pile of Index Cards method.
Discovery Card Icon : Electric Bulb (lightning)Tag : 3rd block Things from my brain, mind, spirit, anything emerge from inside me, are classified into this class. This is the most important and enjoyable cards among the Four Cards. You will see your discoveries emerges in your mind like a water from spring. In fact, the 80% of index cards in my dock is dominated by this Discovery Card.
These are more similar to zettelkasten and commonplacing traditions. They comprise the majority of the system.
Reviewer #1 (Public Review):
Tiedje et al. investigated the transient impact of indoor residual spraying (IRS) followed by seasonal malaria chemoprevention (SMC) on the plasmodium falciparum parasite population in a high transmission setting. The parasite population was characterized by sequencing the highly variable DBL$\alpha$ tag as a proxy for var genes, a method known as varcoding. Varcoding presents a unique opportunity due to the extraordinary diversity observed as well as the extremely low overlap of repertoires between parasite strains. The authors also present a new Bayesian approach to estimating individual multiplicity of infection (MOI) from the measured DBL$\alpha$ repertoire, addressing some of the potential shortcomings of the approach that have been previously discussed. The authors also present a new epidemiological endpoint, the so-called "census population size", to evaluate the impact of interventions.
This study provides a nice example of how varcoding technology can be leveraged, as well as the importance of using diverse genetic markers for characterizing populations, especially in the context of high transmission. The data are robust and clearly show the transient impact of IRS in a high transmission setting, however, some aspects of the analysis are confusing.
1) Approaching MOI estimation with a Bayesian framework is a well-received addition to the varcoding methodology that helps to address the uncertainty associated with not knowing the true repertoire size. It's unfortunate that while the authors clearly explored the ability to estimate the population MOI distribution, they opted to use only MAP estimates. Embracing the Bayesian methodology fully would have been interesting, as the posterior distribution of population MOI could have been better explored.
2) The "census population size" endpoint has unclear utility. It is defined as the sum of MOI across measured samples, making it sensitive to the total number of samples collected and genotyped. This means that the values are not comparable outside of this study, and are only roughly comparable between strata in the context of prevalence where we understand that approximately the same number of samples were collected. In contrast, mean MOI would be insensitive to differences in sample size, why was this not explored? It's also unclear in what way this is a "census". While the sample size is certainly large, it is nowhere near a complete enumeration of the parasite population in question, as evidenced by the extremely low level of pairwise type sharing in the observed data.
3) The extraordinary diversity of DBL$\alpha$ presents challenges to analyzing the data. The authors explore the variability in repertoire richness and frequency over the course of the study, noting that richness rapidly declined following IRS and later rebounded, while the frequency of rare types increased, and then later declined back to baseline levels. The authors attribute this to fundamental changes in population structure. While there may have been some changes to the population, the observed differences in richness as well as frequency before and after IRS may also be compatible with simply sampling fewer cases, and thus fewer DBL$\alpha$ sequences. The shift back to frequency and richness that is similar to pre-IRS also coincides with a similar total number of samples collected. The authors explore this to some degree with their survival analysis, demonstrating that a substantial number of rare sequences did not persist between timepoints and that rarer sequences had a higher probability of dropping out. This might also be explained by the extreme stochasticity of the highly diverse DBL$\alpha$, especially for rare sequences that are observed only once, rather than any fundamental shifts in the population structure.
Author Response
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public Review):
Summary:
The manuscript by Heyndrickx et al describes protein crystal formation and function that bears similarity to Charcot-Leyden crystals made of galectin 10, found in humans under similar conditions. Therefore, the authors set out to investigate CLP crystal formation and their immunological effects in the lung. The authors reveal the crystal structure of both Ym1 and Ym2 and show that Ym1 crystals trigger innate immunity, activated dendritic cells in the lymph node, enhancing antigen uptake and migration to the lung, ultimately leading to induction of type 2 immunity.
Strengths:
We know a lot about expression levels of CLPs in various settings in the mouse but still know very little about the functions of these proteins, especially in light of their ability to form crystal structures. As such data presented in this paper is a major advance to the field.
Resolving the crystal structure of Ym2 and the comparison between native and recombinant CLP crystals is a strength of this manuscript that will be a very powerful tool for further evaluation and understanding of receptor, binding partner studies including the ability to aid mutant protein generation.
The ability to recombinantly generate CLP crystals and study their function in vivo and ex vivo has provided a robust dataset whereby CLPs can activate innate immune responses, aid activation and trafficking of antigen presenting cells from the lymph node to the lung and further enhances type 2 immunity. By demonstrating these effects the authors directly address the aims for the study. A key point of this study is the generation of a model in which crystal formation/function an important feature of human eosinophilic diseases, can be studied utilising mouse models. Excitingly, using crystal structures combined with understanding the biochemistry of these proteins will provide a potential avenue whereby inhibitors could be used to dissolve or prevent crystal formation in vivo.
The data presented flows logically and formulates a well constructed overall picture of exactly what CLP crystals could be doing in an inflammatory setting in vivo. This leaves open a clear and exciting future avenue (currently beyond the scope of this work) for determining whether targeting crystal formation in vivo could limit pathology.
Weaknesses:
Although resolving the crystal structure of Ym2 in particular is a strength of the authors work, the weaknesses are that further work or even discussion of Ym2 versus Ym1 has not been directly demonstrated. The authors suggest Ym2 crystals will likely function the same as Ym1, but there is insufficient discussion (or data) beyond sequence similarity as to why this is the case. If Ym1 and Ym2 crystals function the same way, from an evolutionary point, why do mice express two very similar proteins that are expressed under similar conditions that can both crystalise and as the authors suggest act in a similar way. Some discussion around these points would add further value.
We agree with reviewer. We have further elaborated the discussion section including these points, stating clearly that more research needs to be done using Ym2 crystals before we can draw parallels in vivo.
Additionally, the crystal structure for Ym1 has been previously resolved (Tsai et al 2004, PMID 15522777) and it is unclear whether the data from the authors represents an advance in the 3D structure from what is previously known.
The crystal structure of Ym1 has indeed been previously solved, and we refer to that paper. In addition, we also provide the crystal structure of in vitro grown Ym1, ashowing biosimilarity. This, for the field of crystallography is a major finding, since it validates the concept that crystal structures generated in vitro can reflect in vivo grown structures. Moreover, the in vivo crystallization of Ym2 was unknown prior to this work, and is now clear as revealed by the ex vivo X-ray crystallography. The strength of our story is that we can now compare Ym1 and Ym2 crystals structures in detail.
Whilst also generating a model to understand Charcot-Leyden crystals (CLCs), the authors fail to discuss whether crystal shape may be an important feature of crystal function. CLCs are typically needle like, and previous publications have shown using histology and TEM that Ym1 crystals are also needle like. However, the crystals presented in this paper show only formation of plate like structures. It is unclear whether these differences represent different methodologies (ie histology is 2D slides), or differences in CLP crystals that are intracellular versus extracellular. These findings highlight a key question over whether crystal shape could be important for function and has not been addressed by the authors.
In contrast to the bipyramidal, needle-like CLC crystals formed by human galectin-10 protein (hexagonal space group P6522), the in vivo grown Ym1 and Ym2 crystals we were able to isolate for X-ray diffraction experiments had a plate-like morphology with identical crystallographic parameters as recombinant Ym1/Ym2 crystals (space group P21). We note that depending on the viewing orientation of the thin plate-like Ym1 crystals, they may appear needle-like in histology and TEM images. In addition, we can fully not exclude that both Ym1 or Ym2 may crystallize in vivo in different space groups (which could result in different crystal morphologies for Ym1/Ym2) but we have no data to support this. It is finally also a possibility that plate like structures can break up in vivo along a long axis as a result of mechanical forces, and end up as rod-or needle like shapes.
Ym1/Ym2 crystals are often observed in conditions where strong eosinophilic inflammation is present. However, soluble Ym1 delivery in naïve mice shows crystal formation in the absence of a strong immune response. There is no clear discussion as to the conditions in which crystal formation occurs in vivo and how results presented in the paper in terms of priming or exacerbating an immune response align with what is known about situations where Ym1 and Ym2 crystals have been observed.
Although Ym1 and Ym2 crystals are often observed in mice at sites of eosinophilic inflammation, they are not made by eosinophils, but mainly by macrophages and epithelial cells, respectively. In vitro, protein crystallization typically starts from supersaturated solutions that support crystal nucleation. Several factors such as temperature and pH can affect the solubility of Ym1 and Ym2 in vivo and thus affect the nucleation and crystallization process. For Ym1 and Ym2 we noticed in vitro that a small drop in pH facilitates the crystallization process. Although the physiological pH is 7.4, during inflammation, there is a drop in pH. This drop in pH is the result of the infiltration and activation of inflammatory cells in the tissue, which leads to an increased energy and oxygen demand, accelerated glucose consumption via glycolysis and thus increased lactic acid secretion. In addition, we cannot exclude that in vivo, the nucleation process for Ym1/Ym2 is facilitated by interaction with ligands in the extracellular space (e.g. polysaccharide ligands or other – yet to be identified – specific ligands to Ym1/Ym2).
Reviewer #2 (Public Review):
Summary:
This interesting study addresses the ability of Ym1 protein crystals to promote pulmonary type 2 inflammation in vivo, in mice.
Strengths:
The data are extremely high quality, clearly presented, significantly extending previous work from this group on the type 2 immunogenicity of protein crystals.
Weaknesses:
There are no major weaknesses in this study. It would be interesting to see if Ym2 crystals behave similarly to Ym1 crystals in vivo. Some additional text in the Introduction and Discussion would enrich those sections.
We agree that this would be interesting to investigate, however, we choose to not include recombinant Ym2 crystal data in this report. However, we have further elaborated the discussion section including this point.
Recommendations for the authors:
Reviewer #1 (Recommendations For The Authors):
Suggestions for improved experiments and to strengthen findings:
I think additional data on the ability of Ym2 crystals to induce an immune response would be advantageous. I'm not by any means suggesting the authors repeat all the experiments with Ym2 crystals, but even just the ability to show that Ym2 could promote type 2 immunity in the acute OVA model, would help to strengthen the argument that these crystals in general function in a similar way. Alternatively, a discussion on whether these protein crystals may function in different scenarios/tissues or conditions could help in light of additional data
We agree that this is an interesting point to investigate, however, we choose to not include recombinant Ym2 crystal data in this report. However, we have further elaborated the discussion section including this point.
Measuring IL-33 in lung tissue is difficult to interpret as cells will express intracellular IL-33 that is not active and may explain why the results in Fig 2D are not overly convincing. It could just be that Ym1 crystals are changing the number of cells expressing IL-33 (e.g macrophages, or type 2 pneumocytes) Did the authors also measure active IL-33 release in the BAL fluid which may give a better indication of Ym1's ability to activate DAMPs?
We also measured active IL-33 release in the BAL fluid, but due to the limited sample availability we could only measure this in one of the two repeat experiments, resulting in non-significant results for the BAL fluid. However, certainly for the 6h timepoint we saw a similar trend in the BAL fluid as in the lung tissue, meaning higher levels of IL-33 in the Ym1 crystal group compared to the PBS and soluble Ym1 group.
Crystals in Fig 2F staining with Ym1 appear to be brighter in the soluble Ym1 group. Is this related to increased packing of Ym1 in the crystals formed in vivo as opposed to those formed in vitro? Aside from reduced amount of crystals that form when you give soluble Ym1, could the type of crystal also be influencing the ability of soluble Ym1 crystals to generate an immune response?
Our X-ray diffraction experiments show that the packing of Ym1 is identical for in vivo and in vitro grown crystals. Possibly the apparent difference in brightness is caused by stochastic staining by the antibody. In this regard we note that the crystals formed from soluble Ym1 after 24h also can appear as less bright in a similar fashion as recombinant Ym1 crystals.
Overall, the data and writing of the manuscript is presented to a very high standard
A few minor points:
- Fig 2F - a little unsure what the number in the left top corner of the images represented.
These numbers represent the picture numbers generated by the software, but as they don’t have any added value for the story, we removed these numbers from the images.
- Not clear why two different expression vectors were used - one for Ym1 and one for Ym2?
Because we observed that recombinant Ym2 is more poorly secreted in the mammalian cell culture supernatant as compared to recombinant Ym1, we produced Ym2 with an N-terminal hexahistidine-tag followed by a Tobacco Etch Virus (TEV)-protease cleavage site to facilitate its purification.
Reviewer #2 (Recommendations For The Authors):
The authors briefly outline in their Introduction potential Sources of Ym1/2 in vivo, highlighting monocytes, M2 macrophages, alveolar macrophages, neutrophils and epithelial cells. Do DCs also make detectable/meaningful amounts of Ym1/2 in vivo, particularly in type 2 settings?
In the introduction we only highlighted the main cellular sources of Ym1 and Ym2, but there is literature available stating/showing that Ym1/2 is not only expressed by macrophages, neutrophils, monocytes and epithelial cells, but can also be induced in DCs and mast cells. We added the word ‘mainly’ to this sentence in the introduction, to make clear that macrophages, neutrophils and monocytes are not the only sources of Ym1.
Given the nicely demonstrated similarity of recombinant Ym1 and Ym2 crystals, I think it is important for the authors to include at least initial data on the outcome of recombinant Ym2 crystal admin to mice, in comparison to their Ym1 data.
We agree that this is an interesting point to investigate, however, we choose to not include recombinant Ym2 crystal data in this report. However, we have further elaborated the discussion section including this point.
Given the generation of crystals following in vivo administration of soluble Ym1, albeit at a lower level than when crystals were administered, it would be interesting to see if increased concentrations of soluble material show a dose dependent increase in lung inflammation readouts.
We agree that this would be an interesting point to investigate. Alongside this we could also titrate down the crystal dose, to see if there is a dose dependent decrease in lung inflammation readouts. However, at this time, we choose to not investigate this further.
I couldn't easily follow the authors' Discussion about potential ability of anti Ym-1/2 Abs to dissolve Ym1/2 crystals (similar to what they have demonstrated for Abs vs Gal10 crystals). Have they addressed this possibility experimentally? If so, addition of such data to the manuscript would be extremely interesting, given the obvious potential Ym1/2 crystal dissolving Abs for investigation of the role of these in a range of different murine models of type 2 inflammation.
We agree that the phrasing of this part of the discussion can be unclear/confusing. We rephrased this part to make it clearer. However, we did not address the possibility of Ym1/2 crystal dissolving antibodies experimentally.
In the Results section, the authors briefly comment on the pro-type 2 nature of Ym1 crystals in relation to their previous work with uric acid and Gal10 crystals, proposing that the pulmonary type 2 response may be a 'generic response to crystals of different chemical composition'. The Discussion would be enriched by deeper exploration of this comment.
We have further elaborated the discussion section including this point.
Author Response
The following is the authors’ response to the current reviews.
We agree with the reviewer that the statistics are buried in a dense excel file without a read-me page. We will address this by making a summary excel page for p-values during the production process.
The following is the authors’ response to the original reviews.
eLife assessment
This important study uses genomically-engineered glypican alleles to demonstrate convincingly that Dally (not Dally-like protein [Dlp]) is the key contributor to formation of the Dpp/BMP morphogen gradient in the wing disc of Drosophila. The authors provide solid genetic evidence that, surprisingly, the core domain of Dally appears to suffice to trap Dpp at the cell surface. They conclude with a model according to which Dally modulates the range of Dpp signaling by interfering with Dpp's internalization by the Dpp receptor Thickveins.
Public Reviews:
Reviewer #1 (Public Review):
How morphogens spread within tissues remains an important question in developmental biology. Here the authors revisit the role of glypicans in the formation of the Dpp gradient in wing imaginal discs of Drosophila. They first use sophisticated genome engineering to demonstrate that the two glypicans of Drosophila are not equivalent despite being redundant for viability. They show that Dally is the relevant glypican for Dpp gradient formation. They then provide genetic evidence that, surprisingly, the core domain of Dally suffices to trap Dpp at the cell surface (suggesting a minor role for GAGs). They conclude with a model that Dally modulates the range of Dpp signaling by interfering with Dpp's degradation by Tkv. These are important conclusions, but more independent (biochemical/cell biological) evidence is needed.
As indicated above, the genetic evidence for the predominant role of Dally in Dpp protein/signalling gradient formation is strong. In passing, the authors could discuss why overexpressed Dlp has a negative effect on signaling, especially in the anterior compartment. The authors then move on to determine the role of GAG (=HS) chains of Dally. They find that in an overexpression assay, Dally lacking GAGs traps Dpp at the cell surface and, counterintuitively, suppresses signaling (fig 4 C, F). Both findings are unexpected and therefore require further validation and clarification, as outlined in a and b below.
a. In loss of function experiments (dallyDeltaHS replacing endogenous dally), Dpp protein is markedly reduced (fig 4R), as much as in the KO (panel Q), suggesting that GAG chains do contribute to trapping Dpp at the cell surface. This is all the more significant that, according to the overexpression essays, DallyDeltaHS seems more stable than WT Dally (by the way, this difference should also be assessed in the knock-ins, which is possible since they are YFP-tagged). The authors acknowledge that HS chains of Dally are critical for Dpp distribution (and signaling) under physiological conditions. If this is true, one can wonder why overexpressed dally core 'binds' Dpp and whether this is a physiologically relevant activity.
According to the overexpression assay, DallyDeltaHS seems more stable than WT Dally (Fig. 4B’, E’, 5A’, B’). As the reviewer suggested, we addressed the difference using the two knock-in alleles and found that DallyDeltaHS is more stable than WT Dally (Fig.4 L, M inset), further emphasizing the insufficient role of core protein of Dally for extracellular Dpp distribution.
In summary, we showed that, although Dally interacts with Dpp mainly through its core protein from the overexpression assay (Fig. 4E, I), HS chains are essential for extracellular Dpp distribution (Fig. 4R). Thus, the core protein of Dally alone is not sufficient for extracellular Dpp distribution under physiological conditions. These results raise a question about whether the interaction of core protein of Dally with Dpp is physiologically relevant. Since the increase of HS upon dally expression but not upon dlp expression resulted in the accumulation of extracellular Dpp (Fig. 2) and this accumulation was mainly through the core protein of Dally (Fig. 4E, I), we speculate that the interaction of the core protein of Dally with Dpp gives ligand specificity to Dally under physiological conditions.
To understand the importance of the interaction of core protein of Dally with Dpp under physiological conditions, it is important to identify a region responsible for the interaction. Our preliminary results overexpressing a dally mutant lacking the majority of core protein (but keeping the HS modified region intact) showed that HS chains modification was also lost. Although this is consistent with our results that enzymes adding HS chains also interact with the core protein of Dally (Fig. 4D), the dally mutant allele lacking the core protein would hamper us from distinguishing the role of core protein of Dally from HS chains.
Nevertheless, we can infer the importance of the interaction of core protein of Dally with Dpp using dally[3xHA-dlp, attP] allele, where dlp is expressed in dally expressing cells. Since Dally-like is modified by HS chains but does not interact with Dpp (Fig. 2, 4), dally[3xHA-dlp, attP] allele mimics a dally allele where HS chains are properly added but interaction of core protein with Dpp is lost. As we showed in Fig.3O, S, the allele could not rescue dallyKO phenotypes, consistent with the idea that interaction of core protein of Dally with Dpp is essential for Dpp distribution and signaling and HS chain alone is not sufficient for Dpp distribution.
b. Although the authors' inference that dallycore (at least if overexpressed) can bind Dpp. This assertion needs independent validation by a biochemical assay, ideally with surface plasmon resonance or similar so that an affinity can be estimated. I understand that this will require a method that is outside the authors' core expertise but there is no reason why they could not approach a collaborator for such a common technique. In vitro binding data is, in my view, essential.
We agree with the reviewer that a biochemical assay such as SPR helps us characterize the interaction of core protein of Dally and Dpp (if the interaction is direct), although the biochemical assay also would not demonstrate the interaction under the physiological conditions.
However, SPR has never been applied in the case of Dpp, probably because purifying functional refolded Dpp dimer from bacteria has previously been found to be stable only in low pH and be precipitated in normal pH buffer (Groppe J, et al., 1998)(Matsuda et al., 2021). As the reviewer suggests, collaborating with experts is an important step in the future.
Nevertheless, SPR was applied for the interaction between BMP4 and Dally (Kirkpatrick et al., 2006), probably because BMP4 is more stable in the normal buffer. Although the binding affinity was not calculated, SPR showed that BMP4 directly binds to Dally and this interaction was only partially inhibited by molar excess of exogenous HS, suggesting that BMP4 can interact with core protein of Dally as well as its HS chains. In addition, the same study applied Co-IP experiments using lysis of S2 cells and showed that Dpp and core protein of Dally are co-immunoprecipitated, although it does not demonstrate if the interaction is direct.
In a subsequent set of experiments, the authors assess the activity of a form of Dpp that is expected not to bind GAGs (DppDeltaN). Overexpression assays show that this protein is trapped by DallyWT but not dallyDeltaHS. This is a good first step validation of the deltaN mutation, although, as before, an invitro binding assay would be preferable.
Our overexpression assays actually showed that DppDeltaN is trapped by DallyWT and by dallyDeltaHS at similar levels (Fig. 5C), indicating that interaction of DppDeltaN and HS chains of Dally is largely lost but DppDeltaN can still interact with core protein of Dally.
We thank the reviewer for the suggesting the in vitro experiment. Although we decided not to develop biophysical experiments such as SPR for Dpp in this study due to the reasons discussed above, we would like to point out that our result is consistent with a previous Co-IP experiment using S2 cells showing that DppDeltaN loses interaction with heparin (Akiyama2008).
However, in contrast to our results, the same study also proposed by Co-IP experiments using S2 cells that DppDeltaN loses interaction with Dally (Akiyama2008). Although it is hard to conclude since western blotting was too saturated without loading controls and normalization (Fig. 1C in Akiyama 2008), and negative in vitro experiments do not necessarily demonstrate the lack of interaction in vivo. One explanation why the interaction was missed in the previous study is that some factors required for the interaction of DppDeltaN with core protein of Dally are missing in S2 cells. In this case, in vivo interaction assay we used in this study has an advantage to robustly detect the interaction.
Nevertheless, the authors show that DppDeltaN is surprisingly active in a knock-in strain. At face value (assuming that DeltaN fully abrogates binding to GAGs), this suggests that interaction of Dpp with the GAG chains of Dally is not required for signaling activity. This leads to authors to suggest (as shown in their final model) that GAG chains could be involved in mediating the interactions of Dally with Tkv (and not with Dpp. This is an interesting idea, which would need to be reconciled with the observation that the distribution of Dpp is affected in dallyDeltaHS knock-ins (item a above). It would also be strengthened by biochemical data (although more technically challenging than the experiments suggested above). In an attempt to determine the role of Dally (GAGs in particular) in the signaling gradient, the paper next addresses its relation to Tkv. They first show that reducing Tkv leads to Dpp accumulation at the cell surface, a clear indication that Tkv normally contributes to the degradation of Dpp. From this they suggest that Tkv could be required for Dpp internalisation although this is not shown directly. The authors then show that a Dpp gradient still forms upon double knockdown (Dally and Tkv). This intriguing observation shows that Dally is not strictly required for the spread of Dpp, an important conclusion that is compatible with early work by Lander suggesting that Dpp spreads by free diffusion. These result show that Dally is required for gradient formation only when Tkv is present. They suggest therefore that Dally prevents Tkv-mediated internalisation of Dpp. Although this is a reasonable inference, internalisation assays (e.g. with anti-Ollas or anti-HA Ab) would strengthen the authors' conclusions especially because they contradict a recent paper from the Gonzalez-Gaitan lab.
Thanks for suggesting the internalization assay. As we discussed in the discussion, our results suggest that extracellular Dpp distribution is severely reduced in dally mutants due to Tkv mediated internalization of Dpp (Fig. 6). Thus, extracellular Dpp available for labelling with nanobody is severely reduced in dally mutants, which can explain the reduced internalization of Dpp in dally mutants in the internalization assay. Therefore, we think that the nanobody internalization assay would not distinguish the two contradicting possibilities.
The paper ends with a model suggesting that HS chains have a dual function of suppressing Tkv internalisation and stimulating signaling. This constitutes a novel view of a glypican's mode of action and possibly an important contribution of this paper. As indicated above, further experiments could considerably strengthen the conclusion. Speculation on how the authors imagine that GAG chains have these activities would also be warranted.
Thank you very much!
Reviewer #2 (Public Review):
The authors are trying to distinguish between four models of the role of glypicans (HSPGs) on the Dpp/BMP gradient in the Drosophila wing, schematized in Fig. 1: (1) "Restricted diffusion" (HSPGs transport Dpp via repetitive interaction of HS chains with Dpp); (2) "Hindered diffusion" (HSPGs hinder Dpp spreading via reversible interaction of HS chains with Dpp); (3) "Stabilization" (HSPGs stabilize Dpp on the cell surface via reversible interaction of HS chains with Dpp that antagonizes Tkv-mediated Dpp internalization); and (4) "Recycling" (HSPGs internalize and recycle Dpp).
To distinguish between these models, the authors generate new alleles for the glypicans Dally and Dally-like protein (Dlp) and for Dpp: a Dally knock-out allele, a Dally YFP-tagged allele, a Dally knock-out allele with 3HA-Dlp, a Dlp knock-out allele, a Dlp allele containing 3-HA tags, and a Dpp lacking the HS-interacting domain. Additionally, they use an OLLAS-tag Dpp (OLLAS being an epitope tag against which extremely high affinity antibodies exist). They examine OLLAS-Dpp or HA-Dpp distribution, phospho-Mad staining, adult wing size.
They find that over-expressed Dally - but not Dlp - expands Dpp distribution in the larval wing disc. They find that the Dally[KO] allele behaves like a Dally strong hypomorph Dally[MH32]. The Dally[KO] - but not the Dlp[KO] - caused reduced pMad in both anterior and posterior domains and reduced adult wing size (particularly in the Anterior-Posterior axis). These defects can be substantially corrected by supplying an endogenously tagged YFP-tagged Dally. By contrast, they were not rescued when a 3xHA Dlp was inserted in the Dally locus. These results support their conclusion that Dpp interacts with Dally but not Dlp.
They next wanted to determine the relative contributions of the Dally core or the HS chains to the Dpp distribution. To test this, they over-expressed UAS-Dally or UAS-Dally[deltaHS] (lacking the HS chains) in the dorsal wing. Dally[deltaHS] over-expression increased the distribution of OLLAS-Dpp but caused a reduction in pMad. Then they write that after they normalize for expression levels, they find that Dally[deltaHS] only mildly reduces pMad and this result indicates a major contribution of the Dally core protein to Dpp stability.
Thanks for the comments. We actually showed that compared with Dally overexpression, Dally[deltaHS] overexpression only mildly reduces extracellular Dpp accumulation (Fig. 4I). This indicates a major contribution of the Dally core protein to interaction with Dpp, although the interaction is not sufficient to sustain extracellular Dpp distribution and signaling gradient.
The "normalization" is a key part of this model and is not mentioned how the normalization was done. When they do the critical experiment, making the Dally[deltaHS] allele, they find that loss of the HS chains is nearly as severe as total loss of Dally (i.e., Dally[KO]). Additionally, experimental approaches are needed here to prove the role of the Dally core.
Since the expression level of Dally[deltaHS] is higher than Dally when overexpressed, we normalized extracellular Dpp distribution (a-Ollas staining) against GFP fluorescent signal (Dally or Dally[deltaHS]). To do this, we first extracted both signal along the A-P axis from the same ROI in the previous version. The ratio was calculated by dividing the intensity of a-Ollas staining with the intensity of GFP fluorescent signal at a given position x. The average profile from each normalized profile was generated and plotted using the script described in the method (wingdisc_comparison.py) as other pMad or extracellular staining profiles.
Although this analysis provides normalized extracellular Dpp accumulation at different positions along the A-P axis, we are more interested in the total amount of Dpp or DppDeltaN accumulation upon Dally or dallyDeltaHS expression. Therefore, in the revised ms, we decided to normalize total amount of extracellular Dpp against the level of Dally or Dally[deltaHS] by dividing total signal intensity of extracellular Dpp staining (ExOllas staining) by total GFP fluorescent signal (Dally or Dally[deltaHS]) around the Dpp producing cells in each wing disc. Statistical analysis showed that accumulation of extracellular Dpp is only slightly reduced without HS chains (Fig.4I), indicating that Dally interacts with Dpp mainly through its core protein.
We agree with the reviewer that additional experimental approaches are needed to address the role of the core protein of Dally. As we discussed in the response to the reviewer1, to understand the importance of the interaction of core protein of Dally with Dpp, it is important to identify a region responsible for the interaction. Our preliminary results overexpressing a dally mutant lacking the majority of core protein (but keeping the HS modified region intact) showed that HS chains modification was also lost. Although this is consistent with our results that enzymes adding HS chains also interact with the core protein of Dally (Fig. 4D), the dally mutant allele lacking the core protein would hamper us from distinguishing the role of the core protein of Dally from HS chains.
Nevertheless, we can infer the importance of the interaction of core protein of Dally with Dpp using dally[3xHA-dlp, attP] allele, where dlp is expressed in dally expressing cells. Since Dally-like is modified by HS chains but does not interact with Dpp (Fig. 2, 4), dally[3xHA-dlp, attP] allele mimics a dally allele where HS chains are properly added but interaction of core protein with Dpp is lost. As we showed in Fig.3O, S, the allele could not rescue dallyKO phenotypes, consistent with the idea that interaction of core protein of Dally with Dpp is essential for Dpp distribution and signaling.
Prior work has shown that a stretch of 7 amino acids in the Dpp N-terminal domain is required to interact with heparin but not with Dpp receptors (Akiyama, 2008). The authors generated an HA-tagged Dpp allele lacking these residues (HA-dpp[deltaN]). It is an embryonic lethal allele, but they can get some animals to survive to larval stages if they also supply a transgene called “JAX” containing dpp regulatory sequences. In the JAX; HA-dpp[deltaN] mutant background, they find that the distribution and signaling of this Dpp molecule is largely normal. While over-expressed Dally can increase the distribution of HA-dpp[deltaN], over-expression of Dally[deltaHS] cannot. These latter results support the model that the HS chains in Dally are required for Dpp function but not because of a direct interaction with Dpp.
Our overexpression assays actually showed that both Dally and Dally[deltaHS] can accumulate Dpp upon overexpression and the accumulation of Dpp is comparable after normalization (Fig. 5C), consistent with the idea that interaction of DppdeltaN and HS chains are largely lost. As the reviewer pointed out, these results support the model that the HS chains in Dally are required for Dpp function but not because of a direct interaction with Dpp.
In the last part of the results, they attempt to determine if the Dpp receptor Thickveins (Tkv) is required for Dally-HS chains interaction. The 2008 (Akiyama) model posits that Tkv activates pMad downstream of Dpp and also internalizes and degrades Dpp. A 2022 (Romanova-Michaelides) model proposes that Dally (not Tkv) internalizes Dpp.
To distinguish between these models, the authors deplete Tkv from the dorsal compartment of the wing disc and found that extracellular Dpp increased and expanded in that domain. These results support the model that Tkv is required to internalize Dpp.
They then tested the model that Dally antagonizes Tkv-mediated Dpp internalization by determining whether the defective extracellular Dpp distribution in Dally[KO] mutants could be rescued by depleting Tkv. Extracellular Dpp did increase in the D vs V compartment, potentially providing some support for their model. However, there are no statistics performed, which is needed for full confidence in the results. The lack of statistics is particularly problematic (1) when they state that extracellular Dpp does not rise in ap>tkv RNAi vs ap>tkv RNAi, dally[KO] wing discs (Fig. 6E) or (2) when they state that extracellular Dpp gradient expanded in the dorsal compartment when tkv was dorsally depleted in dally[deltaHS] mutants (Fig. 6I). These last two experiments are important for their model but the differences are assessed only visually. In fact, extracellular Dpp in ap>tkv RNAi, dally[KO] (Fig. 6B) appears to be lower than extracellular Dpp in ap>tkv RNAi (Fig. 6A) and the histogram of Dpp in ap>tkv RNAi, dally[KO] is actually a bit lower than Dpp in ap>tkv RNAi, But the author claim that there is no difference between the two. Their conclusion would be strengthened by statistical analyses of the two lines.
We provided statistics for all the quantifications for pMad and extracellular Dpp distribution as supplementary data. In the previous version, we argued that extracellular Dpp level in ap>tkvRNAi, dallyKO (Fig.6B) does not increase compared with that in ap>tkvRNAi (Fig.6A). Statistical analysis (t-test) showed that the extracellular Dpp level in Fig. 6B is similar to or lower than that in Fig. 6A (Fig. 6E), confirming our conclusion. Statistical analysis (t-test) also confirmed that extracellular Dpp distribution expanded when tkv was knocked down in dallyHS mutants (Fig. 6I).
Strengths:
- New genomically-engineered alleles
A considerable strength of the study is the generation and characterization of new Dally, Dlp and Dpp alleles. These reagents will be of great use to the field.
Thanks. We hope that these resources are indeed useful to the field.
- Surveying multiple phenotypes
The authors survey numerous parameters (Dpp distribution, Dpp signaling (pMad) and adult wing phenotypes) which provides many points of analysis.
Thanks!
Weaknesses:
- Confusing discussion regarding the Dally core vs HS in Dpp stability. They don't provide any measurements or information on how they "normalize" for the level of Dally vs Dally[deltaHS]? This is important part of their model that currently is not supported by any measurements.
We explained how we normalized in the above section and updated the method section in the revised ms.
- Lacking quantifications and statistical analyses:
a. Why are statistical significance for histograms (pMad and Dpp distribution) not supplied? These histograms provide the key results supporting the authors' conclusions but no statistical tests/results are presented. This is a pervasive shortcoming in the current study.
Thanks. We provided t-test analyses together with the raw data as supplementary data.
b. dpp[deltaN] with JAX transgene - it would strengthen the study to supply quantitative data on the percent survival/lethal stage of dpp[deltaN] mutants with or without the JAK transgene
In this study, we are interested in the role of dpp[deltaN] during the wing disc development. Therefore, we decided not to perform the detailed analysis on the percent survival/lethal stage of dpp[deltaN] mutants with or without the JAX transgene in the current study. Nevertheless, the fact that dpp[deltaN] allele is maintained with a balanced stock and JAX;dpp[deltaN] allele can be maintained as homozygous stock indicates that the lethality of dpp[deltaN] allele comes from the early stages. Indeed, our preliminary results showed that pMad signal is severely lost in the dpp[deltaN] embryo without JAX (data not shown), indicating that the allele is lethal at early embryonic stages.
c. The graphs on wing size etc should start at zero.
Thanks. We corrected this in the current ms.
d. The sizes of histograms and graphs in each figure should be increased so that the reader can properly assess them. Currently, they are very small.
Thanks. We changed the sizes in the current ms.
The authors' model is that Dally (not Dlp) is required for Dpp distribution and signaling but that this is not due to a direct interaction with Dpp. Rather, they posit that Dally-HS antagonize Tkv-mediated Dpp internalization. Currently the results of the experiments could be considered consistent with their model, but as noted above, the lack of statistical analyses of some parameters is a weakness.
Thanks. We now performed and provided the statistical analyses in the revised ms.
One problematic part of their result for me is the role of the Dally core protein (Fig. 7B). There is a mis-match between the over-expression results and Dally allele lacking HS (but containing the core). Finally, their results support the idea that one or more as-yet unidentified proteins interact with Dally-HS chains to control Dpp distribution and signaling in the wing disc.
Our results simply suggest that Dpp can interact with Dally mainly through core protein but this interaction is not sufficient to sustain extracellular Dpp gradient formation under physiological conditions (dallyDeltaHS) (Fig. 4Q). We find that the mis-match is not problematic if the role of Dally is not simply mediated through interaction with Dpp. We speculate that interaction of Dpp and core protein of Dally is transient and not sufficient to sustain the Dpp gradient without HS chains of Dally stabilizing extracellular Dpp distribution by blocking Tkv-mediated Dpp internalization.
There is much debate and controversy in the Dpp morphogen field. The generation of new, high quality alleles in this study will be useful to Drosophila community, and the results of this study support the concept that Tkv but not Dally regulate Dpp internalization. Thus the work could be impactful and fuel new debates among morphogen researchers.
Thanks.
The manuscript is currently written in a manner that really is only accessible to researchers who work on the Dpp gradient. It would be very helpful for the authors to re-write the manuscript and carefully explain in each section of the results (1) the exact question that will be asked, (2) the prior work on the topic, (3) the precise experiment that will be done, and (4) the predicted results. This would make the study more accessible to developmental biologists outside of the morphogen gradient and Drosophila communities.
Thanks. We modified texts and changed the order of Fig.5. We hope that the changes make this study more accessible to developmental biologists outside of the field.
Joint Public Review:
The authors are trying to distinguish between four models of the role of glypicans (HSPGs) on the Dpp/BMP gradient in the Drosophila wing, schematized in Fig. 1: (1) "Restricted diffusion" (HSPGs transport Dpp via repetitive interaction of HS chains with Dpp); (2) "Hindered diffusion" (HSPGs hinder Dpp spreading via reversible interaction of HS chains with Dpp); (3) "Stabilization" (HSPGs stabilize Dpp on the cell surface via reversible interaction of HS chains with Dpp that antagonizes Tkv-mediated Dpp internalization); and (4) "Recycling" (HSPGs internalize and recycle Dpp).
To distinguish between these models, the authors generate new alleles for the glypicans Dally and Dally-like protein (Dlp) and for Dpp: a Dally knock-out allele, a Dally YFP-tagged allele, a Dally knock-out allele with 3HA-Dlp, a Dlp knock-out allele, a Dlp allele containing 3-HA tags, and a Dpp lacking the HS-interacting domain. Additionally, they use an OLLAS-tag Dpp (OLLAS being an epitope tag against which extremely high affinity antibodies exist). They examine OLLAS-Dpp or HA-Dpp distribution, phospho-Mad staining, adult wing size.
They find that over-expressed Dally - but not Dlp - expands Dpp distribution in the larval wing disc. They find that the Dally[KO] allele behaves like a Dally strong hypomorph Dally[MH32]. The Dally[KO] - but not the Dlp[KO] - caused reduced pMad in both anterior and posterior domains and reduced adult wing size (particularly in the Anterior-Posterior axis). These defects can be substantially corrected by supplying an endogenously tagged YFP-tagged Dally. By contrast, they were not rescued when a 3xHA Dlp was inserted in the Dally locus. These results support their conclusion that Dpp interacts with Dally but not Dlp.
They next wanted to determine the relative contributions of the Dally core or the HS chains to the Dpp distribution. To test this, they over-expressed UAS-Dally or UAS-Dally[deltaHS] (lacking the HS chains) in the dorsal wing. Dally[deltaHS] over-expression increased the distribution of OLLAS-Dpp but caused a reduction in pMad. They do a critical experiment, making the Dally[deltaHS] allele, they find that loss of the HS chains is nearly as severe as total loss of Dally (i.e., Dally[KO]). These results indicate that the HS are critical for Dally's role in Dpp distribution and signaling.
Prior work has shown that a stretch of 7 amino acids in the Dpp N-terminal domain is required to interact with heparin but not with Dpp receptors (Akiyama, 2008). The authors generated an HA-tagged Dpp allele lacking these residues (HA-dpp[deltaN]). It is an embryonic lethal allele, but they can get some animals to survive to larval stages if they also supply a transgene called "JAK" containing dpp regulatory sequences. In the JAK; HA-dpp[deltaN] mutant background, they find that the distribution and signaling of this Dpp molecule is largely normal. While over-expressed Dally can increase the distribution of HA-dpp[deltaN], over-expression of Dally[deltaHS] cannot. These latter results support the model that the HS chains in Dally are required for Dpp function but not because of a direct interaction with Dpp.
In the last part of the results, they attempt to determine if the Dpp receptor Thickveins (Tkv) is required for Dally-HS chains interaction. The 2008 (Akiyama) model posits that Tkv activates pMad downstream of Dpp and also internalizes and degrades Dpp. A 2022 (Romanova-Michaelides) model proposes that Dally (not Tkv) internalizes Dpp. To distinguish between these models, the authors deplete Tkv from the dorsal compartment of the wing disc and found that extracellular Dpp increased and expanded in that domain. These results support the model that Tkv is required to internalize Dpp. They then tested the model that Dally antagonizes Tkv-mediated Dpp internalization by determining whether the defective extracellular Dpp distribution in Dally[KO] mutants could be rescued by depleting Tkv. Extracellular Dpp did increase in the D vs V compartment, potentially providing some support for their model. The results are statistically significant but the statistics are buried in an excel file without a read-me page. The code for the statistics is available from Github. These p values should be made more readily accessible and/or intelligible to the reader.
Strengthens:<br /> 1. New genomically-engineered alleles<br /> A considerable strength of the study is the generation and characterization of new Dally, Dlp and Dpp alleles. These reagents will be of great use to the field.
2. Surveying multiple phenotypes<br /> The authors survey numerous parameters (Dpp distribution, Dpp signaling (pMad) and adult wing phenotypes) which provides many points of analysis.
Weaknesses (minor):<br /> 1. The results are statistically significant but the statistics are buried in a dense excel file without a read-me page. The code for the statistics is available from Github. These p values should be made more readily accessible to the reader.
An appraisal of whether the authors achieved their aims, and whether the results support their conclusions.<br /> The authors' model is that Dally (not Dlp) is required for Dpp distribution and signaling but that this is not due to a direct interaction with Dpp. Rather, they posit that Dally-HS antagonize Tkv-mediated Dpp internalization. Currently the results of the experiments could be considered consistent with their model. Finally, their results support the idea that one or more as-yet unidentified proteins interact with Dally-HS chains to control Dpp distribution and signaling in the wing disc.
There is much debate and controversy in the Dpp morphogen field. The generation of new, high quality alleles in this study will be useful to Drosophila community, and the results of this study support the concept that Tkv but not Dally regulate Dpp internalization. Thus the work could be impactful and fuel new debates among the morphogen researchers.
Rome they're tagged to the inbox page
Ryder, has a tag called #inbox inside of Roam. All the stuff that he reads, goes inside of ReadWise, and from ReadWise to the Inbox inside of Roam where he processes that information.
Author Response:
Reviewer #1 (Public Review):
[...] Weaknesses:
- I feel the authors need to justify why flow-crushing helps localization specificity. There is an entire family of recent papers that aim to achieve higher localization specificity by doing the exact opposite. Namely, MT or ABC fRMRI aims to increase the localization specificity by highlighting the intravascular BOLD by means of suppressing non-flowing tissue. To name a few:
Priovoulos, N., de Oliveira, I.A.F., Poser, B.A., Norris, D.G., van der Zwaag, W., 2023. Combining arterial blood contrast with BOLD increases fMRI intracortical contrast. Human Brain Mapping hbm.26227. https://doi.org/10.1002/hbm.26227.
Pfaffenrot, V., Koopmans, P.J., 2022. Magnetization Transfer weighted laminar fMRI with multi-echo FLASH. NeuroImage 119725. https://doi.org/10.1016/j.neuroimage.2022.119725
Schulz, J., Fazal, Z., Metere, R., Marques, J.P., Norris, D.G., 2020. Arterial blood contrast ( ABC ) enabled by magnetization transfer ( MT ): a novel MRI technique for enhancing the measurement of brain activation changes. bioRxiv. https://doi.org/10.1101/2020.05.20.106666
Based on this literature, it seems that the proposed method will make the vein problem worse, not better. The authors could make it clearer how they reason that making GE-BOLD signals more extra-vascular weighted should help to reduce large vein effects.
The empirical evidence for the claim that flow crushing helps with the localization specificity should be made clearer. The response magnitude with and without flow crushing looks pretty much identical to me (see Fig, 6d). It's unclear to me what to look for in Fig. 5. I cannot discern any layer patterns in these maps. It's too noisy. The two maps of TE=43ms look like identical copies from each other. Maybe an editorial error?
The authors discuss bipolar crushing with respect to SE-BOLD where it has been previously applied. For SE-BOLD at UHF, a substantial portion of the vein signal comes from the intravascular compartment. So I agree that for SE-BOLD, it makes sense to crush the intravascular signal. For GE-BOLD however, this reasoning does not hold. For GE-BOLD (even at 3T), most of the vein signal comes from extravascular dephasing around large unspecific veins, and the bipolar crushing is not expected to help with this.
The authors would like to clarify that the velocity-nulling gradient is NOT designed to suppress all the contributions from intravascular blood. Instead, we tried to find a balance so that the VN gradient maximally suppressed the macrovascular signal in unspecific veins but minimally attenuated the microvascular signal in specific capillary bed. We acknowledge the reviewer's concern regarding the potential extravascular contributions from large, non-specific vessels. This aspect will be thoroughly evaluated and addressed in the revised manuscript. Additionally, we will make clarifications in other parts that may have cause the reviewer’s misunderstandings.
- The bipolar crushing is limited to one single direction of flow. This introduces a lot of artificial variance across the cortical folding pattern. This is not mentioned in the manuscript. There is an entire family of papers that perform layer-fmri with black-blood imaging that solves this with a 3D contrast preparation (VAPER) that is applied across a longer time period, thus killing the blood signal while it flows across all directions of the vascular tree. Here, the signal cruising is happening with a 2D readout as a "snap-shot" crushing. This does not allow the blood to flow in multiple directions. VAPER also accounts for BOLD contaminations of larger draining veins by means of a tag-control sampling. The proposed approach here does not account for this contamination.
Chai, Y., Li, L., Huber, L., Poser, B.A., Bandettini, P.A., 2020. Integrated VASO and perfusion contrast: A new tool for laminar functional MRI. NeuroImage 207, 116358. https://doi.org/10.1016/j.neuroimage.2019.116358
Chai, Y., Liu, T.T., Marrett, S., Li, L., Khojandi, A., Handwerker, D.A., Alink, A., Muckli, L., Bandettini, P.A., 2021. Topographical and laminar distribution of audiovisual processing within human planum temporale. Progress in Neurobiology 102121. https://doi.org/10.1016/j.pneurobio.2021.102121
If I would recommend anyone to perform layer-fMRI with blood crushing, it seems that VAPER is the superior approach. The authors could make it clearer why users might want to use the unidirectional crushing instead.
We acknowledge that the degree of velocity nulling varies across the cortical folding pattern. We intend to discuss potential solutions to address this variance, and these may be implemented in the revised manuscript as appropriate. Furthermore, we will provide a comprehensive discussion on the advantages and disadvantages of both CBV-based and BOLD-based approaches.
- The comparison with VASO is misleading. The authors claim that previous VASO approaches were limited by TRs of 8.2s. The authors might be advised to check the latest literature of the last years. Koiso et al. performed whole brain layer-fMRI VASO at 0.8mm at 3.9 seconds (with reliable activation), 2.7 seconds (with unconvincing activation pattern, though), and 2.3 (without activation). Also, whole brain layer-fMRI BOLD at 0.5mm and 0.7mm has been previously performed by the Juelich group at TRs of 3.5s (their TR definition is 'fishy' though).
Koiso, K., Müller, A.K., Akamatsu, K., Dresbach, S., Gulban, O.F., Goebel, R., Miyawaki, Y., Poser, B.A., Huber, L., 2023. Acquisition and processing methods of whole-brain layer-fMRI VASO and BOLD: The Kenshu dataset. Aperture Neuro 34. https://doi.org/10.1101/2022.08.19.504502
Yun, S.D., Pais‐Roldán, P., Palomero‐Gallagher, N., Shah, N.J., 2022. Mapping of whole‐cerebrum resting‐state networks using ultra‐high resolution acquisition protocols. Human Brain Mapping. https://doi.org/10.1002/hbm.25855
Pais-Roldan, P., Yun, S.D., Palomero-Gallagher, N., Shah, N.J., 2023. Cortical depth-dependent human fMRI of resting-state networks using EPIK. Front. Neurosci. 17, 1151544. https://doi.org/10.3389/fnins.2023.1151544
The authors are correct that VASO is not advised as a turn-key method for lower brain areas, incl. Hippocampus and subcortex. However, the authors use this word of caution that is intended for inexperienced "users" as a statement that this cannot be performed. This statement is taken out of context. This statement is not from the academic literature. It's advice for the 40+ user base that wants to perform layer-fMRI as a plug-and-play routine tool in neuroscience usage. In fact, sub-millimeter VASO is routinely being performed by MRI-physicists across all brain areas (including deep brain structures, hippocampus etc). E.g. see Koiso et al. and an overview lecture from a layer-fMRI workshop that I had recently attended: https://youtu.be/kzh-nWXd54s?si=hoIJjLLIxFUJ4g20&t=2401
Thus, the authors could embed this phrasing into the context of their own method that they are proposing in the manuscript. E.g. the authors could state whether they think that their sequence has the potential to be disseminated across sites, considering that it requires slow offline reconstruction in Matlab? Do the authors think that the results shown in Fig. 6c are suggesting turn-key acquisition of a routine mapping tool? In my humble opinion, it looks like random noise, with most of the activation outside the ROI (in white matter).
Those literatures will be included and discussed in the revised manuscript. Furthermore, we are considering the exclusion of the LGN results presented in Figure 6, as they may divert attention from the primary focus of the study.
We are enthusiastic about sharing our imaging sequence, provided its usefulness is conclusively established. However, it's important to note that without an online reconstruction capability, such as the ICE, the practical utility of the sequence may be limited. Unfortunately, we currently don’t have the manpower to implement the online reconstruction. Nevertheless, we are more than willing to share the offline reconstruction codes upon request.
- The repeatability of the results is questionable. The authors perform experiments about the robustness of the method (line 620). The corresponding results are not suggesting any robustness to me. In fact, the layer profiles in Fig. 4c vs. Fig 4d are completely opposite. The location of peaks turns into locations of dips and vice versa. The methods are not described in enough detail to reproduce these results. The authors mention that their image reconstruction is done "using in-house MATLAB code" (line 634). They do not post a link to github, nor do they say if they share this code.
It is not trivial to get good phase data for fMRI. The authors do not mention how they perform the respective coil-combination. No data are shared for reproduction of the analysis.
There may have been a misunderstanding regarding the presentation in Figure 4, which illustrates the impact of TEs and the VN gradient. To enhance clarity and avoid further confusion, we will redesign this figure for improved comprehension.
The authors are open to sharing the MATLAB codes associated with our study. However, we were limited by manpower for refining and enhancing the readability of these codes for broader use.
Regarding the coil combination, we utilized an adaptive coil combination approach as described in the paper by Walsh DO, Gmitro AF, and Marcellin MW, titled 'Adaptive reconstruction of phased array MR imagery' (Magnetic Resonance in Medicine 2000; 43:682-690). The MATLAB code for this method was implemented by Dr. Diego Hernando. We will include a link for downloading this code in the revised manuscript for the convenience of interested readers.
- The application of NODRIC is not validated. Previous applications of NORDIC at 3T layer-fMRI have resulted in mixed success. When not adjusted for the right SNR regime it can result in artifactual reductions of beta scores, depending on the SNR across layers. The authors could validate their application of NORDIC and confirm that the average layer-profiles are unaffected by the application of NORDIC. Also, the NORDIC version should be explicitly mentioned in the manuscript.
Akbari, A., Gati, J.S., Zeman, P., Liem, B., Menon, R.S., 2023. Layer Dependence of Monocular and Binocular Responses in Human Ocular Dominance Columns at 7T using VASO and BOLD (preprint). Neuroscience. https://doi.org/10.1101/2023.04.06.535924
Knudsen, L., Guo, F., Huang, J., Blicher, J.U., Lund, T.E., Zhou, Y., Zhang, P., Yang, Y., 2023. The laminar pattern of proprioceptive activation in human primary motor cortex. bioRxiv. https://doi.org/10.1101/2023.10.29.564658
During our internal testing, we observed that the NORDIC denoising process did not alter the activation patterns. These findings will be incorporated into the revised manuscript. The details of NORDIC will be provided as well.
Reviewer #2 (Public Review):
[...] The well-known double peak feature in M1 during finger tapping was used as a test-bed to evaluate the spatial specificity. They were indeed able to demonstrate two distinct peaks in group-level laminar profiles extracted from M1 during finger tapping, which was largely free from superficial bias. This is rather intriguing as, even at 7T, clear peaks are usually only seen with spatially specific non-BOLD sequences. This is in line with their simple simulations, which nicely illustrated that, in theory, intravascular macrovascular signals should be suppressible with only minimal suppression of microvasculature when small b-values of the VN gradients are employed. However, the authors do not state how ROIs were defined making the validity of this finding unclear; were they defined from independent criteria or were they selected based on the region mostly expressing the double peak, which would clearly be circular? In any case, results are based on a very small sub-region of M1 in a single slice - it would be useful to see the generalizability of superficial-bias-free BOLD responses across a larger portion of M1.
Given the individual variations in the location of the M1 region, we opted for manual selection of the ROI. In the revised manuscript, we plan to explore and implement an independent criterion for ROI selection to enhance the objectivity and reproducibility of our methodology.
As repeatedly mentioned by the authors, a laminar fMRI setup must demonstrate adequate functional sensitivity to detect (in this case) BOLD responses. The sensitivity evaluation is unfortunately quite weak. It is mainly based on the argument that significant activation was found in a challenging sub-cortical region (LGN). However, it was a single participant, the activation map was not very convincing, and the demonstration of significant activation after considerable voxel-averaging is inadequate evidence to claim sufficient BOLD sensitivity. How well sensitivity is retained in the presence of VN gradients, high acceleration factors, etc., is therefore unclear. The ability of the setup to obtain meaningful functional connectivity results is reassuring, yet, more elaborate comparison with e.g., the conventional BOLD setup (no VN gradients) is warranted, for example by comparison of tSNR, quantification and comparison of CNR, illustration of unmasked-full-slice activation maps to compare noise-levels, comparison of the across-trial variance in each subject, etc. Furthermore, as NORDIC appears to be a cornerstone to enable submillimeter resolution in this setup at 3T, it is critical to evaluate its impact on the data through comparison with non-denoised data, which is currently lacking.
We appreciate the reviewer’s comments. Those issues will be addressed carefully.
Reviewer #3 (Public Review):
[...] Weaknesses: - Although the VASO acquisition is discussed in the introduction section, the VN-sequence seems closer to diffusion-weighted functional MRI. The authors should make it more clear to the reader what the differences are, and how results are expected to differ. Generally, it is not so clear why the introduction is so focused on the VASO acquisition (which, curiously, lacks a reference to Lu et al 2013). There are many more alternatives to BOLD-weighted imaging for fMRI. CBF-weighted ASL and GRASE have been around for a while, ABC and double-SE have been proposed more recently.
The principal distinction between DW-fMRI and our methodology lies in the level of the b-value employed. DW-fMRI typically measures cellular swelling by utilizing a b-value greater than 1000 s/mm^2 (e.g. 1800). Conversely, our Velocity Nulling functional MRI (VN-fMRI) approach continues to assess hemodynamic responses, utilizing a smaller b-value specifically for the suppression of signals from draining veins. In addition, other layer-fMRI methods will be discussed.
- The comparison in Figure 2 for different b-values shows % signal changes. However, as the baseline signal changes dramatically with added diffusion weighting, this is rather uninformative. A plot of t-values against cortical depth would be much more insightful.
- Surprisingly, the %-signal change for a b-value of 0 is not significantly different from 0 in the gray matter. This raises some doubts about the task or ROI definition. A finger-tapping task should reliably engage the primary motor cortex, even at 3T, and even in a single participant.
- The BOLD weighted images in Figure 3 show a very clear double-peak pattern. This contradicts the results in Figure 2 and is unexpected given the existing literature on BOLD responses as a function of cortical depth.
In our study, the TE in Figure 2 is shorter than that in Figure 3 (33 ms versus 43 ms). It has been reported in the literature that BOLD fMRI with a shorter TE tends to include a greater intravascular contribution. Acknowledging this, we plan to repeat the experiments with a controlled TE to ensure consistency in our results.
- Given that data from Figures 2, 3, and 4 are derived from a single participant each, order and attention affects might have dramatically affected the observed patterns. Especially for Figure 4, neither BOLD nor VN profiles are really different from 0, and without statistical values or inter-subject averaging, these cannot be used to draw conclusions from.
The order of the experiments were randomized to ensure unbiased results.
It is important to note that the error bars presented in Figures 2, 3, and 4 do not represent the standard deviation of the residual fitting error. Instead, they illustrate the variation across voxels within a specific layer. This approach may lead to the error bars being influenced by the selection of the Region of Interest (ROI). In light of this, we intend to refine our statistical methodologies in the revised manuscript to address this issue.
- In Figure 5, a phase regression is added to the data presented in Figure 4. However, for a phase regression to work, there has to be a (macrovascular) response to start with. As none of the responses in Figure 4 are significant for the single participant dataset, phase regression should probably not have been undertaken. In this case, the functional 'responses' appear to increase with phase regression, which is contra-intuitive and deserves an explanation.
- Consistency of responses is indeed expected to increase by a removal of the more variable vascular component. However, the microvascular component is always expected to be smaller than the combination of microvascular + macrovascular responses. Note that the use of %signal changes may obscure this effect somewhat because of the modified baseline. Another expected feature of BOLD profiles containing both micro- and microvasculature is the draining towards the cortical surface. In the profiles shown in Figure 7, this is completely absent. In the group data, no significant responses to the task are shown anywhere in the cortical ribbon.
- Although I'd like to applaud the authors for their ambition with the connectivity analysis, I feel that acquisitions that are so SNR starved as to fail to show a significant response to a motor task should not be used for brain wide directed connectivity analysis.
We agree that exploring brain-wide directed functional connectivity may be overly ambitious at this stage, particularly before the VN-fMRI technique has been comprehensively evaluated and validated. In the revised manuscript, we will focus more on examining the characteristics of the layer-dependent BOLD signal rather than delving into layer-dependent functional connectivity.
Reviewer #1 (Public Review):
Summary:
This study aims to provide imaging methods for users of the field of human layer-fMRI. This is an emerging field with 240 papers published so far. Different than implied in the manuscript, 3T is well represented among those papers. E.g. see the papers below that are not cited in the manuscript. Thus, the claim on the impact of developing 3T methodology for wider dissemination is not justified. Specifically, because some of the previous papers perform whole brain layer-fMRI (also at 3T) in more efficient, and more established procedures.
The authors implemented a sequence with lots of nice features. Including their own SMS EPI, diffusion bipolar pulses, eye-saturation bands, and they built their own reconstruction around it. This is not trivial. Only a few labs around the world have this level of engineering expertise. I applaud this technical achievement. However, I doubt that any of this is the right tool for layer-fMRI, nor does it represent an advancement for the field. In the thermal noise dominated regime of sub-millimeter fMRI (especially at 3T), it is established to use 3D readouts over 2D (SMS) readouts. While it is not trivial to implement SMS, the vendor implementations (as well as the CMRR and MGH implementations) are most widely applied across the majority of current fMRI studies already. The author's work on this does not serve any previous shortcomings in the field.
The mechanism to use bi-polar gradients to increase the localization specificity is doubtful to me. In my understanding, killing the intra-vascular BOLD should make it less specific. Also, the empirical data do not suggest a higher localization specificity to me.
Embedding this work in the literature of previous methods is incomplete. Recent trends of vessel signal manipulation with ABC or VAPER are not mentioned. Comparisons with VASO are outdated and incorrect.
The reproducibility of the methods and the result is doubtful (see below).
I don't think that this manuscript is in the top 50% of the 240 layer-fmri papers out there.
3T layer-fMRI papers that are not cited:<br /> Taso, M., Munsch, F., Zhao, L., Alsop, D.C., 2021. Regional and depth-dependence of cortical blood-flow assessed with high-resolution Arterial Spin Labeling (ASL). Journal of Cerebral Blood Flow and Metabolism. https://doi.org/10.1177/0271678X20982382
Wu, P.Y., Chu, Y.H., Lin, J.F.L., Kuo, W.J., Lin, F.H., 2018. Feature-dependent intrinsic functional connectivity across cortical depths in the human auditory cortex. Scientific Reports 8, 1-14. https://doi.org/10.1038/s41598-018-31292-x
Lifshits, S., Tomer, O., Shamir, I., Barazany, D., Tsarfaty, G., Rosset, S., Assaf, Y., 2018. Resolution considerations in imaging of the cortical layers. NeuroImage 164, 112-120. https://doi.org/10.1016/j.neuroimage.2017.02.086
Puckett, A.M., Aquino, K.M., Robinson, P.A., Breakspear, M., Schira, M.M., 2016. The spatiotemporal hemodynamic response function for depth-dependent functional imaging of human cortex. NeuroImage 139, 240-248. https://doi.org/10.1016/j.neuroimage.2016.06.019
Olman, C.A., Inati, S., Heeger, D.J., 2007. The effect of large veins on spatial localization with GE BOLD at 3 T: Displacement, not blurring. NeuroImage 34, 1126-1135. https://doi.org/10.1016/j.neuroimage.2006.08.045
Ress, D., Glover, G.H., Liu, J., Wandell, B., 2007. Laminar profiles of functional activity in the human brain. NeuroImage 34, 74-84. https://doi.org/10.1016/j.neuroimage.2006.08.020
Huber, L., Kronbichler, L., Stirnberg, R., Ehses, P., Stocker, T., Fernández-Cabello, S., Poser, B.A., Kronbichler, M., 2023. Evaluating the capabilities and challenges of layer-fMRI VASO at 3T. Aperture Neuro 3. https://doi.org/10.52294/001c.85117
Scheeringa, R., Bonnefond, M., van Mourik, T., Jensen, O., Norris, D.G., Koopmans, P.J., 2022. Relating neural oscillations to laminar fMRI connectivity in visual cortex. Cerebral Cortex. https://doi.org/10.1093/cercor/bhac154
Strengths:
See above. The authors developed their own SMS sequence with many features. This is important to the field. And does not leave sequence development work to view isolated monopoly labs. This work democratises SMS.<br /> The questions addressed here are of high relevance to the field: getting tools with good sensitivity, user-friendly applicability, and locally specific brain activity mapping is an important topic in the field of layer-fMRI.
Weaknesses:
1. I feel the authors need to justify why flow-crushing helps localization specificity. There is an entire family of recent papers that aim to achieve higher localization specificity by doing the exact opposite. Namely, MT or ABC fRMRI aims to increase the localization specificity by highlighting the intravascular BOLD by means of suppressing non-flowing tissue. To name a few:
Priovoulos, N., de Oliveira, I.A.F., Poser, B.A., Norris, D.G., van der Zwaag, W., 2023. Combining arterial blood contrast with BOLD increases fMRI intracortical contrast. Human Brain Mapping hbm.26227. https://doi.org/10.1002/hbm.26227.
Pfaffenrot, V., Koopmans, P.J., 2022. Magnetization Transfer weighted laminar fMRI with multi-echo FLASH. NeuroImage 119725. https://doi.org/10.1016/j.neuroimage.2022.119725
Schulz, J., Fazal, Z., Metere, R., Marques, J.P., Norris, D.G., 2020. Arterial blood contrast ( ABC ) enabled by magnetization transfer ( MT ): a novel MRI technique for enhancing the measurement of brain activation changes. bioRxiv. https://doi.org/10.1101/2020.05.20.106666
Based on this literature, it seems that the proposed method will make the vein problem worse, not better. The authors could make it clearer how they reason that making GE-BOLD signals more extra-vascular weighted should help to reduce large vein effects.
The empirical evidence for the claim that flow crushing helps with the localization specificity should be made clearer. The response magnitude with and without flow crushing looks pretty much identical to me (see Fig, 6d).<br /> It's unclear to me what to look for in Fig. 5. I cannot discern any layer patterns in these maps. It's too noisy. The two maps of TE=43ms look like identical copies from each other. Maybe an editorial error?
The authors discuss bipolar crushing with respect to SE-BOLD where it has been previously applied. For SE-BOLD at UHF, a substantial portion of the vein signal comes from the intravascular compartment. So I agree that for SE-BOLD, it makes sense to crush the intravascular signal. For GE-BOLD however, this reasoning does not hold. For GE-BOLD (even at 3T), most of the vein signal comes from extravascular dephasing around large unspecific veins, and the bipolar crushing is not expected to help with this.
2. The bipolar crushing is limited to one single direction of flow. This introduces a lot of artificial variance across the cortical folding pattern. This is not mentioned in the manuscript. There is an entire family of papers that perform layer-fmri with black-blood imaging that solves this with a 3D contrast preparation (VAPER) that is applied across a longer time period, thus killing the blood signal while it flows across all directions of the vascular tree. Here, the signal cruising is happening with a 2D readout as a "snap-shot" crushing. This does not allow the blood to flow in multiple directions.<br /> VAPER also accounts for BOLD contaminations of larger draining veins by means of a tag-control sampling. The proposed approach here does not account for this contamination.
Chai, Y., Li, L., Huber, L., Poser, B.A., Bandettini, P.A., 2020. Integrated VASO and perfusion contrast: A new tool for laminar functional MRI. NeuroImage 207, 116358. https://doi.org/10.1016/j.neuroimage.2019.116358
Chai, Y., Liu, T.T., Marrett, S., Li, L., Khojandi, A., Handwerker, D.A., Alink, A., Muckli, L., Bandettini, P.A., 2021. Topographical and laminar distribution of audiovisual processing within human planum temporale. Progress in Neurobiology 102121. https://doi.org/10.1016/j.pneurobio.2021.102121
If I would recommend anyone to perform layer-fMRI with blood crushing, it seems that VAPER is the superior approach. The authors could make it clearer why users might want to use the unidirectional crushing instead.
3. The comparison with VASO is misleading.<br /> The authors claim that previous VASO approaches were limited by TRs of 8.2s. The authors might be advised to check the latest literature of the last years.<br /> Koiso et al. performed whole brain layer-fMRI VASO at 0.8mm at 3.9 seconds (with reliable activation), 2.7 seconds (with unconvincing activation pattern, though), and 2.3 (without activation).<br /> Also, whole brain layer-fMRI BOLD at 0.5mm and 0.7mm has been previously performed by the Juelich group at TRs of 3.5s (their TR definition is 'fishy' though).
Koiso, K., Müller, A.K., Akamatsu, K., Dresbach, S., Gulban, O.F., Goebel, R., Miyawaki, Y., Poser, B.A., Huber, L., 2023. Acquisition and processing methods of whole-brain layer-fMRI VASO and BOLD: The Kenshu dataset. Aperture Neuro 34. https://doi.org/10.1101/2022.08.19.504502
Yun, S.D., Pais‐Roldán, P., Palomero‐Gallagher, N., Shah, N.J., 2022. Mapping of whole‐cerebrum resting‐state networks using ultra‐high resolution acquisition protocols. Human Brain Mapping. https://doi.org/10.1002/hbm.25855
Pais-Roldan, P., Yun, S.D., Palomero-Gallagher, N., Shah, N.J., 2023. Cortical depth-dependent human fMRI of resting-state networks using EPIK. Front. Neurosci. 17, 1151544. https://doi.org/10.3389/fnins.2023.1151544
The authors are correct that VASO is not advised as a turn-key method for lower brain areas, incl. Hippocampus and subcortex. However, the authors use this word of caution that is intended for inexperienced "users" as a statement that this cannot be performed. This statement is taken out of context. This statement is not from the academic literature. It's advice for the 40+ user base that wants to perform layer-fMRI as a plug-and-play routine tool in neuroscience usage. In fact, sub-millimeter VASO is routinely being performed by MRI-physicists across all brain areas (including deep brain structures, hippocampus etc). E.g. see Koiso et al. and an overview lecture from a layer-fMRI workshop that I had recently attended: https://youtu.be/kzh-nWXd54s?si=hoIJjLLIxFUJ4g20&t=2401
Thus, the authors could embed this phrasing into the context of their own method that they are proposing in the manuscript. E.g. the authors could state whether they think that their sequence has the potential to be disseminated across sites, considering that it requires slow offline reconstruction in Matlab?<br /> Do the authors think that the results shown in Fig. 6c are suggesting turn-key acquisition of a routine mapping tool? In my humble opinion, it looks like random noise, with most of the activation outside the ROI (in white matter).
4. The repeatability of the results is questionable.<br /> The authors perform experiments about the robustness of the method (line 620). The corresponding results are not suggesting any robustness to me. In fact, the layer profiles in Fig. 4c vs. Fig 4d are completely opposite. The location of peaks turns into locations of dips and vice versa.<br /> The methods are not described in enough detail to reproduce these results.<br /> The authors mention that their image reconstruction is done "using in-house MATLAB code" (line 634). They do not post a link to github, nor do they say if they share this code.
It is not trivial to get good phase data for fMRI. The authors do not mention how they perform the respective coil-combination.<br /> No data are shared for reproduction of the analysis.
5. The application of NODRIC is not validated.<br /> Previous applications of NORDIC at 3T layer-fMRI have resulted in mixed success. When not adjusted for the right SNR regime it can result in artifactual reductions of beta scores, depending on the SNR across layers. The authors could validate their application of NORDIC and confirm that the average layer-profiles are unaffected by the application of NORDIC. Also, the NORDIC version should be explicitly mentioned in the manuscript.
Akbari, A., Gati, J.S., Zeman, P., Liem, B., Menon, R.S., 2023. Layer Dependence of Monocular and Binocular Responses in Human Ocular Dominance Columns at 7T using VASO and BOLD (preprint). Neuroscience. https://doi.org/10.1101/2023.04.06.535924
Knudsen, L., Guo, F., Huang, J., Blicher, J.U., Lund, T.E., Zhou, Y., Zhang, P., Yang, Y., 2023. The laminar pattern of proprioceptive activation in human primary motor cortex. bioRxiv. https://doi.org/10.1101/2023.10.29.564658
Yeah, same for status and same for 12 questions.
Use your 12 questions as a tag in your project.
I disagree. What is expressed is an attempt to solve X by making something that should maybe be agnostic of time asynchronous. The problem is related to design: time taints code. You have a choice: either you make the surface area of async code grow and grow or you treat it as impure code and you lift pure synchronous logic in an async context. Without more information on the surrounding algorithm, we don't know if the design decision to make SymbolTable async was the best decision and we can't propose an alternative. This question was handled superficially and carelessly by the community.
superficially and carelessly?
The problem with this pile of questions is that, instead of helping the OP get out of the X Y problem, people stay focussed on Y, mark the question as a duplicate of Y in a matter of minutes and X is never properly addressed.
sticking too much to policy/habit instead of addressing the specific needs of individuals? too much eagerness to close / mark as duplicate?
Joint Public Review:
Summary:
Cincotta et al set out to investigate the presence of glucocorticoid receptors in the male and female embryonic germline. They further investigate the impact of tissue specific genetically induced receptor absence and/or systemic receptor activation on fertility and RNA regulation. They are motivated by several lines of research that report inter and transgenerational effects of stress and or glucocorticoid receptor activation and suggest that their findings provide an explanatory mechanism to mechanistically back parental stress hormone exposure induced phenotypes in the offspring.
Strengths:
- A chronological immunofluorescent assessment of GR in fetal and early life oocyte and sperm development.<br /> - RNA seq data that reveal novel cell type specific isoforms validated by q-RT PCR E15.5 in the oocyte.<br /> - 2 alternative approaches to knock out GR to study transcriptional outcomes. Oocytes: systemic GR KO (E17.5) with low input 3-tag seq and germline specific GR KO (E15.5) on fetal oocyte expression via 10X single cell seq and 3-cap sequencing on sorted KO versus WT oocytes - both indicating little impact on polyadenylated RNAs -<br /> - 2 alternative approaches to assess the effect of GR activation in vivo (systemic) and ex vivo (ovary culture): here the RNA seq did show again some changes in germ cells and many in the soma.<br /> - They exclude oocyte specific GR signaling inhibition via beta isoforms<br /> - Perinatal male germline shows differential splicing regulation in response to systemic Dex administration, results were backed up with q-PCR analysis of splicing factors.
Weaknesses:
- Sequencing techniques used are not Total RNA but either are focused on all polyA transcripts (10x) - effects on non-polyA-transcripts are left unexplored.<br /> The number of replicates in the low input seq is very low and hence this might be underpowered. Since Dex treatment showed some (modest) changes in oocyte RNA effects of GR depletion might only become apparent upon Dex treatment as an interaction. Meaning GR activation in the presence of GR shows changes, upon GR depletion those changes are abolished --> statistically speaking an interaction --> conclusion: there are germline GR effects that get abolished when there is no GR hinting on germline GR autonomous effects.<br /> - Effects in oocytes following systemic Dex might be indirect due to GR activation in the soma. The changes observed might be irrelevant to meiosis and thus in the manuscript are deemed irrelevant, but they could still lead to settle consequences. in other terms.
Even though ex vivo culture of ovaries shows GR translocation to nucleus it is not sure whether the in vivo systemic administration does the same. The authors argue in their rebuttal that GR is already nuclear in fetal oocytes hence the<br /> conclusion that fetal oocytes are resistant to GR manipulation is understandable, at least for the readouts that were considered. Yet the question arises: If GR is already nuclear (active) in the absence of additional Dex treatment why does GR knock out not elicit any changes in the considered readouts -> what are we missing.
This work is a good reference point for researchers interested in glucocorticoid hormone signaling fertility and RNA splicing. It might spark further studies on germline-specific GR functions and the impact of GR activation on alternative splicing.<br /> The study provides a characterization of GR and some aspects of GR perturbation, and the negative findings in this study do help to rule out a range of specific roles of GR in the germline. This will help the study of unexplored options. The authors do acknowledge the unexplored options in their discussion.<br /> The intro of the study eludes to implications for intergenerational effects via epigenetic modifications in the germline and points out additional potential indirect effects of reproductive tissue GR signaling on the germline. Future studies might hence focus on further exploration of epigenetic modifications and/or indirect effects.
<body>
a semantic tag that contains what is visible to the user
Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Public Review):
This work challenges previously published results regarding the presence and abundance of 6mA in the Drosophila genome, as well as the claim that the TET or DMAD enzyme serves as the "eraser" of this DNA methylation mark and its roles in development. This information is needed to clarify these questions in the field. I am less familiar with the biochemical approaches in this work, so my comments are mainly on the genetic analyses. Generally speaking, the methods for fly husbandry and treatment seem to be in accordance with those established in the field.
Response : We thank the reviewer for his/her work and positive assessment of our manuscript.
Reviewer #2 (Public Review):
DNA adenine methylation (6mA) is a rediscovered modification that has been described in a wide range of eukaryotes. However, 6mA presence in eukaryote remains controversial due to the low abundance of its modification in eukaryotic genome. In this manuscript, Boulet et al. re-investigate 6mA presence in drosophila using axenic or conventional fly to avoid contaminants from feeding bacteria. By using these flies, they find that 6mA is rare but present in the drosophila genome by performing LC/MS/MS. They also find that the loss of TET (also known as DMAD) does not impact 6mA levels in drosophila, contrary to previous studies. In addition, the authors find that TET is required for fly development in its enzymatic activity-independent manner.
The strength of this study is, that compared to previous studies of 6mA in drosophila, the authors employed axenic or conventional fly for 6mA analysis. These fly strains make it possible to analyze 6mA presence in drosophila without bacterial contaminant. Therefore, showing data of 6mA abundance in drosophila by performing LC-MS/MS in this manuscript is more convincing as compared with previous studies. Intriguingly, the authors find that the conserved iron-binding motif required for the catalytic activity of TET is dispensable for its function. This finding could be important to reveal TET function in organisms whose genomic 5mC levels are very low.
The manuscript in this paper is well written but some aspects of data analysis and discussion need to be clarified and extended.
It is convincing that an increase in 6mA levels is not observed in TETnull presented in Fig1. But it seems 6mA levels are altered in Ax.TET1/2 compared with Ax.TETwt and Ax.TETnull presented in Fig1f (and also WT vs TET1/2 presented in Fig1g). Is it sure that no statistically significant were not observed between Ax.TET1/2 and Ax.TETwt?
The representing data of in vitro demethylation assay presented in Fig.3 is convincing, but it is not well discussed and analyzed why these results are contrary to previous reports (Yao et al., 2018 and Zhang et al., 2015).
We thank the reviewer for his/her work and positive assessment of our manuscript.
(1) We repeated our statistical analyses and confirmed that there is no significant difference between wildtype and tet1/2 mutant embryos in axenic conditions (Welch two sample t-test : p=0.075).
(2) We added some elements in the revised manuscript to discuss the possible reasons for the discrepancies with previous reports. Notably both studies performed the in vitro demethylation assays over a much longer time course and with different sources of recombinant proteins. Zhang et al. purified TET catalytic domain from human cells (HEK293T) and observed around 2.5% of 6mA demethylation at 30 min and less than 25% after 10 hours of incubation as measured by HPLC-MS/MS analyses. Yao et al. incubated recombinant TET catalytic domain with 6mA DNA for 3h and observed a 25% decrease in 6mA levels as measured by dot blot. These results suggest that drosophila TET may oxidize 6mA, but with a much lower affinity than 5mC since with observed a near complete oxidation of 5mC after 1 minute and no decrease in 6mA levels after 30 minutes of reaction (for identical concentrations of substrate and enzyme). It is possible too that the preparation of TET catalytic domain in different systems changes its enzymatic activity, potentially in relation with distinct post-translational modifications. Still, as already mentioned in our manuscript, extensive biochemical analyses of the distant TET homolog from the fungus Coprinopsis cinerea (Mu et al., Nature Chem Biol 2022) strongly argue that TET enzymes do not harbor the residues required to serve as 6mA demethylase.
Reviewer #1 (Recommendations For The Authors):
Here are one comment (#1) and a couple of questions (#2-3) that could be addressed in the future, in order to understand the roles of 6mA and TET. Even though #2 and #3 are likely beyond the scope of this paper, #1 should be addressed within the scope of this work and compared with previous reports.
- The phenotypic analyses in Fig. 4 should use tet_null/Deficiency and tet_CD/Deficiency for their potential phenotypes. This needs to be addressed since both the tet_null and the tet_CD were generated using the same starting fly line (GFP knock-in). Using a deficiency chromosome and testing these alleles in hemizygotes would be helpful to eliminate any secondary effects due to genetic background issues.
Thanks for this comment. Actually, tet_null and tet_CD were not generated using the same starting lines. Whereas tet_cd was generated (by CRISPR) using the tet-GFP knock-in line, tet_null was generated by FRT site recombination between two PBac insertions (Delatte et al. 2016). As for tet1 and tet2 (used in allelic combination in Fig 4 J-L), they correspond to two distinct mutant alleles generated by CRISPR (Zhang et al. 2015). We have clarified this in the M&M (page 9).
- Regarding the estimated "200 to 400 methylated adenines per haplogenome", is there any insight into where are they located in the genome?
It is an interesting question and we initially used SMRT-seq sequencing to obtain this kind of information. As it turned out that this technique gives a high level of false positive, we should consider with caution the interpretation of these data and we decided not to include them in the manuscript. Still, we characterized the genomic features of the 6mA detected using stringent criteria (mQV>100, cov>25x in the fusion dataset and triplicated across samples of the same genotype). Both in wild type and tet_null, 6mA were dispersed along each chromosome although few of them were found on chromosome X. In both cases there appeared to be a higher accumulation of 6mAs on the histone locus and the transposon-rich tip of chromosome X, but 6mA density remained below 1.3/kb in other genomic regions. Comparisons with annotated genomic regions indicated that 6mA were enriched in long interspersed nuclear elements (LINEs) and satellite repeats, and depleted in 3’UTR and exons, but there was no significant difference in their repartition between the two genetic contexts. Besides, motif analyses showed similar enrichments in both conditions, with GAG triplet accounting for more than one quarter of all the sites. Whether this reflects the specificity of a putative adenine methylase or a technical bias associated the with SMTR-seq technology remains to be established.
- The TET-GFP and TET-CD-GFP knock-in lines give proper nuclear localization and could be used to identify genomic regions bound with full-length TET and TET-CD using anti-GFP for ChIP-seq or CUT&RUN (or CUT&TAG).
Indeed, this is a line of research that we are following up and will be part of another study. Actually, our ChIP-seq experiments indicate that they bind on the same genomic regions.
Reviewer #2 (Recommendations For The Authors):
- I think the major findings of this paper are showing 6mA present in drosophila by using xenic or conventional breeding conditions and finding that TET function independently of its catalytic activity is essential for fly development. The authors could have been more precise in title and abstract to emphasize these findings.
We have now modified the abstract to try to emphasize these findings.
- The authors claim that any increase of 6mA levels was not observed in both TETnull and TET1/2, but it is not sufficiently convincing. Because it seems 6mA levels were increased in Ax. tet1/2 embryo as compared with in Ax.wt embryo (Fig.1). In this scenario, 6mA abundance in both TETnull and TET1/2 mutant are supposed to be the same. It would be better to re-analyze data carefully and discuss if 6mA levels were significantly increased in TET1/2, and why 6mA levels are different between TETnull and TET1/2. Additionally, the authors describe that the TET null mutant is pupal lethal, while the TET1/2 survivor is available. The text suggests that TET1/2 could have partial functionality on fly development (Fig.4). It would be better to check whether the N-terminus of TET is expressed in the TET1/2 mutant.
Indeed, the increase in 6mA levels in Ax. tet1/2 embryo seems consequent (although it is not statistically significant) and no increase was observed in Ax tet_null embryos. Thus, the putative effect on 6mA levels in tet1/2 embryos may not be directly due to the absence of TET function. We now mention in the revised manuscript (page 6) that “the apparent increase in 6mA levels in tet1/2 axenic embryos was not reproduced in tet_null embryos, suggesting that it does not simply reflect the tet loss of function, and that it was not statistically significant”. Besides, we do not have an antibody to check whether the N-terminus of TET is expressed in the tet1/2 mutants, but the western blot published by Zhang et al 2015 shows that tet2 mutation leads to the expression of TET N-terminal domain. This N-terminal domain could have partial TET functionality and/or interfere with the function of other factors (notably those implicated in 6mA metabolism).
- The authors show that SMRT-seq data did not reveal an increase in 6mA levels in loss of TET (Fig.2). It is convincing that total 6mA abundance was not altered by loss of TET. But were 6mA-accumulated locus/regions observed in WT not altered by loss of TET?
Please refer to our answer to reviewer 1 on that point.
- It remains unclear that the TET proteins the authors prepared do not exhibit 6mA demethylate activity in vitro, contrary to what was reported in previous papers (Fig.3). I think the preparation of recombinant proteins may make different results between this and previous papers. Yao et al., 2018 and Zhang et al., 2015 used recombinant proteins purified from Human cells or insect cells, while the author purified them from E.Coli. Additionally, it's mentioned that VK Rao et al., 2020 demonstrated cdk5-mediated phosphorylation of Tet3 increases its in catalytic activity in vitro. These previous reports suggest modification of TET could change demethylase activity. More analysis and discussion are needed to support the conclusion.
Thanks for your insights. This in an important point and we added the following elements in the revised manuscript to discuss possible reasons for the discrepancies with previous reports (pages 7-8): “Our results contrast with previous reports showing that recombinant drosophila TET demethylates 6mA on dsDNA in vitro (Yao et al. 2018; Zhang et al., 2015a). However, both studies ran much longer reactions (up to 10 hours) and used different sources of recombinant protein (drosophila TET catalytic domain purified from human HEK293T cells). Notably, Zhang et al. (2015a) only found around 2.5% of 6mA demethylation at 30 min and less than 25% after 10 hours of incubation as measured by HPLC-MS/MS analyses. These results suggest that drosophila TET may oxidize 6mA, but with a much lower affinity than 5mC since with observed a near complete oxidation of 5mC after 1 min. and no significant decrease in 6mA levels after 30 min. of reaction (for identical concentrations of substrate and enzyme). It is possible too that the preparation of TET catalytic domain in different systems changes its enzymatic activity, potentially in relation to distinct post-translational modifications.”
Reviewer #1 (Public Review)
This work challenges previously published results regarding the presence and abundance of 6mA in Drosophila genome, as well as the claim that the TET or DMAD enzyme serves as the "eraser" of this DNA methylation mark and its roles in development. This information is needed to clarify these questions in the field. Generally speaking, the methods for fly husbandry and treatment seem to be in accordance with those established ones in the field.
Here are a couple of suggestions that could be discussed with the current work and addressed in the future, in order to better understand the roles of 6mA and TET.
1. Regarding the estimated "200 to 400 methylated adenines per haplogenome", some insights regarding where they are enriched in the genome could inform the potential target sites regulated by 6mA.
2. The TET-GFP and TET-CD-GFP knock-in lines give proper nuclear localization and could be used to identify genomic regions bound with full-length TET and TET-CD using anti-GFP for ChIP-seq or CUT&RUN (or CUT&TAG).
negative effects of lost instructional time for those students who were suspended and positive effects of reduced number of disruptive peers in the classroom for students who were not.
this creates a gap between the class, some people are way above the mark, while others can barely tag along in class
Reviewer #1 (Public Review):
In this study, the authors examined the role of IBTK, a substrate-binding adaptor of the CRL3 ubiquitin ligase complex, in modulating the activity of the eiF4F translation initiation complex. They find that IBTK mediates the non-degradative ubiquitination of eiF4A1, promotes cap-dependent translational initiation, nascent protein synthesis, oncogene expression, and tumor cell growth. Correspondingly, phosphorylation of IBTK by mTORC1/ S6K1 increases eIF4A1 ubiquitination and sustains oncogenic translation.
Strengths:
This study utilizes multiple biochemical, proteomic, functional, and cell biology assays to substantiate their results. Importantly, the work nominates IBTK as a unique substrate of mTORC1, and further validates eiF4A1 ( a crucial subunit of the ei44F complex) as a promising therapeutic target in cancer. Since IBTK interacts broadly with multiple members of the translational initial complex - it will be interesting to examine its role in eiF2alpha-mediated ER stress as well as eiF3-mediated translation. Additionally, since IBTK exerts pro-survival effects in multiple cell types, it will be of relevance to characterize the role of IBTK in mediating increased mTORC1 mediated translation in other tumor types, thus potentially impacting their treatment with eiF4F inhibitors.
Limitations/Weaknesses:
The findings are mostly well supported by data, but some areas need clarification and could potentially be enhanced with further experiments:
1) Since eiF4A1 appears to function downstream of IBTK1, can the effects of IBTK1 KO/KD in reducing puromycin incorporation (in Fig 3A), cap-dependent luciferase reporter activity (Fig 3G), reduced oncogene expression ( Fig 4A) or 2D growth/ invasion assays (Fig 4) be overcome or bypassed by overexpressing eiF4A1? These could potentially be tested in future studies.
2) The decrease in nascent protein synthesis in puromycin incorporation assays in Figure 3A suggest that the effects of IBTK KO are comparable to and additive with silvesterol. It would be of interest to examine whether silvesterol decreases nascent protein synthesis or increases stress granules in the IBTK KO cells stably expressing IBTK as well.
3) The data presented in Figure 5 regarding the role of mTORC1 in IBTK-mediated eiF4A1 ubiquitination needs further clarification on several points:
- It is not clear if the experiments in Figure 5F with Phos-tag gels are using the FLAG-IBTK deletion mutant or the peptide containing the mTOR sites as it is mentioned on line 517, page 19 "To do so, we generated an IBTK deletion mutant (900-1150 aa) spanning the potential mTORC1-regulated phosphorylation sites" This needs further clarification.
-It may be of benefit to repeat the Phos tag experiments with full-length FLAG-IBTK and/or endogenous IBTK with molecular weight markers indicating the size of migrated bands.
-Additionally, torin or Lambda phosphatase treatment may be used to confirm the specificity of the band in separate experiments.
-Phos-tag gels with the IBTK CRISPR KO line would also help confirm that the non-phosphorylated band is indeed IBTK.
-It is unclear why the lower, phosphorylated bands seem to be increasing (rather than decreasing) with AA starvation/ Rapa in Fig 5H.
Evernote 的筆記標籤、連結等無法直接轉移,需要一邊執行一邊重新建立。
WAH? Evernote tags cannot be imported into Upnote??? Seriously?
This video contradicts and mentions tags can be imported:
後記:根據影片說法,這是新功能,所以電腦玩物當初可能不知道。
the new ability to import tags from Evernote I get a lot of questions about importing tags from Evernote
Awesome! This is so important. I wonder why Esor in his intro to Upnote said Evernote tags can't be imported to Upnote.
Evernote標籤可匯入Upnote。(牴觸Esor電腦玩物的說法)
Reviewer #1 (Public Review):
This is a review of the manuscript entitled "Pharmacologic hyperstabilisation of the HIV-1 capsid lattice induces capsid failure" by Faysal et al., in this manuscript the authors used an elegant single virion fluorescence assay based on TIRF to measure the stability of mature HIV cores. Virions were biotinylated and captured onto glass coverslips through specific Biotin-Avidin interactions. Immobilized virions were then introduced to the imaging buffer which contained the pore-forming protein DLY, and fluorescently labeled CypA. Mature virions were identified through the binding of CypA which had a red fluorescent tag allowing them to measure the dynamics of GFP trapped within the mature cores as well as the CypA bound outside the core. The authors show that the addition of LEN starting from about 50nM stabilized the mature cores even after cores have ruptured and released their internal GFP. Higher concentration of Len results in ultrastabilization of the cores and rapid rupture leading to the release of GFP at an earlier timepoint. A biochemical assembly assay was performed which showed uM quantities of Len synergized with IP6 to promote CA assembly. Purified mature virions were also treated with 700nM of Len and analyzed by CryoET, this analysis showed an increased representation of irregular cores within the Len-treated sample. Putting all of this together, the authors concluded that Len facilitates core rupture through hyperstabilization of HIV cores, as described in the title.
While I have found this work technically well performed and well explained, I do not believe that the presented data supports the conclusions reached by the authors.
the robots meta tag instructs search engines not to show the page in search results
Remove from search engine results
Author Response
The following is the authors’ response to the original reviews.
eLife assessment
This study reports important findings regarding the systemic function of hemocytes controlling whole-body responses to oxidative stress. The evidence in support of the requirement for hemocytes in oxidative stress responses as well as the hemocyte single-nuclei analyses in the presence or absence of oxidative stress are convincing. In contrast, the genetic and physiological analyses that link the non-canonical DDR pathway to upd3/JNK expression and high susceptibility, and the inferences regarding the function of hemocytes in systemic metabolic control are incomplete and would benefit from more rigorous approaches. The work will be of interest to cell and developmental biologists working on animal metabolism, immunity, or stress responses.
We would like to thank the editorial team for these positive comments on our manuscript and the constructive suggestions to improve our manuscript. We are now happy to send you our revised manuscript, which we improved according to the suggestions and valuable comments of the referees.
Public Reviews:
Reviewer #1 (Public Review):
The study examines how hemocytes control whole-body responses to oxidative stress. Using single cell sequencing they identify several transcriptionally distinct populations of hemocytes, including one subset that show altered immune and stress gene expression. They also find that knockdown of DNA Damage Response (DDR) genes in hemocytes increases expression of the immune cytokine, upd3, and that both upd3 overexpression in hemocytes and hemocyte knockdown of DDR genes leads to increased lethality upon oxidative stress.
Strengths
The single cell analyses provide a clear description of how oxidative stress can cause distinct transcriptional changes in different populations of hemocytes. These results add to the emerging them in the field that there functionally different subpopulations of hemocytes that can control organismal responses to stress.
The discovery that DDR genes are required upon oxidative stress to limit cytokine production and lethality provides interesting new insight into the DDR may play non-canonical roles in controlling organismal responses to stress.
We are grateful to referee 1 to point out the importance and novelty of our snRNA-seq data and our findings on the role of DNA damage-modulated cytokine release by hemocytes during oxidative stress. We further extended these analyses in the revised manuscript by looking deeper into the transcriptomic alterations in fat body cells upon oxidative stress (Figure 4, Figure S4). We further provide additional data to support the connection of DNA damage signaling and regulation of upd3 release from hemocytes (Figure 6F). Here we show that upd3-deficiency can abrogate the increased susceptibility of flies with mei41 and tefu knockdown in hemocytes. In line with this finding, we also show that upd3null mutants show a reduced but not abolished susceptibility to oxidative stress overall (Figure 6F), underlining the role of upd3 as a mediator of oxidative stress response.
Weaknesses
- In some ways the authors interpretation of the data - as indicated, for example, in the title, summary and model figure - don't quite match their data. From the title and model figure, it seems that the authors suggest that the DDR pathway induces JNK and Upd3 and that the upd3 leads to tissue wasting. However, the data suggest that the DDR actually limits upd3 production and susceptibility to death as suggested by several results:
According to the referee’s suggestion, we revised the manuscript and adjusted our title, abstract and graphical summary to be more precise that DNA damage signaling seem to have a modulatory or regulatory effect on upd3 release. Furthermore, we provide now additional data to support the connection between DNA damage signaling and upd3 release. For example, we added several genetic “rescue” experiments to strengthen the epistasis that modulation of DNA damage signaling and the higher susceptibility of the fly is connected to altered upd3 levels (Figure 6F). We now provide additional data showing that the loss of upd3 rescues the susceptibility to oxidative stress in flies, which are deficient for DDR components in hemocytes.
a. PQ normally doesn't induce upd3 but does lead to glycogen and TAG loss, suggesting that upd3 isn't connected to the PQ-induced wasting.
Even though in our systemic gene expression analysis of upd3 expression, we could not detect a significant induction of upd3 upon PQ feeding. However, we found upd3 expression within our snRNAseq data in a distinct cluster of immune-activated hemocytes (Figure 3B, Cluster 6). Upon knockdown of the DNA damage signaling in hemocytes, the levels then increase to a detectable level in the whole fly. This supports our assumption that upd3 is needed upon oxidative stress to induce energy mobilization from the fat body, but needs to be tightly controlled to balance tissue wasting for energy mobilization. Furthermore, we found evidence in our new analysis of the snRNA-seq data of the fat body cells, that indeed we can find Jak/STAT activation in one cell cluster here, which could speak for an interaction of Cluster 6 hemocytes with cluster 6 fat body cells. A hypothesis we aim to explore in future studies.
b. knockdown of DDR upregulates upd3 and leads to increased PQ-induced death. This would suggest that activation of DDR is normally required to limit, rather than serve as the trigger for upd3 production and death.
Our data support the hypothesis that DDR signaling in hemocytes “modulates” upd3 levels upon oxidative stress. We now carefully revised the text and the graphical summary of the manuscript to emphasize that oxidative stress causes DNA damage, which subsequently induces the DNA damage signaling machinery. If this machinery is not sufficiently induced, for example by knockdown of tefu and mei-41, non-canonical DNA damage signaling is altered which induces JNK signaling and induces release of pro-inflammatory cytokines, including upd3. Whereas DNA damage itself is only slightly increase in the used DDR deficient lines (Figure 5C) and hemocytes do not undergo apoptosis (unaltered cell number on PQ (Figure 5B)), we conclude that loss of tefu, mei-41, or nbs1 causes dysregulation of inflammatory signaling cascades via non-canonical DNA damage signaling. However, oxidative stress itself seems to also induce upd3 release and DNA damage signaling in the same cell cluster, as shown by our snRNA-seq data (Figure 3B). Hence, we think that DNA damage signaling is needed as a rate-limiting step for upd3 release.
c. hemocyte knockdown of either JNK activity or upd3 doesn't affect PQ-induced death, suggesting that they don't contribute to oxidative stress-induced death. It’s only when DDR is impaired (with DDR gene knockdown) that an increase in upd3 is seen (although no experiments addressed whether JNK was activated or involved in this induction of upd3), suggesting that DDR activation prevents upd3 induction upon oxidative stress.
Whereas the double knockdown of upd3 or bsk and DDR genes was resulting in insufficient knockdown efficiencies, we added a rescue experiment where we combined upd3null mutants with knockdown of tefu and mei-41 in hemocytes and found a reduced susceptibility of DDR-deficient flies to oxidative stress.
- The connections between DDR, JNK and upd3 aren't fully developed. The experiments show that susceptibility to oxidative stress-induced death can be caused by a) knockdown of DDR genes, b) genetic overexpression of upd3, c) genetic activation of JNK. But whether these effects are all related and reflect a linear pathway requires a little more work. For example, one prediction of the proposed model is that the increased susceptibility to oxidative stress-induced death in the hemocyte DDR gene knockdowns would be suppressed (perhaps partially) by simultaneous knockdown of upd3 and/or JNK. These types of epistasis experiments would strengthen the model and the paper.
As mentioned before, we had some technical difficulties combining the knockdown of bsk or upd3 with DDR genes. However, we added a new experiment in which we show that upd3null mutation can rescue the higher susceptibility of hemocytes with tefu and mei41 knockdown.
- The (potential) connections between DDR/JNK/UPD3 and the oxidative stress effects on depletion of nutrient (lipids and glycogen) stores was also not fully developed. However, it may be the case that, in this paper, the authors just want to speculate that the effects of hemocyte DDR/upd3 manipulation on viability upon oxidative stress involve changes in nutrient stores.
In the revised version of the manuscript, we now provide a more thorough snRNA-seq analysis in the fat body upon PQ treatment to give more insights on the changes in the fat body upon PQ treatment. We added additional histological images of the abdominal fat body on control food and PQ food, to demonstrate the elimination of triglycerides from fat body with Oil-Red-O staining (Figure S1). We also analyzed now hemocyte-deficient (crq-Gal80ts>reaper) flies for their levels of triglycerides and carbohydrates during oxidative stress, to support our hypothesis that hemocytes are key players in the regulation of energy mobilization during oxidative stress. Loss of hemocytes (and therefore also their regulatory input on energy mobilization from the fat body) results in increased triglyceride storage in the fat body during steady state with a decreased consumption of these triglycerides on PQ food compared to control flies (Figure 1J). In contrast, glycogen storage and mobilization, which is mostly done in muscle, is not altered in these flies during oxidative stress (Figure 1L). Interestingly, free glucose levels are drastically reduced in hemocyte-deficient flies, which could be due to insufficient energy mobilization from the fat body and subsequently results in a higher susceptibility of these flies on oxidative stress (Figure 1K). Additionally, we aim to point out here that “functional” hemocytes are needed for effective response to oxidative stress, but this response has to be tightly balanced (see also new graphical abstract).
Reviewer #2 (Public Review):
Hersperger et al. investigated the importance of Drosophila immune cells, called hemocytes, in the response to oxidative stress in adult flies. They found that hemocytes are essential in this response, and using state-of-the-art single-cell transcriptomics, they identified expression changes at the level of individual hemocytes. This allowed them to cluster hemocytes into subgroups with different responses, which certainly represents very valuable work. One of the clusters appears to respond directly to oxidative stress and shows a very specific expression response that could be related to the observed systemic metabolic changes and energy mobilization. However, the association of these transcriptional changes in hemocytes with metabolic changes is not well established in this work. Using hemocyte-specific genetic manipulation, the authors convincingly show that the DNA damage response in hemocytes regulates JNK activity and subsequent expression of the JAK/STAT ligand Upd3. Silencing of the DNA damage response or excessive activation of JNK and Upd3 leads to increased susceptibility to oxidative stress. This nicely demonstrates the importance of tight control of JNK-Upd3 signaling in hemocytes during oxidative stress. However, it would have been nice to show here a link to systemic metabolic changes, as the authors conclude that it is tissue wasting caused by excessive Upd3 activation that leads to increased susceptibility, but metabolic changes were not analyzed in the manipulated flies.
We thank the referee for the suggestion to better connect upd3 cytokine levels to energy mobilization from the fat body. We agree that this is an important point to support our hypothesis. First, we added now a detailed analysis of fat body cells in our snRNA-seq data to evaluate the changes induced in the fat body upon oxidative stress. We further added additional metabolic analyses of hemocyte-deficient flies (crq-Gal80ts>reaper) to support our hypothesis that hemocytes are key players in the regulation of energy mobilization during oxidative stress (see also answer to referee 1). Loss of the regulatory role of hemocytes in the energy mobilization and redistribution leads to a decreased consumption of these triglycerides on PQ food compared to control flies (Figure 1J). In contrast, glycogen storage and mobilization from muscle, is not affected in hemocyte-deficient flies during oxidative stress (Figure 1L). Interestingly, free glucose levels are drastically reduced in hemocyte-deficient flies compared to controls, which could be due to insufficient energy mobilization from the fat body resulting in a higher susceptibility to oxidative stress (Figure 1K). This data supports our assumption that “functional” hemocytes are needed for effective response to oxidative stress, but this response has to be tightly balanced (see also new graphical summary).
The overall conclusion of this work, as presented by the authors, is that Upd3 expression in hemocytes under oxidative stress leads to tissue wasting, whereas in fact it has been shown that excessive hemocyte-specific Upd3 activation leads to increased susceptibility to oxidative stress (whether due to increased tissue wasting remains a question). The DNA damage response ensures tight control of JNK-Upd3, which is important. However, what role naturally occurring Upd3 expression plays in a single hemocyte cluster during oxidative stress has not been tested. What if the energy mobilization induced by this naturally occurring Upd3 expression during oxidative stress is actually beneficial, as the authors themselves state in the abstract - for potential tissue repair? It would have been useful to clarify in the manuscript that the observed pathological effects are due to overactivation of Upd3 (an important finding), but this does not necessarily mean that the observed expression of Upd3 in one cluster of hemocytes causes the pathology.
We agree with the referee that the pathological effects and increased susceptibility to oxidative stress are mediated by over-activated hemocytes and enhanced cytokine release, including upd3 during oxidative stress. We edited the revised manuscript accordingly to imply a “regulatory” role of upd3, which we suspect and suggest as an important mediator for inter-organ communication between hemocytes and fat body. Whereas our used model for oxidative stress (15mM Paraquat feeding) is a severe insult from which most of the flies will not recover, we could not account and test how upd3 might influence tissue repair after injury, insults and infection. We believe that this is an important factor, we aim to explore in future studies.
Reviewer #3 (Public Review):
In this study, Kierdorf and colleagues investigated the function of hemocytes in oxidative stress response and found that non-canonical DNA damage response (DDR) is critical for controlling JNK activity and the expression of cytokine unpaired3. Hemocyte-mediated expression of upd3 and JNK determines the susceptibility to oxidative stress and systemic energy metabolism required for animal survival, suggesting a new role for hemocytes in the direct mediation of stress response and animal survival.
Strength of the study:
This study demonstrates the role of hemocytes in oxidative stress response in adults and provides novel insights into hemocytes in systemic stress response and animal homeostasis.
The single-cell transcriptome profiling of adult hemocytes during Paraquat treatment, compared to controls, would be of broad interest to scientists in the field.
We are grateful to these positive comments on our data and are excited that the referee pointed out the importance of our provided snRNA-seq analysis of hemocytes and other cell types during oxidative stress. In the revised, version we now extended this analysis and looked not only into hemocytes but also highlighted induced changes in the fat body (Figure 4).
Weakness of the study:
- The authors claim that the non-canonical DNA damage response mechanism in hemocytes controls the susceptibility of animals through JNK and upd3 expression. However, the link between DDR-JNK/upd3 in oxidative stress response is incomplete and some of the descriptions do not match their data.
In the revised manuscript, we aimed to strengthen the weaknesses pointed out by the referee. We now included additional genetic crosses to validate the connection of DDR signaling in hemocytes with upd3 release. For example, we added now survival studies where we show that upd3null mutation can rescue the higher susceptibility of flies with tefu and mei41 knockdown in hemocytes during oxidative stress. Furthermore, we added additional data to highlight the importance of hemocytes themselves as essential regulators of susceptibility to oxidative stress. We analyzed the hemocyte-deficient flies (crq-Gal80ts>reaper) for their triglyceride content and carbohydrate levels during oxidative stress (Figure 1 I-L). As outlined above, loss of hemocytes leads to a decreased consumption of these triglycerides on PQ food compared to control flies (Figure 1J). In contrast, glycogen storage and mobilization from muscle, is not affected in hemocyte-deficient flies during oxidative stress (Figure 1L). Interestingly, free glucose levels are drastically reduced in hemocyte-deficient flies, which could be due to insufficient energy mobilization from the fat body resulting in a higher susceptibility to oxidative stress (Figure 1K).
- The schematic diagram does not accurately represent the authors' findings and requires further modifications.
We carefully revised the text throughout the manuscript describing our results and edited the graphical abstract to display that upd3 levels and hemocytes are essential to balance and modulate response to oxidative stress.
Reviewer #1 (Recommendations For The Authors):
The summary doesn't say too much about what the specific discoveries and results of the study are. The description is limited to just one sentence saying, "Here we describe the responses of hemocytes in adult Drosophila to oxidative stress and the essential role of non-canonical DNA damage repair activity in direct "responder" hemocytes to control JNK-mediated stress signaling, systemic levels of the cytokine upd3 and subsequently susceptibility to oxidative stress" which doesn't provide sufficient explanation of what the results were.
In the revised version of our manuscript, we now provide further information for the reader to outline the findings of our study in a concise way in the summary.
Reviewer #2 (Recommendations For The Authors):
- To strengthen the conclusion that the DDR response suppresses JNK, and thus Upd3, rescue of DDR by upd3 null mutation would help (knockdown by Hml>upd3IR might not work, RNAi seems problematic).
We would like to thank the referee for this suggestion and included now a genetic experiment where we combined upd3null mutants with hemocyte-specific knockdown of mei-41 and tefu to test their susceptibility to oxidative stress. Our data indeed provide evidence that loss of upd3 rescues the higher susceptibility of flies with hemocyte-specific knockdown for tefu and mei-41 (Figure 6F). Furthermore, we see that upd3null mutants show a diminished susceptibility to oxidative stress compared to control flies (Figure 6F).
- To link the observed effects to systemic metabolic changes, it would be useful to measure glycogen and triglycerides in these flies as well:
crq-Gal80ts>reaper to see what role hemocytes play in the observed metabolic changes.
Hml-Upd3 overexpression and Upd3 null mutant (Upd3 RNAi seems to be problematic, we have similar experiences) to see if Upd3 overexpression leads to even more profound changes as suggested, and if Upd3 mutation at least partially suppresses the observed changes.
We agree with the referee that analyzing the connection of hemocyte activation to metabolic changes should be demonstrated in our manuscript to support our claim that hemocytes are important regulators of energy mobilization during oxidative stress. Hence, we analyzed triglycerides and carbohydrate levels in hemocyte-deficient flies (crq-Gal80ts>reaper) during oxidative stress. Indeed, we found substantial differences in energy mobilization in these flies supporting the assumption that the higher susceptibility of hemocyte-deficient flies could be caused by substantial decrease in free glucose and inefficient lysis of triglycerides from the fat body (Figure 1I-K).
- To test whether the cause of the increased susceptibility to oxidative stress is due to Upd3 overactivation induced by DDR silencing, the authors should attempt to rescue DDR silencing with an Upd3 null mutation.
The suggestion of the reviewer was included in the revised manuscript and as outlined above we now added this data set to our manuscript (Figure 6F). Indeed, we can now provide evidence that upd3null mutation rescues the higher susceptibility of flies with DDR knockdown in hemocytes.
- Lethality after PQ treatment varies widely (sometimes from 10 to 90%! as in Figure 5D) - is this normal? In some experiments the variability was much lower. In particular, Figure 5D is very problematic and for example the result with upd3 null mutant compared to control is not very convincing. This could be an important result to test whether Upd3, with normal expression likely coming from cluster 6, actually plays a beneficial role, whereas overexpression with Hml leads to pathology.
We agree with the referee that it would be more convincing if the variation cross of survival experiments would be less. However, we included a lot of flies and vials in many individual experiments to test our hypothesis and variation in these survivals was always the case. These effects can be caused by many factors for example the amount of food intake by the flies, genetic background or inserted transgenes. The n-number is quite high across our survivals; so that we are convinced, the seen effects are valid. This reflects also the power of using Drosophila melanogaster as a model organism for such survivals. The high n-number in our data falls into a normal Gauss distribution with a distinct mean susceptibility between the genotypes analyzed.
- I like the conclusion at the end of the results: line 413: "We show that this oxidative stressmediated immune activation seems to be controlled by non-canonical DNA damage signaling resulting in JNK activation and subsequent upd3 expression, which can render the adult fly more susceptible to oxidative stress when it is over-activated." This is actually a more appropriate conclusion, but in the summary, introduction and discussion along with the overall schematic illustration, this is not actually stated as such, but rather as Upd3 released from cluster 6 causes the pathology. For example: line 435 "Hence, we postulate that hemocyte-derived upd3, most likely released by the activated plasmatocyte cluster C6 during oxidative stress in vivo and subsequently controlling energy mobilization and subsequent tissue wasting upon oxidative stress."
We thank the referee for this suggestion and edited our manuscript and conclusions accordingly.
Reviewer #3 (Recommendations For The Authors):
- In Figure 2, the authors claim showed that PQ treatment changes the hemocyte clusters in a way that suppresses the conventional Hml+ or Pxn+ hemocytes (cluster1) while expanding hemocyte clusters enriched with metabolic genes such as Lpin, bmm etc. It is not clear whether these cells are comparable to the fat body and if these clusters express any of previously known hemocyte marker genes to claim that these are bona fide hemocytes.
We now included a new analysis of our snRNA-seq data in Figure S4, where we clearly show that all identified hemocyte clusters do not have a fat body signature and are hemocytes, which seem to undergo metabolic adaptations (Figure S4A). Furthermore, we show that the identified fat body cells have a clear fat body signature (Figure S4B) and do not express specific hemocyte markers (Figure S4C).
- In Figure 4C, the authors showed that comet assays of isolated hemocytes result in a statistically significant increase in DNA damage in DDR-deficient flies before and after PQ treatment. However, the authors conclude that, in lines 324-328, the higher susceptibility of DDR-deficient flies is not due to an increase in DNA damage. To explicitly conclude that "non-canonical" DNA damage response, without any DNA damage, is specifically upregulated during PQ treatment, the authors require further support to exclude the potential activation of canonical DDR.
The referee is correct that we do not provide direct evidence for non-canonical DNA damage signaling. Therefore, we also decided to tune down our statement here a bit and removed that claim from the title. Increase in DNA damage can of course also increase the non-canonical DNA damage signaling pathway, loss of DNA damage signaling genes such as tefu and mei-41 seem to only have minor impacts on the overall amount of DNA damage acquired in hemocytes by oxidative stress. We therefore concluded that the induction in immune activation is most unlikely only caused by increased DNA damage but might be connected to dysregulation in non-canonical DNA damage signaling. Canonical DNA damage signaling leads essentially to DDR, which could be slow in adult hemocytes because they post-mitotic, or to apoptosis, which we could not observe in the analyzed time window in our experiments. Hemocyte number remained stable over the 24h PQ treatment without reduction in cell number (Figure 1H).
- From Figure 4D-F, the authors showed that loss of DDR in hemocytes induces the expression of unpaired 2 and 3, Socs36E, which represent the JAK/STAT pathway, and thor, InR, Pepck in the InR pathway, and a JNK readout, puc. These results indicate that the DDR pathway normally inhibits the upd-mediated JAK/STAT activation upon PQ treatment, compared to wild-type animals during PQ treatment in Figure 1B-C, which in turn protects the animal during oxidative stress responses. However, the authors claim that "enhanced DNA damage boosts immune activation and therefore susceptibility to oxidative stress (lines 365-366); we show that this oxidative stress-mediated immune activation seems to be controlled by non-canonical DNA damage signaling resulting in JNK activation and subsequent upd3 expression (line 413-416)". These conclusions are not compatible with the authors' data and may require additional data to support or can be modified.
In the revised manuscript, we carefully revised now the text and our statements that it seems that DNA damage signaling in hemocytes has regulatory or modulatory effect on the immune response during oxidative stress. Accordingly, we also adjusted our graphical summary. We agree with the referee and used the term “non-canonical” DNA damage signaling more carefully throughout the manuscript. The slight increase in DNA damage seen after PQ treatment can contribute to immune activation but seems to be not correlative to the induced cytokine levels or the susceptibility of the flies to oxidative stress.
- In Fig 1I, the authors showed that genetic ablation of hemocytes using UAS-repear induces susceptibility to PQ treatment. It is possible that inducing cell death in hemocytes itself causes the expression of cytokine upd3 or activates the JNK pathway to enhance the basal level of upd3/JNK even without PQ treatment. If this phenotype is solely mediated by the loss of hemocytes, the results should be repeated by reducing the number of hemocytes with alternative genetic backgrounds.
In the different genotypes analyzed across our manuscript we did not detect cell death of hemocytes or a dramatic reduction in hemocytes number (see Figure 1H, Figure 5B, Figure 6C). The higher susceptibility if hemocyte-deficient flies during oxidative stress is most likely caused by the loss of their regulatory role during energy mobilization. We tested triglyceride levels in hemocyte-deficient flies and found a decreased triglyceride consumption (lipolysis), with reduced levels of circulating glucose levels. This findings support our hypothesis that hemocytes are needed to balance the response to oxidative stress. In contrast, the flies with DDR-deficient hemocytes show higher systemic cytokine levels, which most likely enhance energy mobilization from the fat body and therefore result in a higher susceptibility of the fly to oxidative stress. Hence, we claim that hemocytes and their regulation of systemic cytokine levels are important to balance the response to oxidative stress and guarantee the survival of the organism.
- Lethality of control animals in PQ treatment is variable and it is hard to estimate the effect of animal susceptibility during 15mM PQ feeding. For example, Fig1A shows that control animals exhibit ~10% death during 15mM PQ which is further enhanced by crq-Gal80>reaper expression to 40% (Fig 1I). However, in Fig 5D-E, the basal lethality of wild-type controls already reaches 40~50%, which makes them hard to compare with other genetic manipulations. Related to this, the authors demonstrated that the expression of upd3 in hemocytes is sufficient to aggravate animal survival upon PQ treatment; however, upd3 null mutants do not rescue the lethality, which indicates that upd3 is not required for hampering animal mortality. These data need to be revisited and analyzed.
As outlined above, we find the variability of susceptibility to oxidative stress across all of our experiments. This could be due to different effects such as food intake but also transgene insertion and genetic background. Crq-gal80ts>reaper flies are healthy, but show a shortened life span on normal food (Kierdorf et al., 2020) due to enhanced loss of proteostasis in muscles. We show in the revised manuscript that these flies have a higher susceptibility to oxidative stress and that this effect could be mediated by defects in energy mobilization and redistribution as shown by less triglyceride lysis from the fat body and decreasing levels in free glucose. This would explain the high mortality rate of these flies at 7 days after eclosion. Paraquat treatment (15mM) is a severe inducer of oxidative stress, which results in death of most flies when they are maintained for longer time windows on PQ food. Hence, it is a model, which is not suitable to examine and monitor recovery from this detrimental insult. upd3null mutants were extensively reexamined in this manuscript, and even though we could not see a full protection of these flies from oxidative stress induced death, we found a reduced susceptibility compared to control flies (Figure 6F). Furthermore, when we combined upd3null mutants with flies deficient for tefu and mei-41 in hemocytes, the increased susceptibility to oxidative stress was rescued.
Reviewer #3 (Public Review):
Summary:<br /> This manuscript describes some biochemical experiments on the crucial virulence factor EsxA (ESAT-6) of Mycobacterium tuberculosis. EsxA is secreted via the ESX-1 secretion system. Although this system is recognized to be crucial for virulence the actual mechanisms employed by the ESX-1 substrates are still mostly unknown. The EsxA substrate is attracting the most attention as the central player in virulence, especially phagosomal membrane disruption. EsxA is secreted as a dimer together with EsxB. The authors show that EsxA is also able to form homodimers and even tetramers, albeit at very low pH (below 5). Furthermore, the addition of a nanobody that specifically binds EsxA blocks intracellular survival, as well as if the nanobody is produced in the cytosol of the infected macrophages.
Strengths:<br /> -Decent biochemical characterization of EsxA and identification of a new and interesting tool to study the function of EsxA (nanobody).
-The manuscript is well-written.
Weaknesses:<br /> The findings are not critically evaluated using extra experiments or controls.
For instance, tetrameric EsxA in itself is interesting and could reveal how EsxA works. But one would say that this is a starting point to make small point mutations that specifically affect tetramer formation and then evaluate what the effect is on phagosomal membrane lysis. Also one would like to see experiments to indicate whether these structures can be produced under in vitro conditions, especially because it seems that this mainly happens when the pH is lower than 5, which is not normally happening in phagosomes that are loaded with M. tuberculosis.
Also, the fact that the addition of the nanobody, either directly to the bacteria or produced in the cytosol of macrophages is interesting, but again it is the starting point for further experimentation. As a control, one would like to see the effect on an Esx-1 secretion mutant. Furthermore, does cytosolic production or direct addition of the nanobody affect phagosomal escape? What happens if an EsxA mutant is produced that does not bind the nanobody?
Finally, it is a bit strange that the authors use a non-native version of esxA that has not only an additional His-tag but also an additional 12 amino acids, which makes the protein in total almost 20% bigger. Of course, these additions do not have to alter the characteristics, but they might. On the other hand, they easily discard the natural acetylation of EsxA by mycobacteria itself (proven for M. marinum) as not relevant for the function because it might not happen in (the close homologue) M. tuberculosis.
Lovely. I guess what I'm trying to define is some methodology for practicing. Many times I simply resort to my exhaustive method, which has worked for me in the past simply due to brute force.Thank you for taking the time to respond and for what look like some very interesting references.
reply to u/ethanzanemiller at https://www.reddit.com/r/Zettelkasten/comments/185xmuh/comment/kb778dy/?utm_source=reddit&utm_medium=web2x&context=3
Some of your methodology will certainly depend on what questions you're asking, how well you know your area already, and where you'd like to go. If you're taking notes as part of learning a new area, they'll be different and you'll treat them differently than notes you're collecting on ideas you're actively building on or intriguing facts you're slowly accumulating. Often you'll have specific questions in mind and you'll do a literature review to see what's happing around that area and then read and take notes as a means of moving yourself closer to answering your particular questions.
Take for example, the frequently asked questions (both here in this forum and by note takers across history): how big is an idea? what is an atomic note? or even something related to the question of how small can a fact be? If this is a topic you're interested in addressing, you'll make note of it as you encounter it in various settings and see that various authors use different words to describe these ideas. Over time, you'll be able to tag them with various phrases and terminologies like "atomic notes", "one idea per card", "note size", or "note lengths". I didn't originally set out to answer these questions specifically, but my interest in the related topics across intellectual history allowed such a question to emerge from my work and my notes.
Once you've got a reasonable collection, you can then begin analyzing what various authors say about the topic. Bring them all to "terms" to ensure that they're talking about the same things and then consider what arguments they're making about the topic and write up your own ideas about what is happening to answer those questions you had. Perhaps a new thesis emerges about the idea? Some have called this process having a conversation with the texts and their authors or as Robert Hutchins called it participating in "The Great Conversation".
Almost anyone in the forum here could expound on what an "atomic note" is for a few minutes, but they're likely to barely scratch the surface beyond their own definition. Based on the notes linked above, I've probably got enough of a collection on the idea of the length of a note that I can explore it better than any other ten people here could. My notes would allow me a lot of leverage and power to create some significant subtlety and nuance on this topic. (And it helps that they're all shared publicly so you can see what I mean a bit more clearly; most peoples' notes are private/hidden, so seeing examples are scant and difficult at best.)
Some of the overall process of having and maintaining a zettelkasten for creating material is hard to physically "see". This is some of the benefit of Victor Margolin's video example of how he wrote his book on the history of design. He includes just enough that one can picture what's happening despite his not showing the deep specifics. I wrote a short piece about how I used my notes about delving into S.D. Goitein's work to write a short article a while back and looking at the article, the footnotes, and links to my original notes may be illustrative for some: https://boffosocko.com/2023/01/14/a-note-about-my-article-on-goitein-with-respect-to-zettelkasten-output-processes/. The exercise is a tedious one (though not as tedious as it was to create and hyperlink everything), but spend some time to click on each link to see the original notes and compare them with the final text. Some of the additional benefit of reading it all is that Goitein also had a zettelkasten which he used in his research and in leaving copies of it behind other researchers still actively use his translations and notes to continue on the conversation he started about the contents of the Cairo Geniza. Seeing some of his example, comparing his own notes/cards and his writings may be additionally illustrative as well, though take care as many of his notes are in multiple languages.
Another potentially useful example is this video interview with Kathleen Coleman from the Thesaurus Linguae Latinae. It's in the realm of historical linguistics and lexicography, but she describes researchers collecting masses of data (from texts, inscriptions, coins, graffiti, etc.) on cards which they can then study and arrange to write their own articles about Latin words and their use across time/history. It's an incredibly simple looking example because they're creating a "dictionary", but the work involved was painstaking historical work to be sure.
Again, when you're done, remember to go back and practice for yourself. Read. Ask questions of the texts and sources you're working with. Write them down. Allow your zettelkasten to become a ratchet for your ideas. New ideas and questions will emerge. Write them down! Follow up on them. Hunt down the answers. Make notes on others' attempts to answer similar questions. Then analyze, compare, and contrast them all to see what you might have to say on the topics. Rinse and repeat.
As a further and final (meta) example, some of my answer to your questions has been based on my own experience, but the majority of it is easy to pull up, because I can pose your questions not to my experience, but to my own zettelkasten and then quickly search and pull up a variety of examples I've collected over time. Of course I have far more experience with my own zettelkasten, so it's easier and quicker for me to query it than for you, but you'll build this facility with your own over time.
Good luck. 🗃️
Reviewer #1 (Public Review):
Summary:
This manuscript by Warfvinge et al. reports the results of CITE-seq to generate single-cell multi-omics maps from BM CD34+ and CD34+CD38- cells from nine CML patients at diagnosis. Patients were retrospectively stratified by molecular response after 12 months of TKI therapy using European Leukemia Net (ELN) recommendations. They demonstrate heterogeneity of stem and progenitor cell composition at diagnosis, and show that compared to optimal responders, patients with treatment failure after 12 months of therapy demonstrate increased frequency of molecularly defined primitive cells at diagnosis. These results were validated by deconvolution of an independent previously published dataset of bulk transcriptomes from 59 CML patients. They further applied a BCR-ABL-associated gene signature to classify primitive Lin-CD34+CD38- stem cells as BCR:ABL+ and BCR:ABL-. They identified variability in the ratio of leukemic to non-leukemic primitive cells between patients, showed differences in the expression of cell surface markers, and determined that a combination of CD26 and CD35 cell surface markers could be used to prospectively isolate the two populations. The relative proportion of CD26-CD35+ (BCR:ABL-) primitive stem cells was higher in optimal responders compared to treatment failures, both at diagnosis and following 3 months of TKI therapy.
Strengths:
The studies are carefully conducted and the results are very clearly presented. The data generated will be a valuable resource for further studies. The strengths of this study are the application of single-cell multi-omics using CITE-Seq to study individual variations in stem and progenitor clusters at diagnosis that are associated with good versus poor outcomes in response to TKI treatment. These results were confirmed by deconvolution of a historical bulk RNAseq data set. Moreover, they are also consistent with a recent report from Krishnan et al. and are a useful confirmation of those results. The major new contribution of this study is the use of gene expression profiles to distinguish BCR-ABL+ and BCR-ABL- populations within CML primitive stem cell clusters and then applying antibody-derived tag (ADT) data to define molecularly identified BCR:ABL+ and BCR-ABL- primitive cells by expression of surface markers. This approach allowed them to show an association between the ratio of BCR-ABL+ vs BCR-ABL- primitive cells and TKI response and study dynamic changes in these populations following short-term TKI treatment.
Weaknesses:
One of the limitations of the study is the small number of samples employed, which is insufficient to make associations with outcomes with confidence. Although the authors discuss the potential heterogeneity of primitive stem, they do not directly address the heterogeneity of hematopoietic potential or response to TKI treatment in the results presented. Another limitation is that the BCR-ABL + versus BCR-ABL- status of cells was not confirmed by direct sequencing for BCR-ABL. The BCR-ABL status of cells sorted based on CD26 and CD35 was evaluated in only two samples. We also note that the surface markers identified were previously reported by the same authors using different single-cell approaches, which limits the novelty of the findings. It will be important to determine whether the GEP and surface markers identified here are able to distinguish BCR-ABL+ and BCR-ABL- primitive stem cells later in the course of TKI treatment. Finally, although the authors do describe differential gene expression between CML and normal, BCR:ABL+ and BCR:ABL-, primitive stem cells they have not as yet taken the opportunity to use these findings to address questions regarding biological mechanisms related to CML LSC that impact on TKI response and outcomes.
documented evidence of oral transmission of index card use as a method
reply to u/atomicnotes at https://www.reddit.com/r/Zettelkasten/comments/1843k2w/comment/kaypbk2/?utm_source=reddit&utm_medium=web2x&context=3
I'm reasonably certain that most of the transmission of the traditions was specifically from person to person rather than from text to person. Yours is an interesting and important (and rare oral) example of person to person zettelkasten transmission, of which I've been collecting some scant examples. (Other examples appreciated, inquire within.)
Interestingly a lot of this transmission is still happening every day (though now more "visibly" online) in fora like Reddit, zettelkasten.de, Discord, in social media, and even smaller group courses. As Annie Murphy Paul indicates in The Extended Mind, people like to imitate rather than innovate. Perhaps Luhmann, being on his own outside of the establishment, was more likely to innovate because he was on his own and took Heyde's advice, but evolved it to his needs rather than asking questions on Reddit?
Reviewer #1 (Public Review):
Summary:<br /> The study by Klug et al. investigated the pathway specificity of corticostriatal projections, focusing on two cortical regions. Using a G-deleted rabies system in D1-Cre and A2a-Cre mice to retrogradely deliver channelrhodopsin to cortical inputs, the authors found that M1 and MCC inputs to direct and indirect pathway spiny projection neurons (SPNs) are both partially segregated and asymmetrically overlapping. In general, corticostriatal inputs that target indirect pathway SPNs are likely to also target direct pathway SPNs, while inputs targeting direct pathway SPNs are less likely to also target indirect pathway SPNs. Such asymmetric overlap of corticostriatal inputs has important implications for how the cortex itself may determine striatal output. Indeed, the authors provide behavioral evidence that optogenetic activation of M1 or MCC cortical neurons that send axons to either direct or indirect pathway SPNs can have opposite effects on locomotion and different effects on action sequence execution. The conclusions of this study add to our understanding of how cortical activity may influence striatal output and offer important new clues about basal ganglia function.
The conceptual conclusions of the manuscript are supported by the data, but the details of the magnitude of afferent overlap and causal role of asymmetric corticostriatal inputs on behavioral outcomes were not yet fully resolved.
After virally labeling either direct pathway (D1) or indirect pathway (D2) SPNs to optogenetically tag pathway-specific cortical inputs, the authors report that a much larger number of "non-starter" D2-SPNs from D2-SPN labeled mice responded to optogenetic stimulation in slices than "non-starter" D1 SPNs from D1-SPN labeled mice did. Without knowing the relative number of D1 or D2 SPN starters used to label cortical inputs, it is difficult to interpret the exact meaning of the lower number of responsive D2-SPNs in D1 labeled mice (where only ~63% of D1-SPNs themselves respond) compared to the relatively higher number of responsive D1-SPNs (and D2-SPNs) in D2 labeled mice. While relative differences in connectivity certainly suggest that some amount of asymmetric overlap of inputs exists, differences in infection efficiency and ensuing differences in detection sensitivity in slice experiments make determining the degree of asymmetry problematic.
It is also unclear if retrograde labeling of D1-SPN- vs D2-SPN- targeting afferents labels the same densities of cortical neurons. This gets to the point of specificity in the behavioral experiments. If the target-based labeling strategies used to introduce channelrhodopsin into specific SPN afferents label significantly different numbers of cortical neurons, might the difference in the relative numbers of optogenetically activated cortical neurons itself lead to behavioral differences?
In general, the manuscript would also benefit from more clarity about the statistical comparisons that were made and sample sizes used to reach their conclusions.
Homestar

Author Response
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public Review):
Trebino et al. investigated the BRAF activation process by analysing the interactions of BRAF N-terminal regulatory regions (CRD, RBD, and BSR) with the C-terminal kinase domain and with the upstream regulators HRAS and KRAS. To this end, they generated four constructs comprising different combinations of N-terminal domains of BRAF and analysed their interaction with HRAS as well as conformational changes that occur. By HDX-MS they confirmed that the RBD is indeed the main mediator of interaction with HRAS. Moreover, they observed that HRAS binding leads to conformational changes exposing the BSR to the environment. Next, the authors used OpenSPR to determine the binding affinities of HRAS to the different BRAF constructs. While BSR+RBD, RBD+CRD, and RBD bound HRAS with nanomolar affinity, no binding was observed with the construct comprising all three domains. Based on these experiments, the authors concluded that BSR and CRD negatively regulate binding to HRAS and hypothesised that BSR may confer some RAS isoform specificity. They corroborated this notion by showing that KRAS bound to BRAF-NT1 (BSR+RBD+CRD) while HRAS did not. Next, the authors analysed the autoinhibitory interaction occurring between the N-terminal regions and the kinase domain. Through pulldown and OpenSPR experiments, they confirm that it is mainly the CRD that makes the necessary contacts with the kinase domain. In addition, they show that the BSR stabilizes these interactions and that the addition of HRAS abolishes them. Finally, the D594G mutation within the KD of BRAF is shown to destabilise these autoinhibitory interactions, which could explain its oncogenic potential.
Overall, the in vitro study provides new insights into the regulation of BRAF and its interactions with HRAS and KRAS through a comprehensive in vitro analysis of the BRAF N-terminal region. Also, the authors report the first KD values for the N- and C-terminal interactions of BRAF and show that the BSR might provide isoform specificity towards KRAS. While these findings could be useful for the development of a new generation of inhibitors, the overall impact of the manuscript could probably be enhanced if the authors were to investigate in more detail how the BSR-mediated specificity of BRAF towards certain RAS isoforms is achieved. Moreover, though the very "clean" in vitro approach is appreciated, it also seems useful to examine whether the observed interactions and conformational changes occur in the full-length BRAF molecule and in more physiological contexts. Some of the results could be compared with studies including full-length constructs.
Public Response: We would like to express our gratitude for your valuable feedback on our manuscript. Your insightful suggestions have significantly improved the quality and completeness of our research. In response to your comments, we have conducted additional experiments and incorporated new data into the revised manuscript.
To gain a deeper understanding of how the BSR-mediated specificity of BRAF towards certain RAS isoforms is achieved, we performed HDX-MS to investigate the impact of KRAS interactions on the BSR. Our findings indicate that when KRAS is bound to BRAF NT2, there is no significant difference in hydrogen-deuterium exchange rates in the BSR compared to the apo-NT2 state (Figure 4). This observation contrasts with the effect of HRAS binding, where peptides from the BRAF-BSR exhibit an increased rate change, suggesting that HRAS induces a conformationally more dynamic state (Figure 2).
Our results align with the conclusions of Terrell et al. in their 2019 publication, which propose that isoform preferences in the RAS-RAF interaction are driven by opposite charge attractions between BRAF-BSR and KRAS-HVR, promoting the interaction.1 Our data offers a potential mechanistic explanation, suggesting that HRAS disrupts the conformational stability of the BSR provided by the RBD, while KRAS-HVR restores stability and enhances interaction favorability. It is important to note that our results do not directly confirm a long-lasting interaction between the BRAF-BSR and KRAS-HVR, but they do not rule out the possibility of a transient, low-affinity interaction or close proximity between the two.
Furthermore, our binding kinetics measurements conducted using OpenSPR support these findings. Particularly, in the case of NT1, when the CRD accompanies the BSR and RBD, no interactions with HRAS were observed. Additionally, we quantified the binding affinities between NT3:KRAS and NT4:KRAS, demonstrating that they are equally strong and that the presence of the BSR or CRD does not singularly affect the primary RBD interaction, consistent with HRAS. The BSR appears to exert an inhibitory effect on HRAS when the entire N-terminal region (BSR+RBD+CRD) is present. The BSR-mediated specificity is achieved through a coordinated interplay with the CRD.
Moreover, we have addressed your concern regarding the physiological relevance of our conclusions. In response, we utilized active, full-length (FL) BRAF purified from HEK293F cells in OpenSPR experiments. Our findings indicate that FL-BRAF behaves similarly to BRAF-NT1, as it does not bind to HRAS but binds to KRAS with a deviation comparable to NT1. We have demonstrated that post-translational modifications or native intramolecular interactions do not alter our initial results. Several literature sources, employing cell systems or expressing proteins from insect or mammalian cells, further support the findings presented in our study.2–5
Thank you once again for your constructive feedback, which has contributed significantly to the refinement of our work.
For the author:
Major points:
- Figure 1D: Negative control is missing.
Response: We have incorporated the negative control into this figure as suggested.
- Figure 3F and G: negative controls (GST only) are missing.
Response: We have incorporated the negative control into this figure as suggested.
- The authors demonstrate that BRAF NT1 (BSR+RBD+CRD) interacts with KRAS but not HRAS in SPR experiments (Figure 4). What about the conformational change that affects the positioning of BSR when NT2 (BSR+RBD) binds to HRAS (Figure 2)? Does it also occur with KRAS or not? When a rate change is observed between free protein and bound protein in HDX, particularly when this rate change results in a sigmoidal curve that closely parallels the reference curve, it signifies that all residues within the peptide share a uniform protection factor. This suggests that they collectively undergo conformational changes at the same rate, likely due to a concerted opening as a cohesive unit. In the context of our time plots, we observe this distinctive characteristic in the curves derived from the BSR peptides, indicating that HRAS binding perturbs this region, alters its flexibility, and induces a coordinated conformational shift. This compelling evidence strongly supports our assertion that HRAS instigates a reorientation of the BSR.
Response: In response to the reviewer's comments, we conducted additional experiments to explore whether KRAS elicits any comparable alterations in the H-D exchange of the BSR within BRAF-NT2. Our findings indicate that KRAS does not induce a similar conformational change in the BSR. We have detailed these results in the Results section under the heading "BSR Differentiates the BRAF-KRAS Interaction from the BRAF-HRAS Interaction" and have included corresponding panels in Figure 4 to visually illustrate these observations.
- Related to point 3: The authors mention that the HVR domain is responsible for isoform-specific differences. Does the BSR interact with the HVR domain of KRAS (but not HRAS)?
Response: It has been suggested by Terrell and colleagues1 that the BRAF-BSR and KRASHVR are directly responsible for the isoform specific interactions. We have no direct evidence confirming an interaction between the HVR and BSR. However, we deduce the possibility of such interaction based on previous research findings. Our HDX-MS experiments have demonstrated that the BRAF-BSR does not engage with HRAS. In our new HDX-MS experiments involving KRAS, we observed that the presence of KRAS does not lead to any discernible increase or decrease in the rate of deuterium exchange within the BRAF-BSR. It is important to emphasize that the absence of a rate change does not necessarily negate the occurrence of binding; rather, it might indicate a transient interaction with an affinity level below the detection threshold of HDX-MS.
Given that the only major difference between H- and K-RAS isoforms is the HVR, we hypothesize that binding differences between BRAF and RAS isoforms can be attributed to the HVR. Notably, BRAF-NT3 resembles CRAF, which also behaves in line with the findings from Terrell et al. in which the BSR is not present to impact RAS-RAF association. We have updated some of the discussion section to include the new results and draw relevant conclusion.
We mention in the text in the results section, “The HVR is an important region for regulating RAS isoform differences, like membrane anchoring, localization, RAS dimerization, and RAF interactions6… These results, combined with HDX-MS results, which showed that the BSR is exposed when bound to HRAS, suggest that the electrostatic forces surrounding the BSR promote BRAF autoinhibition and the specificity of RAF-RAS interactions.”
We also write in the discussion, “However, BRET assays suggest that CRAF does not show preference for either H- or KRAS, while BRAF appears to prefer KRAS.1 This preference is suggested to result from the potential favorable interactions between the negatively charged BSR of BRAF and the positively charged, poly-lysine region of the HVR of KRAS1… Our binding data provide additional examples of isoform-specific activity. We speculate that diminished BRAF-NT1 binding to HRAS and increased BSR exposure upon HRAS binding may be due to electrostatic repulsion between HRAS and the BSR. Our full-length KRAS and its interaction with NT1 support the hypothesis that the BSR attenuates fast binding to HRAS but not to KRAS.”
- The authors might consider including NRAS in their study to give more weight to this interesting aspect.
Response: While this suggestion is intriguing and could contribute to the expanding body of literature on RAS signaling, particularly in the context of NRAS-mutant tumors, we believe that delving into this topic would be beyond the scope of the present manuscript.
- Figure 6A: In this pulldown experiment the authors wish to demonstrate that binding of HRAS abolishes the autoinhibitory binding between NT1 and the kinase domain. However, the experimental design (i.e., pulldown of RAS) does not allow us to assess whether NT1 and KD are bound to each other in these conditions at all. The authors should rather pull down the KD and show that the interaction with NT1 is abolished when RAS is added.
Response: We appreciate your suggestion. The experimental design for this study was intentionally structured to focus on the specific subset of NT1 that interacts with HRAS. The BRAF N-terminal region has the capacity to bind both HRAS and KD, resulting in two distinct populations within BRAF-NT1: NT1:KD and NT1:HRAS, although we believe the ratio between those two populations is not 1:1. If we were to design the experiment by isolating either the KD or NT1, it would lead to the observation of both populations simultaneously, making it challenging to distinguish between them. Our pulldown experiments are performed under the same conditions (i.e. all the proteins were maintained in a molar ratio of 1:1 and exposed to the same buffer components), and we rely on pulldown assays, such as those depicted in Figure 5, to clearly demonstrate the binding interactions between NT1 and KD.
- The authors have chosen a purely in vitro approach for their interaction studies, which initially makes sense for the addressed questions. However, since the BRAF constructs studied are only fragments and neither BRAF nor K/HRAS has any posttranslational modifications, the question arises to what extent the findings obtained hold up in vivo. Therefore, the manuscript would greatly benefit from monitoring the described interactions in full-length proteins and in cells or at least with proteins purified from cells.
Response: Thank you for your valuable suggestion, which we take very seriously to enhance the quality of our manuscript. Upon carefully reviewing your comments, we conducted additional experiments involving full-length, wild-type BRAF (FL-BRAF) that was purified from mammalian cells, encompassing the post-translational modifications and scaffolding proteins such as 14-3-3 (Supplementary Fig 8A). We have incorporated the findings from these OpenSPR experiments into the revised manuscript within the Results Section titled "BSR Differentiates the BRAF-KRAS Interaction from the BRAFHRAS Interaction" and Figure 4. In summary, our results with FL-BRAF affirm the extension of our initial observations. Both NT1 and FL-BRAF interact with KRAS with comparable affinities, and neither NT1 nor FL-BRAF demonstrates an interaction with HRAS using OpenSPR. These results underscore that BRAF fragments accurately represent active, fully processed BRAF, lending support to our in vitro approach.
Moreover, the conserved interactions we report in this manuscript are supported by literature. The interaction between RAF-RBD and RAS has been extensively documented, spanning investigations conducted in both insect and mammalian cell lines. For instance, Tran et al. (2021) utilized mammalian expression systems to explore the role of RBD in mediating BRAF activation through RAS interaction, identifying the same binding surfaces that we highlighted using HDX-MS.2 They quantified the KRAS-CRAF interaction yielding binding affinities in the low nanomolar range, similar to our findings for BRAF-NT:KRAS OpenSPR.2 In the manuscript text, we compared the binding affinity of BRAF residues 1245 purified from insect cells3 to our BRAF 1-227 (NT2 from E. coli), noting that the published value falls within the standard deviation of our experimental value. Additionally, our results align with the autoinhibited FL-BRAF:MEK:14-3-3 structure, which was expressed in Sf9 insect cells and reveals the central role of the CRD in maintaining autoinhibition through interactions with KD.4 In 2005, Tran and colleagues revealed specific domains within the BRAF N-terminal region are involved in binding to KD through Co-IP experiments conducted in mammalian cells.5
While we are fully aware of the limitations of taking a purely in vitro approach to study the role of BRAF regulatory domains in RAS-RAF interactions and autoinhibition, as well as to quantify the affinity of these interactions, we emphasize that this approach enables us to dissect and examine the specific regions of RAF that are under investigation. As we write in the manuscript: “Our in vitro studies were conducted using proteins purified from E. coli, which lack the membrane, post-translational modifications, and regulatory, scaffolding, or chaperone proteins that are involved in BRAF regulation. Nonetheless, our study provides a direct characterization of the intra- and inter-molecular protein-protein interactions involved in BRAF regulation, without the complications that arise in cell-based assays.” We have added the following comment to clarify the advantages of our in vitro approach and the challenges associated with cell-based assays: “… without the complications and false-positives that can arise in cell-based assays, which often cannot distinguish between proximity and biochemical interactions.”
Once again, we appreciate your insight feedback, which has contributed significantly to the improvement of our manuscript.
Minor:
- Page 7, paragraph 2, line 6: It should probably read "BRAF autoinhibition" not "BRAF autoinhibitory".
Response: Thank you for bringing this to our attention. We have fixed this typo.
- Figure 3G: In the first lane (time point 0 min) there is no input band for His/MBP-NT1. Probably a mistake when cropping the image from the original photo.
Response: We sincerely appreciate your diligence in identifying cropping errors, and we have taken comprehensive measures to review the manuscript and correct any such errors. Regarding this specific figure, it is important to note that NT1 was not added at the "0" minute time point, which explains the absence of an input band at that stage. To avoid any confusion, we have revised the notation from "0" to "-" for clarity.
Reviewer #2 (Public Review):
In the manuscript entitled 'Unveiling the Domain-Specific and RAS Isoform-Specific Details of BRAF Regulation', the authors conduct a series of in vitro experiments using Nterminal and C-terminal BRAF fragments (SPR, HDX-MS, pull-down assays) to interrogate BRAF domain-specific autoinhibitory interactions and engagement by H- and KRAS GTPases. Of the three RAF isoforms, BRAF contains an extended N-terminal domain that has yet to be detected in X-ray and cryoEM reconstructions but has been proposed to interact with the KRAS hypervariable region. The investigators probe binding interactions between 4 N-terminal (NT) BRAF fragments (containing one more NT domain (BRS, RBD, and CRD)), with full-length bacterial expressed HRAS, KRAS as well as two BRAF C-terminal kinase fragments to tease out the underlying contribution of domainspecific binding events. They find, consistent with previous studies, that the BRAF BSR domain may negatively regulate RAS binding and propose that the presence of the BSR domain in BRAF provides an additional layer of autoinhibitory constraints that mediate BRAF activity in a RAS-isoform-specific manner. One of the fragments studied contains an oncogenic mutation in the kinase domain (BRAF-KDD594G). The investigators find that this mutant shows reduced interactions with an N-terminal regulatory fragment and postulate that this oncogenic BRAF mutant may promote BRAF activation by weakening autoinhibitory interactions between the N- and C-terminus.
While this manuscript sheds light on B-RAF specific autoinhibitory interactions and the identification and partial characterization of an oncogenic kinase domain (KD) mutant, several concerns exist with the vitro binding studies as they are performed using taggedisolated bacterial expressed fragments, 'dimerized' RAS constructs, lack of relevant citations, controls, comparisons and data/error analysis. Detailed concerns are listed below.
- Bacterial-expressed truncated BRAF constructs are used to dissect the role of individual domains in BRAF autoinhibition. Concerns exist regarding the possibility that bacterial expression of isolated domains or regions of BRAF could miss important posttranslational modifications, intra-molecular interactions, or conformational changes that may occur in the context of the full-length protein in mammalian cells. This concern is not addressed in the manuscript.
Response: Reviewer 1 raised a similar concern, and we have duplicated our response below for your reference:
Thank you for your valuable suggestion, which we take very seriously to enhance the quality of our manuscript. Upon carefully reviewing your comments, we conducted additional experiments involving full-length, wild-type BRAF (FL-BRAF) that was purified from mammalian cells, encompassing the post-translational modifications and scaffolding proteins such as 14-3-3 (Supplementary Fig 8A). We have incorporated the findings from these OpenSPR experiments into the revised manuscript within the Results Section titled "BSR Differentiates the BRAF-KRAS Interaction from the BRAF-HRAS Interaction" and Figure 4. In summary, our results with FL-BRAF affirm the extension of our initial observations. Both NT1 and FL-BRAF interact with KRAS with comparable affinities, and neither NT1 nor FL-BRAF demonstrates an interaction with HRAS using OpenSPR. These results underscore that BRAF fragments accurately represent active, fully processed BRAF, lending support to our in vitro approach.
Moreover, the conserved interactions we report in this manuscript are supported by literature. The interaction between RAF-RBD and RAS has been extensively documented, spanning investigations conducted in both insect and mammalian cell lines. For instance, Tran et al. (2021) utilized mammalian expression systems to explore the role of RBD in mediating BRAF activation through RAS interaction, identifying the same binding surfaces that we highlighted using HDX-MS.2 They quantified the KRAS-CRAF interaction yielding binding affinities in the low nanomolar range, similar to our findings for BRAF-NT:KRAS OpenSPR.2 In the manuscript text, we compared the binding affinity of BRAF residues 1245 purified from insect cells3 to our BRAF 1-227 (NT2 from E. coli), noting that the published value falls within the standard deviation of our experimental value. Additionally, our results align with the autoinhibited FL-BRAF:MEK:14-3-3 structure, which was expressed in Sf9 insect cells and reveals the central role of the CRD in maintaining autoinhibition through interactions with KD.4 In 2005, Tran and colleagues revealed specific domains within the BRAF N-terminal region are involved in binding to KD through Co-IP experiments conducted in mammalian cells.5
While we are fully aware of the limitations of taking a purely in vitro approach to study the role of BRAF regulatory domains in RAS-RAF interactions and autoinhibition, as well as to quantify the affinity of these interactions, we emphasize that this approach enables us to dissect and examine the specific regions of RAF that are under investigation. As we write in the manuscript: “Our in vitro studies were conducted using proteins purified from E. coli, which lack the membrane, post-translational modifications, and regulatory, scaffolding, or chaperone proteins that are involved in BRAF regulation. Nonetheless, our study provides a direct characterization of the intra- and inter-molecular protein-protein interactions involved in BRAF regulation, without the complications that arise in cell-based assays.” We have added the following comment to clarify the advantages of our in vitro approach and the challenges associated with cell-based assays: “… without the complications and false-positives that can arise in cell-based assays, which often cannot distinguish between proximity and biochemical interactions.”
Once again, we appreciate your insight feedback, which has contributed significantly to the improvement of our manuscript.
- The experiments employ BRAF NT constructs that retain an MBP tag and RAS proteins with a GST tag. Have the investigators conducted control experiments to verify that the tags do not induce or perturb native interactions?
Response: Thank you for highlighting this important issue. We have conducted control experiments whenever feasible, particularly in cases where tags were not required for visualization, immobilization, or where cleave sites were present. We have subsequently included these control experiments in the supplementary figures and accompanying text within the manuscript.
It is essential to note that many of the techniques employed in this manuscript rely on tags, such as immobilizing proteins onto NTA OpenSPR sensors and employing various resins/beads for pulldown assays. Utilizing tags for protein immobilization in OpenSPR applications offers distinct advantages, including homogeneous and site-specific immobilization of the protein, ensuring that binding sites remain accessible for the study of protein-protein interactions (PPIs) of interest. Furthermore, in all BRAF-RAS SPR experiments, the MBP protein serves as the reference channel "blocking" protein. This reference channel is instrumental in mitigating any potential false-positive signals resulting from binding interactions with the MBP protein. Any such signal is subsequently subtracted out during data analysis.
To provide a comprehensive understanding of these aspects, we have incorporated these details into the manuscript text for clarity:
“Maltose bind protein (MBP) is immobilized on the OpenSPR reference channel, which accounts for any non-specific binding or impacts to the native PPIs that may result from the presence of tags. Kinetic analysis is performed on the corrected binding curves, which subtracts any response in the reference channel.”
We describe the control experiment to examine whether His/MBP-tag affects NT1 binding with BRAF-KD: “Similarly, we removed the His/MBP-tag from BRAF-NT1 through a TEV protease cleavage reaction and flowed over untagged NT1. Kinetic analysis confirmed that the interaction is preserved with the KD=13 nM (Supplemental Figure 6F).”
We show that the GST-tag does not affect KRAS interactions with NTs in supplemental figure 6. We purified full-length, His/MBP-KRAS and subsequently removed the tag through TEV cleavage. BRAF-NT interactions are preserved with untagged KRAS. GST alone, also does not interact with BRAF-NTs. We updated the text in the results section “BSR differentiates the BRAF-KRAS interaction from the BRAF-HRAS interaction.”
Additionally, Vojtek and colleagues used the same fusion-protein combinations (GSTRAS and MBP-RAF) in pulldown experiments and also found no perturbations from these tags.8
- The investigators state that the GST tag on the RAS constructs was used to promote RAS dimerization, as RAS dimerization is proposed to be key for RAF activation. However, recent findings argue against the role of RAS dimers in RAF dimerization and activation (Simanshu et al, Mol. Cell 2023). Moreover, while GST can dimerize, it is unclear whether this promotes RAS dimerization as suggested. In methods for the OpenSPR experiments probing NT BRAF:RAS interactions, it is stated that "monomeric KRAS was flowed...". This terminology is a bit confusing. How was the monomeric state of KRAS determined and what was the rationale behind the experiment? Is there a difference in binding interactions between "monomeric vs dimeric KRAS"?
Response: Thank you for conducting such a comprehensive review of our manuscript and for identifying the mention of "monomeric KRAS" in the experimental section, which was inadvertently included and should not have been present. This terminology originally referred to a series of experiments involving "monomeric" KRAS that were initially considered for inclusion in the main body of the manuscript but were subsequently removed before submission. Furthermore, we adjusted the terminology to prevent any confusion or unwarranted implications.
To clarify, this "monomeric" construct refers to the tagless, full-length KRAS variant that was confirmed to exist in a monomeric state through Size Exclusion Chromatography, eluting at a volume equivalent to 21 kDa. We have incorporated the findings from experiments involving this untagged KRAS variant into the supplementary figures to provide supporting evidence, particularly in response to comment #2, that the GST-tag does not interfere with native interactions. Supplementary Figure 1 illustrates that both GST-HRAS (45 kDa) and GST-KRAS (45 kDa) elute as dimers in solution, at approximately 90 kDa. It is important to note that the main text figures primarily feature the GST-tagged, "dimeric" RAS constructs. Our research results do not suggest any significant differences between "monomeric," untagged KRAS and "dimeric" GST-tagged KRAS, indicating that the binding kinetics between RAS and RAF are not influenced by oligomerization state (Supplementary Fig 6). To mitigate any potential confusion, we have made the necessary distinctions in the text and have revised the methods description to accurately reflect these aspects.
While the recent findings summarized by Simanshu and colleagues were published concurrently with our manuscript submission, we would like to address this comment in the following manner. The authors assert that RAS does not engage in dimerization through the G domain, a hypothesis that contrasts with certain prior research findings. Instead, they propose that the plasma membrane plays a pivotal role in the clustering of RAS. Furthermore, the authors mention the involvement of RAS "dimerization" in RAF dimerization and activation in the subsequent statements:
“Recruitment of two RAF proteins by RAS proteins in close proximity facilitate RAF activation but are not required for RAF dimerization.”
“However, the PM recruitment of two RAF proteins by two non-dimerized but co- localized RAS proteins would serve equally well to promote RAF dimerization. Moreover, recent work on the activation cycle of RAF dimers (ref 20–23) argues strongly against a role for RAS dimers while revealing regulation by the 14-3-3 and SHOC2-MRAS- PP1C complexes. (Ref 24)”
The primary focus of our study centers on elucidating the intricate details of the RAS-RAF interaction and the mechanisms underlying RAF autoinhibition, rather than emphasizing RAF dimerization as the sole pathway to RAF activation. It is important to recognize that RAF activation encompasses multiple steps, including RAS-mediated relief of RAF autoinhibition.
To mimic physiological conditions as closely as possible, we employed a GST-tag on RAS in our experiments. It's worth noting that GST has a dimerization property,9 which brings RAS molecules into close proximity to one another, effectively emulating conditions akin to the plasma membrane. Our primary objective is not solely to facilitate interactions by bringing RAS into close proximity. Instead, our aim is to replicate cellular conditions to the greatest extent feasible, especially within the predominantly in vitro framework of our studies. Furthermore, we have revised the sentence pertaining to HRAS as follows: “As verified by size exclusion chromatography (Supplementary Fig 1A), the GST-tag dimerizes and forces HRAS into close proximity to recapitulate physiological conditions. (ref. 35)”
- The investigators determine binding affinities between GST-HRAS and NT BRAF domains (NT2 7.5 {plus minus} 3.5; NT3 22 {plus minus} 11 nM) by SPR, and propose that the BRS domain has an inhibitory role HRAS interactions with the RAF NT. However, it is unclear whether these differences are statistically meaningful given the error.
Response: Thank you for bringing up this matter for further discussion. We are fully aware that these distinctions (NT2 and NT3), considering the overlapping error, lack statistical significance. Our conclusion points toward the most notable differences occurring when comparing NT1 to either NT2 or NT3, highlighting that the presence of the BSR has an inhibitory effect, particularly when the CRD is also present. It's important to note that we did not directly compare NT2 and NT3 to each other. Our comparison primarily elucidates that BSR without the CRD, and conversely, CRD without the BSR, do not exhibit the inhibitory effect. This collective evidence leads to the conclusion that all three domains collaboratively play a role in negatively regulating BRAF against HRAS.
- It is unclear why NT1 (BSR+RBD+CRD) was not included in the HDX experiments, which makes it challenging to directly compare and determine specific contributions of each domain in the presence of HRAS. Including NT1 in the experimental design could provide a more comprehensive understanding of the interplay between the domains and their respective roles in the HRAS-BRAF interaction. Further, excluding certain domains from the constructs, such as the BSR or CRD, may overlook potential domain-domain interactions and their influence on the conformational changes induced by HRAS binding.
Response: We acknowledge that incorporating NT1 into the HDX experiments would have provided clearer insights into the specific contributions of each domain. Originally, it was our intention to include NT1 in these experiments. Unfortunately, we encountered challenges with the HDX experiments when it came to BRAF-NT1, as it yielded a significantly low sequence coverage after MS/MS analysis. We made multiple attempts to address this issue, which included additional protein purifications involving reducing agents, increasing the concentration of reaction buffer components, and extending the incubation time with reducing agents before injection. Despite these efforts, we were unable to obtain the desired sequence coverage for NT1. Consequently, we switched our approach to analyze NT2 and NT3 as the next best alternative.
- The authors perform pulldown experiments with BRAF constructs (NT1: BSR+RBD+CRD, NT2: BSR+RBD, NT3: RBD+CRD, NT4: RBD alone), in which biotinylated BRAF-KD was captured on streptavidin beads and probed for bound His/MBP-tagged BRAF NTs. Western blot results suggest that only NT1 and NT3 bind to the KD (Figure 5). However, performing a pulldown experiment with an additional construct, CRD alone, it would help to determine whether the CRD alone is sufficient for the interaction or if the presence of the RBD is required for higher affinity binding. This additional experiment would strengthen the authors' arguments and provide further insights into the mechanism of BRAF autoinhibition.
Response: We are grateful for this valuable suggestion, and in response, we have taken the initiative to clone and purify a CRD-only construct (NT5) to strengthen our arguments. Subsequently, we conducted OpenSPR experiments to measure the binding affinity between NT5 and KD. Our findings clearly indicate that the CRD alone is not sufficient to mediate the autoinhibitory interactions and that the presence of the RBD is indeed necessary. These results have been incorporated into Figure 5 and are described within the Results Section for enhanced clarity and support.
- While the investigators state that their findings indicate that H- and KRAS differentially interact with BRAF, most of the experiments are focused on HRAS, with only a subset on KRAS. As SPR & pull-down experiments are only conducted on NT1 and NT2, evidence for RAS isoform-specific interactions is weak. It is unclear why parallel experiments were not conducted with KRAS using BRAF NT3 & NT4 constructs.
Response: We sincerely appreciate your suggestion, which has contributed to enhancing the overall robustness of the evidence regarding isoform-specific differences between H- and K-RAS. In response, we performed additional experiments involving NT3 and NT4. The outcomes of these experiments have been integrated into Figure 4, and we have provided a comprehensive description of these results within the Results section “BSR differentiates the BRAF-KRAS interaction from the BRAF-HRAS interaction” of the manuscript.
- The investigators do not cite the AlphaFold prediction of full-length BRAF (AFP15056-F1) or the known X-ray structure of the BRAF BRS domain. Hence, it is unclear how Alpha-Fold is used to gain new structural information, and whether it was used to predict the structure of the N-terminal regulatory or the full-length protein.
Response: We greatly appreciate the reviewer’s commitment to upholding good scientific practices and ensuring the inclusion of relevant citations in publications. In our original manuscript, we employed the UniProt ID P15056 to reference the specific AlphaFold structure used in our study. This was clarified as follows: "Since the full-length structure of BRAF is still unresolved, we applied the AlphaFold Protein Structure Database for a model of BRAF to display the conformation of the N-terminal domains and the HDX-MS results.40,41” Additionally, we referenced AlphaFold using the two citations recommended on their website (references 35 and 36 in the original manuscript). To prevent any potential confusion in the future, we have incorporated "AF-P15056-F1," as suggested.
We are sorry for any misunderstanding that may have arisen regarding the use of AlphaFold for gaining new structural insights. Our sole intention was to utilize AlphaFold as a tool for modeling HDX, as a full-length structure of BRAF, encompassing the entire N-terminal domain, remains unavailable. We have taken steps to clarify our objectives in the manuscript to ensure the purpose of our AlphaFold utilization is unambiguous.
Furthermore, we wish to emphasize that our utilization of AlphaFold was never intended to exclude the known X-ray structure of the BRAF-BSR domain. In our revised text, we have added clarity to our purposes and cited the Lavoie et al. Nature publication from 2018, which provides alignment between the X-ray structure and the AlphaFold model, thereby enhancing the confidence in the latter.
- In HDX-MS experiments, it is unclear how the authors determine whether small differences in deuterium uptake observed for some of the peptide fragments are statistically significant, and why for some of the labeling reaction times the investigators state " {plus minus} HRAS only" for only 3 time points?
Response: First, in reference to the question about " ‘{plus minus} HRAS only’ for only 3 time points,” we write:
“Both constructs were incubated with and without GMPPNP-HRAS in D2O buffer for set labeling reaction times (NT3: 2 sec [NT3 ± HRAS only], 6 sec [NT3 ± HRAS only], 20 sec, 30 sec [NT3 ± HRAS only], 60 sec, 5 min, 10 min, 30 min, 90 min, 4.5 h, 15 h, and 24 h)...”
We realize how this can be confusing. To avoid such confusion, we fixed the text to read instead:<br /> “Both constructs were incubated with and without GMPPNP-HRAS in D2O buffer for set labeling reaction times (NT3: 2 sec, 6 sec, 20 sec, 30 sec, 60 sec, 5 min, 10 min, 30 min, 90 min, 4.5 h, 15 h, 45 h and 24 h at RT; NT2: 20 sec, 60 sec, 5 min, 10 min, 30 min, 90 min, 4.5 h, 15 h, and 24 h at RT)...”
Next, with regard to assessing significance, we determine it by closely examining a consistent trend in smooth time course plots. To establish this trend, we rely on the presence of more than four overlapping peptides, each with multiple charge states, within a specific sequence range. When we observe multiple peptides showing even a small difference in rate exchange, we can confidently infer that structural changes have taken place. This confidence stems from the inherent reliability and redundancy in the data analysis approach we have employed.11,12 It is noteworthy that our focus is primarily on reporting the binding or no binding, rather than quantifying the magnitude of exchange. As such, conducting multiple replicates or statistical testing is not deemed necessary.13,14 This is true for multiple reasons:
1) Instead of small deuterium changes (y-axis), we are focusing on the x-axis changes, which provides a slowing factor and how much that H-D exchange rate has changed.
In a publication investigating the ideal HDX-MS data set, the author explains, “with the availability of high resolution HDX-MS raw data, it may be the time to shift the data analysis paradigm from determination of centroid values and presentation of deuteration levels to deconvolution of isotope envelopes and presentation of exchange rates.” 15
Presentation of data through rate changes provides a physical chemistry measurement, as opposed to a relative measurement with percent deuteration. For example, slowing with a factor of 10 equates to the energy in 1 kCal. By quick visual estimation, we see a slowing factor of about 2 when RAS is bound to the BRAF-RBD.
We made some changes to the text to clear up any confusion about measuring D uptake vs rate.
2) Looking at sigmoidal curves only—the “smooth time course” shows that the timedependent deuterium changes are not random, artifacts, or false positives/negatives. When parallel sigmoidal curves are present, any x-axis change is a measure of H-D exchange. Only plots with a smooth time course are used to make conclusions about BRAF’s conformational changes or binding interfaces.
3) Wide time range- the extended time also confirms that any observed difference is reliable and accurate. This extended time frame provides coverage for deuteration levels from 0 to 100% for peptides. A smooth time course is present in complete coverage.
4) The rate change is observed at multiple time points (at least 4 for each peptide), which are all independent reactions, and show reproducibility of change
5) Many overlapping peptides show the same pattern- the exchange rate difference is observed in at least 4 peptide time plots without contradictory evidence within the sequence range.
6) The many other peptide time plots that do not show any difference with and without RAS is a form of reproducibility, that no difference means no difference.
- The investigators find that KRAS binds NT1 in SPR experiments, whereas HRAS does not. However, the pull-down assays show NT1 binding to both KRAS and HRAS. SI Fig 5 attributes this to slow association, yet both SPR (on/off rates) and equilibrium binding measurements are conducted. This data should be able to 'tease' out differences in association.
Response: Thank you for bringing up this important point. It's crucial to note that the experiments conducted at slow flow rates generated low responses, making it challenging to perform kinetic analyses effectively. Consequently, we are unable to provide accurate equilibrium binding measurements (on/off rates) for NT1 and HRAS. Regrettably, comparing the association rates between KRAS and HRAS is not feasible due to the differing flow rates employed. We have addressed this limitation in the manuscript as follows:
“We therefore immobilized NT1 and flowed over HRAS at a much slower flow rate (5 µL/min), during which we saw minimal but consistent binding (Supplementary Fig 5A). The low response and long timeframe of each injection, however, makes the dissociation constant (KD) unmeasurable and incomparable to our other NT-HRAS OpenSPR results.”
- The model in Figure 7B highlights BSR interactions with KRAS, however, BSR interactions with the KRAS HVR (proximal to the membrane) are not shown, as supported by Terrell et al. (2019).
Response: Thank you for the suggestion. We reoriented the BSR closer to HVR of KRAS rather than G-domain.
- The investigators state that 'These findings demonstrate that HRAS binding to BRAF directly relieves BRAF autoinhibition by disrupting the NT1-KD interaction, providing the first in vitro evidence of RAS-mediated relief of RAF autoinhibition, the central dogma of RAS-RAF regulation. However, in Tran et al (2005) JBC, they report pulldown experiments using N-and C-terminal fragments of BRAF and state that 'BRAF also contains an N-terminal autoinhibitory domain and that the interaction of this domain with the catalytic domain was inhibited by binding to active HRAS'. This reference is not cited.
Response: We appreciate the concern raised regarding our statement. We want to clarify that it was never our intention to disregard this JBC publication, and we apologize for any misunderstanding caused by our phrasing. We recognize that our initial statement was contentious, and we have removed the word "first" from the phrase "first in vitro evidence." In the section of the discussion where we originally cited the Tran et al. (2005) publication, we have revised the language to eliminate "first" and have rephrased the sentence, as provided below:
“Our in vitro binding studies align with previous implications that RAS relieves RAF autoinhibition shown through cell-based coIP’s.5”
- In Fig 2, panels A and C, it is unclear what the grey dotted line in is each plot.
Response: Thank you for drawing our attention to the additional explanation needed here. The gray dotted lines represent the maximum deuterium exchange. We added the following description to the figure 2 legend:
“Gray dotted lines represent the theoretical exchange behavior for specified peptide that is fully unstructured (top) or for specified peptide with a uniform protection factor (fraction of time the residue is involved in protecting the H-bond) of 100 (lower).”
- In Fig 3, error analysis is not provided for panel E.
Response: We added the standard deviation values to this panel. We additionally added these for Fig 4C and Fig 5B.
- How was RAS GMPPNP loading verified?
Response: Ras loading is a well-established protocol with a solid foundation in the literature.16– 21 We followed this accepted method for nucleotide exchange. Our controls, as evident in pulldown and OpenSPR experiments (fig 1C, 4E), unequivocally demonstrate that GMPPNPloaded RAS is active, while unloaded RAS is inactive, as evidenced by the absence of no binding. We also added supplemental figure 6E to show that inactive (unloaded) GST-KRAS does not bind to BRAF during OpenSPR analysis. To exemplify this, we included binding curves of 1 µM GST-KRAS- GMPPNP and -GDP flowed over NTA-immobilized BRAF-NT2 at a flow rate of 30 µl/min.
References
(1) Terrell, E. M.; Durrant, D. E.; Ritt, D. A.; Sealover, N. E.; Sheffels, E.; Spencer-Smith, R.; Esposito, D.; Zhou, Y.; Hancock, J. F.; Kortum, R. L.; Morrison, D. K. Distinct Binding Preferences between Ras and Raf Family Members and the Impact on Oncogenic Ras Signaling. Mol. Cell 2019, 76 (6), 872-884.e5. https://doi.org/10.1016/j.molcel.2019.09.004.
(2) Tran, T. H.; Chan, A. H.; Young, L. C.; Bindu, L.; Neale, C.; Messing, S.; Dharmaiah, S.; Taylor, T.; Denson, J. P.; Esposito, D.; Nissley, D. V.; Stephen, A. G.; McCormick, F.; Simanshu, D. K. KRAS Interaction with RAF1 RAS-Binding Domain and Cysteine-Rich Domain Provides Insights into RAS-Mediated RAF Activation. Nat. Commun. 2021, 12 (1176), 1–16. https://doi.org/10.1038/s41467-021-21422-x.
(3) Fischer, A.; Hekman, M.; Kuhlmann, J.; Rubio, I.; Wiese, S.; Rapp, U. R. B- and C-RAF Display Essential Differences in Their Binding to Ras: The Isotype-Specific N Terminus of B-RAF Facilitates Ras Binding. J. Biol. Chem. 2007, 282 (36), 26503–26516. https://doi.org/10.1074/jbc.M607458200.
(4) Park, E.; Rawson, S.; Li, K.; Kim, B. W.; Ficarro, S. B.; Pino, G. G. Del; Sharif, H.; Marto, J. A.; Jeon, H.; Eck, M. J. Architecture of Autoinhibited and Active BRAF–MEK1–14-3-3 Complexes. Nature 2019, 575 (7783), 545–550. https://doi.org/10.1038/s41586-0191660-y.
(5) Tran, N. H.; Wu, X.; Frost, J. A. B-Raf and Raf-1 Are Regulated by Distinct Autoregulatory Mechanisms. J. Biol. Chem. 2005, 280 (16), 16244–16253. https://doi.org/10.1074/jbc.M501185200.
(6) Prior, I. A.; Hancock, J. F. Ras Trafficking, Localization and Compartmentalized Signalling. Semin. Cell Dev. Biol. 2012, 23 (2), 145–153.
(7) Herrmann, C.; Martin, G. A.; Wittinghofer, A. Quantitative Analysis of the Complex between P21 and the Ras-Binding Domain of the Human Raf-1 Protein Kinase. J. Biol. Chem. 1995, 270 (7), 2901–2905. https://doi.org/10.1074/jbc.270.7.2901.
(8) Vojtek, A. B.; Hollenberg, S. M.; Cooper, J. A. Mammalian Ras Interacts Directly with the Serine/Threonine Kinase Raf. Cell 1993, 74 (1), 205–214. https://doi.org/10.1016/00928674(93)90307-C.
(9) Parker, M. W.; Bello, M. Lo; Federici, G. Crystallization of Glutathione S-Transferase from Human Placenta. J. Mol. Biol. 1990, 213 (2), 221–222. https://doi.org/10.1016/S00222836(05)80183-4.
(10) Inouye, K.; Mizutani, S.; Koide, H.; Kaziro, Y. Formation of the Ras Dimer Is Essential for Raf-1 Activation. J. Biol. Chem. 2000, 275 (6), 3737–3740. https://doi.org/10.1074/JBC.275.6.3737.
(11) Z. Y. Kan, X. Ye, J. J. Skinner, L. Mayne, S. W. E. ExMS2: An Integrated Solution for Hydrogen-Deuterium Exchange Mass Spectrometry Data Analysis. Anal Chem 2019, 91 (11), 7474–7481.
(12) Mayne, L.; Kan, Z. Y.; Sevugan Chetty, P.; Ricciuti, A.; Walters, B. T.; Englander, S. W. Many Overlapping Peptides for Protein Hydrogen Exchange Experiments by the Fragment Separation-Mass Spectrometry Method. J. Am. Soc. Mass Spectrom. 2011, 22 (11), 1898–1905. https://doi.org/10.1007/S13361-011-0235-4.
(13) Ye, X.; Lin, J.; Mayne, L.; Shorter, J.; Englander, S. W. Hydrogen Exchange Reveals Hsp104 Architecture, Structural Dynamics, and Energetics in Physiological Solution. Proc. Natl. Acad. Sci. 2019, 116 (15), 7333–7342. https://doi.org/10.1073/pnas.1816184116.
(14) Ye, X.; Lin, J.; Mayne, L.; Shorter, J.; Englander, S. W. Structural and Kinetic Basis for the Regulation and Potentiation of Hsp104 Function. Proc. Natl. Acad. Sci. 2020, 117 (17), 9384–9392. https://doi.org/10.1073/pnas.1921968117.
(15) Hamuro, Y. Determination of Equine Cytochrome c Backbone Amide Hydrogen/Deuterium Exchange Rates by Mass Spectrometry Using a Wider Time Window and Isotope Envelope. J. Am. Soc. Mass Spectrom. 2017, 28 (3), 486–497. https://doi.org/10.1007/s13361-016-1571-1.
(16) Herrmann, C.; Horn, G.; Spaargaren, M.; Wittinghofer, A. Differential Interaction of the Ras Family GTP-Binding Proteins H-Ras, Rap1A, and R-Ras with the Putative Effector Molecules Raf Kinase and Ral-Guanine Nucleotide Exchange Factor. J. Biol. Chem. 1996, 271 (12), 6794–6800. https://doi.org/10.1074/jbc.271.12.6794.
(17) Miller, A. F.; Halkides, C. J.; Redfield, A. G. An NMR Comparison of the Changes Produced by Different Guanosine 5’-Triphosphate Analogs in Wild-Type and Oncogenic Mutant P21ras. Biochemistry 1993, 32 (29), 7367–7376. https://doi.org/10.1021/bi00080a006.
(18) Amendola, C. R.; Mahaffey, J. P.; Parker, S. J.; Ahearn, I. M.; Chen, W. C.; Zhou, M.; Court, H.; Shi, J.; Mendoza, S. L.; Morten, M. J.; Rothenberg, E.; Gottlieb, E.; Wadghiri, Y. Z.; Possemato, R.; Hubbard, S. R.; Balmain, A.; Kimmelman, A. C.; Philips, M. R. KRAS4A Directly Regulates Hexokinase 1. Nature 2019. https://doi.org/10.1038/s41586019-1832-9.
(19) John, J.; Sohmen, R.; Feuerstein, J.; Linke, R.; Wittinghofer, A.; Goody, R. S. Kinetics of Interaction of Nucleotides with Nucleotide-Free H-Ras P21. Biochemistry 1990, 29 (25), 6058–6065. https://doi.org/10.1021/bi00477a025.
(20) Dharmaiah, S.; Tran, T. H.; Messing, S.; Agamasu, C.; Gillette, W. K.; Yan, W.; Waybright, T.; Alexander, P.; Esposito, D.; Nissley, D. V.; McCormick, F.; Stephen, A. G.; Simanshu, D. K. Structures of N-Terminally Processed KRAS Provide Insight into the Role of N-Acetylation. Sci. Reports 2019 91 2019, 9 (1), 1–15. https://doi.org/10.1038/s41598-019-46846-w.
(21) Rathinaswamy, M. K.; Gaieb, Z.; Fleming, K. D.; Borsari, C.; Harris, N. J.; Moeller, B. E.; Wymann, M. P.; Amaro, R. E.; Burke, J. E. Disease-Related Mutations in PI3Kγ Disrupt Regulatory C-Terminal Dynamics and Reveal a Path to Selective Inhibitors. Elife 2021, 10. https://doi.org/10.7554/eLife.64691.
Reviewer #2 (Public Review):
In this study, the investigators describe an unbiased phosphoproteomic analysis of cardiac-specific overexpression of adenylyl cyclase type 8 (TGAC8) mice that was then integrated with transcriptomic and proteomic data. The phosphoproteomic analysis was performed using tandem mass tag-labeling mass spectrometry of left ventricular (LV) tissue in TGAC8 and wild-type mice. The initial principal component analysis showed differences between the TGAC8 and WT groups. The integrated analysis demonstrated that many stress-response, immune, and metabolic signaling pathways were activated at transcriptional, translational, and/or post-translational levels.
The authors are to be commended for a well-conducted study with quality control steps described for the various analyses. The rationale for following up on prior transcriptomic and proteomic analyses is described. The analysis appears thorough and well-integrated with the group's prior work. Confirmational data using Western blot is provided to support their conclusions. Their findings have the potential of identifying novel pathways involved in cardiac performance and cardioprotection.
It would have been fantastic to eschew this ridiculousness, because we all make fun of branded vulnerabilities too, but this was not the right time to make that stand.
UN-Generalsekretär Guterres hat die Untätigkeit der politischen Führungen scharf angegriffen. Er spricht davon, dass man die vergifteten Wurzeln der Klimakrise, die fossilen Energien, endlich ausreißen muss. Der Emission Gap Report 2023 zeigt, dass keiner der G20-Staaten eine Dekarbonisierungspolitik betreibt, die mit den Zielen des Pariser Abkommens vereinbar ist. https://www.liberation.fr/environnement/climat/le-monde-va-faire-face-a-un-rechauffement-de-25-c-a-29-c-dici-2100-alerte-lonu-avant-la-cop-28-20231120_QXFYQM3CJNHALBWI4PV5KR4NIY/
Mehr zum Emissions Gap Report 2023: https://hypothes.is/search?q=tag%3A%22Emissions%20Gap%20Report%202023%22
Hitzebedingte Todesfälle bei über 65-Jährigen haben seit den 90ern um 85% zugenommen. Senior:innen sind – wie kleine Kinder – zweimal soviel Hitzewellen-Tagen ausgesetzt wie 1986-2005. Extreme Hitze führte 2022 zu Produktivitätsverlusten von ca. 863 Milliarden USD. Alle Indikatoren für öffentliche Gesundheit haben sich in den letzten 9 Jahren verschlechtert. – Die NYT stellt den 2023 Report des Lancet Countdown ausführlich dar. https://www.nytimes.com/2023/11/14/climate/climate-change-health-effects-lancet.html
Mehr zum Rreport: https://hypothes.is/search?q=tag%3A%222023%20report%20of%20the%20Lancet%20Countdown%20on%20health%20and%20climate%20change%22
Die Klimaökonomin Claudia Kemfert hat wegen der Entscheidung des Bundesverfassungsgerichts, die Verwendung von Corona -Rücklagen für den Klima- und Transformationsfonds zu untersagen, von einem schwarzen Tag für den Klimaschutz gesprochen. Sie schlägt vor, den Klimanotstand auszurufen oder fossile Subventionen massiv zu kürzen. https://taz.de/Nach-Karlsruher-Urteil-zum-Bundesetat/!5969938/
Eine neue Studie ergibt, dass sich das Abschmelzen des westantarktischen Eisschilds selbst dann fortsetzen wird, wenn die Erderhitzung auf 1,5° begrenzt wird. Das Schelfeis stellt ein Kipppelememt dar. Der Abschmelzvorgang verstärkt sich selbst und führt zu einer unaufhaltsamen Erhöhung des Meeresspiegels, weil er den Weg für das hinter dem Schelfeis gelegene Gletschereis frei macht. https://www.derstandard.at/story/3000000192327/meterhoher-meeresanstieg-durch-abschmelzen-des-westantarktischen-eisschelfs
Studie: https://www.nature.com/articles/s41558-023-01818-x
Mehr zur Studie: https://hypothes.is/search?q=tag%3A%27report%3A+Unavoidable+future+increase+in+West+Antarctic+ice-shelf+melting%27
Reviewer #1 (Public Review):
Summary: The paper by McGinnis et al. uses a combination of genetic and biochemical approaches to understand how the conserved 5'-3' RNA exonuclease Xrn1 affects autophagy in response to methionine starvation in S. cerevisiae. The authors present evidence Xrn1 affects autophagy primarily via its effect on regulating TORC1 signaling. They present some evidence that Xrn1's effect on TORC1 singnaling is via its physical interaction with the SEACIT complex.
Strengths: The experiments in general for this paper are clear and have proper controls.
Weaknesses:<br /> The authors seem to try and fit the data to a simplistic model rather than embrace the complexity of the data. I will give some examples below.
1) Figure 1 clearly shows that xrn1d results in loss of tight repression of autophagy. Specifically, the 0 timepoint has increased autophagy in both the idh-GFP and ALP assays. However, it is incorrect to say that it is related in any way to methionine deprivation. The same basic pattern of regulation occurs in WT and xrn1d strains. The only difference is the "leakiness" of repression at t=0.
2) Figure 2 shows that catalytically inactive Xrn1 has the same autophagy phenotype as a deletion, indicating that Xrn1 enzymatic activity is important for function. However, it is also clear that xrn1-deletion cells expressing wt Xm1-flag do not repress autophagy as well as XRN+ cells, even though the amount of expressed protein seems similar. Does this imply the flag-tag may be a less active version of the protein? This should be discussed.
3) Figure 3 shows Xrn1-loss effects TORC signaling and that npr2-deletion inhibits autophagy. The surprising result is that a xrn1d/npr2d behaves like WT with regards to autophagy. This needs to be discussed. To me, this seems to strongly suggest that methionine repression of autophagy is occurring downstream of both xrn1 and npr2. Measuring p-S6 in the double mutant may be informative.
4) Figure 4 appears to show that even in the absence of GTR1, autophagy is repressed in rich media, active in YPL-SL, but still responds to methionine repression. This does not seem consistent with the model presented in Figure 5. Shouldn't loss of GTR1 result in repressed Torc1? The GTP and GDP-lock mutants are either all on, or all off. Why is deletion different? This needs to be explained and discussed. Also, the Figure legend does not match figures (problem after Fig4b).
5) Figure 5B shows GTR1 IP with Xrn1-FLAG. However, there are no negative controls in this experiment, so the result could be background. RNAaseA and RNA addition experiments are convincing.
6) Line 254-255. The lead sentence is simply not supported by the data. There is no evidence that Xrn1 actually affects the regulation of Gtr1/2 binding states.
7) Line 259-260. This is again overstated. Just because a mutant can be rescued by Gtr1-GTP-locked, does not say anything about RNA decay. In fact, the double mutant has extra high levels of some ATG RNA's, so I have no idea how the Gtr1 rescues.
8) Line 268-281. Your model here ignores the fact that methionine regulation takes place in the absence of both xrn1 and npr2. Therefore the model, as proposed, can't be correct.
9) Line 290-300. The slow growth rate of Xrn1 mutants may be affecting the metabolite levels. I felt that this entire paragraph was overly speculative.
This is, like THE foundational keystone document to the problem I'm trying to communicate for "builds and burdens".
Author Response
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public Review):
Anderson, Henikoff, Ahmad et al. performed a series of genomics assays to study Drosophila spermatogenesis. Their main approaches include (1) Using two different genetic mutants that arrest male germ cell differentiation at distinct stages, bam and aly mutant, they performed CUT&TAG using H3K4me2, a histone modification for active promoters and enhancers; (2) Using FACS sorted pure spermatocytes, they performed CUT&TAG using antibodies against RNA PolII phosphorylated Ser 2, H4K16ac, H3K9me2, H3K27me3, and ubH2AK118. They also compare these chromatin profiling results with the published single-cell and single-nucleus RNA-seq data. Their analyses are across the genome but the major conclusions are about the chromatin features of the sex chromosomes. For example, the X chromosome is lack of dosage compensation as well as inactivation in spermatocytes, while Y chromosome is activated but enriched with ubH2A in spermatocytes. Overall, this work provides high-quality epigenome data in testes and in purified germ cells. The analyses are very informative to understand and appreciate the dramatic chromatin structure change during spermatogenesis in Drosophila. Some new analyses and a few new experiments are suggested here, which hopefully further take advantage of these data sets and make some results more conclusive.
Major comments: 1. The step-wise accumulation of H3K4me2 in bam, aly and wt testes are interesting. Is it possible to analyse the cis-acting sequences of different groups of genes with distinct H3K4me2 features, in order to examine whether there is any shared motif(s), suggesting common trans-factors that potentially set up the chromatin state for activating gene expression in a sequential manner?
While the histone H3K4me2 mark is low and more widespread at genes active in late spermatocytes and in spermatids (shown in Figure 2C and some examples in Figure 1C-D), we suggest that this may be due to a general decrease in the importance of this modification in late spermatogenesis rather than a specific feature of those genes. We point this out in lines 146-152. This idea is supported by the widespread change in RNAPII distribution in all genes in the germline, shown in Figure 3F and supplementary Figure 2.
- Pg. 4, line 141-142: "we cannot measure H3K4me2 modification at the bam promoter in bam mutant testes or at the aly promoter in aly mutant testes", what are the allelic features of the bam mutant and aly mutant? Are the molecular features of these mutations preventing the detection of H3K4me2 at the endogenous genes' promoters? Also, the references cited (Chen et al., 2011) and (Laktionov et al., 2018) are not the original research papers where these two mutants were characterized.
We have corrected these citations to the original papers. We clarified in the text that the bamΔ86 allele is a deletion of almost all of the coding sequence (reported in Bopp, D., Horabin, J.I., Lersch, R.A., Cline, T.W., Schedl, P. (1993). Expression of the Sex-lethal gene is controlled at multiple levels during Drosophila oogenesis. Development 118(3): 797--812.). The aly1 allele is also a P element-induced mutation; it is not molecularly characterized (it was first described here: Lin, T.Y., Viswanathan, S., Wood, C., Wilson, P.G., Wolf, N., Fuller, M.T. (1996). Coordinate developmental control of the meiotic cell cycle and spermatid differentiation in Drosophila males. Development 122(4): 1331--1341.) We noticed a lack of reads for various histone modifications in aly mutants in part of the gene, suggesting that the deletion is limited to the promoter and the first exon. Signal for the H3K4me2 modification is at background levels for the distal portion of aly, suggesting that the deletion inactivates the gene.
- The original paper that reported the Pc-GFP line and its localization is: Chromosoma 108, 83 (1999).
We are citing the first published description of this marker in the male germline (lines 291-293).
The Pc-GFP is ubiquitously expressed and almost present in all cell types. In Figure 6B, there is no Pc-GFP signals in bam and aly mutant cells.
We apologize, our labeling of the figure was easily overlooked - the bam and aly genotypes do not carry the PcGFP marker, since we didn’t need it for staging the germline nuclei. We have clarified this in the figure.
According to the Method "one testis was dissected", does it mean that only one testis was prepared for immunostaining and imaging? If so, definitely more samples should be used for a more confident conclusion.
We corrected the text to make it clear that all cytological examinations were repeated at least times (lines 438-439).
Also, why use 3rd instar larval testes instead of adult testes?
Generally, we find that immunostaining of the larval testes is cleaner, and we now mention this in the Methods (lines 439-440). We have immunostained both larval and adult testes for these markers with consistent results.
Finally, it is better to compare fixed tissue and live tissue, as the Pc-GFP signal could be lost during fixation and washing steps. Please refer to the above paper [Chromosoma 108, 83 (1999)] for Pc-GFP in spermatogonial cells and Development 138, 2441-2450 (2011) for Pc-GFP localization in aly mutant.
We are using PcGFP staining for staging with antibody detection of other chromatin features, which requires fixed material, although we have compared PcGFP signal in both live and fixed tissue. We have added the 1999 reference for nuclear staging in the male germline.
- Ubiquitinylation of histone H2A is typically associated with gene silencing, here it has been hypothesized that ubH2A contributes to the activation of Y chromosome. This conclusion is strenuous, as it entirely depends on correlative results.
We agree that this is a correlation. We cite in the text examples where uH2A is associated with gene activation. We have added a comment to clarify that this is a correlation (lines 318-320), and now present an alternative that uH2A on the Y chromosome may be moderating expression from these highly active genes (lines 405-407).
For example, the lack of co-localization of ubH2A immunostaining and Pc-GFP are not convincing evidence that ubH2A is not resulting from PRC1 dRing activity. It would be a lot stronger conclusion by using genetic tools to show this. For example, if dRing is knocked down (using RNAi driven by a late-stage germline driver such as bam-Gal4) or mutated in spermatocytes (using mitotic clonal analysis), would they detect changes of ubH2A levels?
We have tested multiple constructs to knockdown dRING using the bam-GAL4 driver although we have not reported it in the manuscript. These knockdowns have no effect on uH2A staining in the testis, on motile sperm production, or on male fertility, although these RNAi constructs do produce Polycomb phenotypes when expressed in somatic cells from an en-GAL4 driver. This is the reason why we point out in the text that there are multiple alternative candidates for an H2A ubiquitin ligase in the Drosophila genome and that in other species RING1 is not responsible for sex body uH2A in the male germline (lines 394-396).
- Regarding "X chromosome of males is thought to be upregulated in early germline cells", it has been shown that male-biased genes are deprived on the X chromosome [Science 299:697-700 (2003); Genome Biol 5:R40 (2004); Nature 450:238-241 (2007)], so are the differentiation genes of spermatogenesis [Cell Research 20:763-783 (2010)]. It would be informative to discuss the X chromatin features identified in this work with these previous findings.
We now mention that the Drosophila X chromosome is moderately depleted of male germline-expressed genes (lines 362-363).
For example, the lack of RNAPII on X chromosome in spermatocytes could be due to a few differentiation genes expressed in spermatocytes located on the X chromosome.
We show in Figure 3B that there is a minor non-significant reduction in RNAPII on the X chromosome in spermatocytes. This small reduction might be due to the moderate paucity of male germline-expressed genes on this chromosome, but since it is non-significant we have not discussed it.
Reviewer #2 (Public Review):
Anderson et al profiled chromatin features, including active chromatin marks, RNA polymerase II distribution, and histone modifications in the sex chromosomes of spermatogenic cells in Drosophila. The results are new and the experiments and analyses look well done, including with appropriate numbers of replicates. Results were parsed by comparing them among two arrest mutants and wildtype, as well as in FACS-sorted spermatocytes. The authors also profiled larval wing discs to serve as reference-somatic cells, which allowed them to focus only on features in their testis data that were associated with germ cells. Their results were further refined by categorizing the genes of interest based on available single nucleus RNA seq expression profiles. The authors document interesting phenomena, such as differences in the distribution of RNAPIIS2p on some genes in germ cells vs somatic cells, the presence of a uH2A body beginning in early spermatocytes, and high levels of uH2A on the Y chromosome and little or none on the X. The former is intriguing because this modification is usually associated with silencing, yet the Y chromosome is active in spermatogenic cells. The authors interpret some of their data as implying a lack of dosage compensation of the X chromosome in spermatocytes.
The data are believable and new, but it is not fully clear how to interpret them. The paper's interpretations rely on subtractive logic to parse results from mixtures of cells down to cell type, extracting spermatogonia, spermatocyte, etc. features by comparing bam mutants (only spermatogonia) to aly mutants (spermatogonia and early spermatocytes but no later stages) to wildtype (all spermatogenic stages), and extracting testis germline data by comparison to wing disc soma; their FACS sorted spermatocytes also have heterogeneity. I recognize that the present paper was a lot of work and am not suggesting that the authors redo their study using methods that give more purity and precision of stage (https://doi.org/10.1126/science.aal3096, https://doi.org/10.1101/gad.335331.119), but they should be aware of them and of their results.
The pulse-release system that the reviewer points to is an interesting system, but more limited in material and in useable markers than the systems we used here. We have added to our discussion of the the limitations of subtractive comparisons between arrest genotypes, both in regards to using mutants that may alter gene expression programs, and to how subtractive comparisons may limit our detection of differences between cell types (lines 143-147).
The conclusions about dosage compensation are indirect, but are consistent with the current model documented in the studies cited by the authors, as well as earlier studies (doi: 10.1186/jbiol30).
We disagree; our data directly speaks to the molecular mechanisms at play. Our profiling of the H4K16acetylation mark and RNAPII in isolated spermatocytes (Figure 4) demonstrates that current models are correct, and so are useful for settling this point in the literature.
Reviewer #1 (Recommendations For The Authors):
Throughout the manuscript, it is better to cite the original research papers.
We have added citations for the original characterizations of bam and aly alleles used, for the descriptions of PCGFP in spermatocytes, and for issues raised by reviewer comments.
Minor comments:
Pg.2, line 70-71: "Germline stem cells at the apical tip of the testis asymmetrically divide to birth spermatogonia", should be gonialblast.
Fixed (line 71).
Pg.2, line 71: "four rapid mitotic divisions", the spermatogonial cell cycle lasts several hours-- "rapid" is subjective and relative, better to leave this word out.
Fixed (line 71).
Reviewer #2 (Recommendations For The Authors):
Other than the major issue raised in the public review this paper only needs a few minor modifications, listed by line number below. The first one would be considered essential by this reviewer.
27: In the sentence that ends on this line, please add the word testis after Drosophila.
Fixed (line 27).
119: It must be known from the Fly Cell Atlas data whether these genes do begin to express in spermatogonia.
Collated expression values from the FCA are provided in Supplementary Table 2. In many cases there is detectable expression of these genes in spermatogonia, although transcript abundance peaks in early spermatocytes.
198: remove "distribution of".
Fixed (line 200).
311: enrichment relative to what?
Fixed (line 313). It is relative to signal in wing discs.
344: other aspects could be regulated such as elongation, termination.
We have added caveats to our speculations in this sentence (lines 340-356). The increased signal we see in gene bodies could be due to slower RNAPII elongation, but we don’t see a way that changes in termination would produce this pattern.
369: This part of the paper seems overly speculative, given the many molecular differences between dosage compensation mechanisms of Drosophila vs mammals, and studies that indicate that MSCI does occur in Drosophila (DOI: 10.3390/genes12111796).
We disagree, and this is a central point in our manuscript. The paper referred to here does not directly assess MSCI in Drosophila, instead they argue that MSCI could be the force driving the evolutionary depletion of male-germline-expressed genes they describe. These and many studies in the literature have conflated the effects of a lack of X dosage compensation and of MSCI in the male germline. Our direct measurements of RNAPII in spermatocytes demonstrates that there is no dosage compensation nor is there MSCI. Further, profiling of histone modifications associated with Drosophila somatic dosage compensation (H4K16ac) or with mammalian MSCI (uH2A, H3K9me2) show that the molecular mechanisms found in these other settings are not in play in the Drosophila male germline. As we have established these biological differences between mammals and Drosophila, it is appropriate to now speculate on why these differences may be, which we do on lines 374-384.
(several lines): Can the authors justify their assumption that chromatin features of larval wing disc cells will match those of somatic cells of adult testes?
We don’t only compare germline features to somatic cells of the wing disc, but also to genes with somatic expression in the testes annotated by FCA expression data (H3K4me2 in Figure 2C, RNAPII in Figure 3F). Note in Supplementary Figure 2 the distribution of RNAPII in whole testes (which includes somatic cells) is similar to that of larval wing discs, confirming that the differences we describe are specific to germline cells.
Reviewer #1 (Public Review):
Anderson, Henikoff and Ahmad et al. performed a series of genomics assays to study Drosophila spermatogenesis. Their main approaches include (1) Using two different genetic mutants that arrest male germ cell differentiation at distinct stages, bam and aly mutant, they performed CUT&TAG using H3K4me2, a histone modification for active promoters and enhancers; (2) Using FACS sorted pure spermatocytes, they performed CUT&TAG using antibodies against RNA PolII phosphorylated Ser 2, H4K16ac, H3K9me2, H3K27me3, and ubH2AK118. They also compare these chromatin profiling results with the published single-cell and single-nucleus RNA-seq data. Their analyses are across the genome but the major conclusions are about the chromatin features of the sex chromosomes. For example, the X chromosome is lack of dosage compensation as well as inactivation in spermatocytes, while Y chromosome is activated but enriched with ubH2A in spermatocytes. Overall, this work provides high quality epigenome data in testes and in purified germ cells. The analyses are very informative to understand and appreciate the dramatic chromatin structure change during spermatogenesis in Drosophila.
What a great tour the summary can be found in the timestamps for the video, but I'd like to point out a few things. Through Maggie's daily tag. She has a field for what's on her mind. This makes it really easy to scan through the list and see how that's changed. Over previous weeks, we show an advanced way to set it up, which is how she originally set it up as well as a simpler way to capture similar benefits.
What a nice feature
Reviewer #3 (Public Review):
Summary:<br /> This study aims to investigate the stoichiometric effect between core factors and partners forming the heterodimeric transcription factor network in living cells at endogenous expression levels. Using state-of-the-art single-molecule analysis techniques, the authors tracked individual RARα and RXRα molecules labeled by HALO-tag knock-in. They discovered an asymmetric response to the overexpression of counter-partners. Specifically, the fact that an increase in RARα did not lead to an increase in RXRα chromatin binding is incompatible with the previous competitive core model. Furthermore, by using a technique that visualizes only molecules proximal to partners, they directly linked transcription factor heterodimerization to chromatin binding.
Strengths:<br /> The carefully designed experiments, from knock-in cell constructions to single-molecule imaging analysis, strengthen the evidence of the stoichiometric perturbation response of endogenous proteins. The novel finding that RXR, previously thought to be a target of competition among partners, is in excess provides new insight into key factors in dimerization network regulation. By combining the cutting-edge single-molecule imaging analysis with the technique for detecting interactions developed by the authors' group, they have directly illustrated the relationship between the physical interactions of dimeric transcription factors and chromatin binding. This has enabled interaction analysis in live cells that was challenging in single-molecule imaging, proving it is a powerful tool for studying endogenous proteins.
Weaknesses:<br /> As the authors have mentioned, they have not investigated the effects of other T2NRs or RXR isoforms. These invisible factors leave room for interpretation regarding the origin of chromatin binding of endogenous proteins (Recommendations 4). In the PAPA experiments, overexpressed factors are visualized, but changes in chromatin binding of endogenous proteins due to interactions with the overexpressed proteins have not been investigated. This might be tested by reversing the fluorescent ligands for the Sender and Receiver. Additionally, the PAPA experiments are likely to be strengthened by control experiments (Recommendations 5).
Die WHO will bei der kommenden COP28 das Thema der Gesundheit offensiv in den Mittelpunkt stellen. Zum ersten Mal wird ein ganzer Tag Gesundheitsthemen gewidmet sein. Der Guardian hat dazu mit Maria Neira. der WHO-Verantwortlichen für Umweltgesundheit, gesprochen. https://www.theguardian.com/world/2023/nov/04/politicians-who-delay-climate-action-must-live-with-consequences-says-who-expert
Non-aims (but may happen anyway):
Author Response
The following is the authors’ response to the original reviews.
Public Reviews:
Reviewer #1 (Public Review):
Summary: Hansen et al. dissect the molecular mechanisms of bacterial ice nucleating proteins mutating the protein systematically. They assay the ice nucleating ability for variants changing the R-coils as well as the coil capping motifs. The ice nucleation mechanism depends on the integrity of the R-coils, without which the multimerization and formation of fibrils are disrupted.
Strengths: The effects of mutations are really dramatic, so there is no doubt about the effect. The variants tested are logical and progressively advance the story. The authors identify an underlying mechanism involving multimerization, which is plausible and compatible with EM data. The model is further shown to work in cells by tomography.
Weaknesses: The theoretical model presented for how the proteins assemble into fibrils is simple, but not supported by much data.
Agreed. This theoretical INP multimer model was introduced to promote discussion and elicit ideas on how to prove or disprove it. The length and width of the fibres are defined by cryo-ET results, in which the narrow width is just sufficient to accommodate a dimer of the INPs, and the long length requires that several INPs are joined end to end. Their antiparallel arrangement produces identical ends to the dimer and avoids steric clash of the C-terminal cap structures as well as the C-terminal GFP tag. This model can accommodate the wide range of INPs lengths seen in nature (due to different numbers of water-organizing coils) and introduced in mutagenesis experiments (Forbes et al. 2022). It defines a critical role for the R-coil subdomain in joining the dimers together and explains why this region cannot be shortened by more than a few coils either in nature or by experimentation.
In response to specific criticisms of the model (Fig. 9), we have redesigned this to be less schematic and to incorporate several copies of the AlphaFold-predicted structure.
Reviewer #2 (Public Review):
Summary:
This paper further investigates the role of self-assembly of ice-binding bacterial proteins in promoting ice-nucleation. For the P. borealis Ice Nucleating Protein (PbINP) studied here, earlier work had already determined clearly distinct roles for different subdomains of the protein in determining activity. Key players are the water-organizing loops (WO-loops) of the central beta-solenoid structure and a set of non-water-organizing C-terminal loops, called the R-loops in view of characteristically located arginines. Previous mutation studies (using nucleation activity as a read-out) had already suggested the R-loops interact with the WO loops, to cause self-assembly of PbINP, which in turn was thought to lead to enhanced ice-nucleating activity. In this paper, the activities of additional mutants are studied, and a bioinformatics analysis on the statistics of the number of WO- and R-loops is presented for a wide range of bacterial ice-nucleating proteins, and additional electron-microscopy results are presented on fibrils formed by the non-mutated PbINP in E coli lysates.
Strengths:
-A very complete set of additional mutants is investigated to further strengthen the earlier hypothesis.
-A nice bioinformatics analysis that underscores that the hypothesis should apply not only to PbINP but to a wide range of (related) bacterial ice-nucleating proteins.
-Convincing data that PbINP overexpressed in E coli forms fibrils (electron microscopy on E coli lysates).
Weaknesses:
-The new data is interesting and further strengthens the hypotheses put forward in the earlier work. However, just as in the earlier work, the proof for the link between self-assembly and ice-nucleation remains indirect. Assembly into fibrils is shown for E coli lysates expressing non-mutated pbINP, hence it is indeed clear that pbINP self-associates. It is not shown however that the mutations that lead to loss of ice-nucleating activity also lead to loss of self-assembly. A more quantitative or additional self-assembly assay could shine light on this, either in the present or in future studies.
The control cryo-ET experiment where the R-coils were deleted and INP fibres were not seen is consistent with a link between the loss of ice-nucleating activity and the loss of self-assembly. However, we agree that a more direct measurement of the physical state of INP molecules is needed to prove the link.
-Also the "working model" for the self-assembly of the fibers remains not more than that, just as in the earlier papers, since the mutation-activity relationship does not contain enough information to build a good structural model. Again, a better model would require different kinds of experiments, that yield more detailed structural data on the fibrils.
Reviewer #1 also raised these criticisms of the model, which we have responded to (above). Testing the model is a focus of our continuing experiments on INPs.
Reviewer #3 (Public Review):
Summary: in this manuscript, Hansen and co-authors investigated the role of R-coils in the multimerization and ice nucleation activity of PbINP, an ice nucleation protein identified in Pseudomonas borealis. The results of this work suggest that the length, localization, and amino acid composition of R-coils are crucial for the formation of PbINP multimers.
Strengths: The authors use a rational mutagenesis approach to identify the role of the length, localisation, and amino acid composition of R-coils in ice nucleation activity. Based on these results, the authors hypothesize a multimerization model. Overall, this is a multidisciplinary work that provides new insights into the molecular mechanisms underlying ice nucleation activity.
Weaknesses: Several parts of the work appear cryptic and unsuitable for non-expert readers. The results of this work should be better described and presented.
In revising the manuscript for reposting we have rewritten sections to make it more accessible to the non-expert. Incorporating the detailed recommendations of the reviewers has been helpful in this effort.
Recommendations for the authors: please note that you control which revisions to undertake from the public reviews and recommendations for the authors
Reviewer #1 (Recommendations For The Authors):
Introduction: Curiously, there is no mention at all in the introduction of what the biological function of these ice-nucleating proteins is.
We added the following text to the first paragraph of the Introduction: ”INP-producing bacteria are widespread in the environment where they are responsible for initiating frost (4) and atmospheric precipitation (5). As such, these bacteria play a significant role in the Earth’s hydrological cycle and in agricultural productivity.”
Line 70: TXT, SLT, and Y motifs are mentioned, but only the first is described. Also, TXT name alternates between TXT and TxT in the manuscript. (I think the latter is more correct).
These putative water-organizing motifs are introduced in the preceding paper (new ref 8). We now use TxT consistently throughout the manuscript and have converted SLT to SxT because L is an inward-pointing residue that is not directly involved in water organization.
Line 236: A construct with repeats deleted is tested for thermostability, but it is not really explained what hypothesis this experiment is supposed to test.
This is an observation that adds information about the stability of the INP multimers and will need to be explained by the structure.
Line 267: The authors test a mutant where the N-terminal coil is disrupted and find a big effect. Nevertheless, no conclusion is drawn. What does this result mean?
On the contrary, INP activity is not appreciably affected by N-terminal deletion.
Line 269: The CryoEM begins rather abruptly with technical details. Consider introducing the paragraph with a brief statement about what you want to investigate. Also, the analysis seems a little half-hearted.
Given that the authors describe other EM studies of fibrils of the same protein it would be nice with a clear statement about what is new in their study and how it compares to previous studies.
We have added this statement about why we used Cryo-EM: “The idea that INPs must assemble into larger structures to be effective at ice nucleation has persisted since their discovery (6). In the interim the resolving power of cryo-EM has immensely improved. Here we elected to use cryo-electron tomography to view the INP multimers in situ and avoid any perturbation of their superstructure during isolation.”
Fig. 7B: Single-letter amino acid codes are always capitalized.
We have revised this figure to use capital letters for the amino acids.
Fig. 9: This figure is really hard to read even though it is very simplistic. I would consider making a figure with several copies of the AlphaFold model instead. Especially panel D, I do not know what is supposed to show.
We have followed this advice and have completely revised the figure using copies of the AlphaFold model. Panel D (now C) shows two cross-sections through the AlphaFold model.
Line 355 onwards: The model of the INP is the weakest part of the manuscript. This reviewer considers that the model is crude and it is unclear what information the model is supported by. The authors might want to consider running an AlphaFold multimer to get a better model of at least the dimer.
Our objective now is to validate or disprove the model by experimentation using protein-protein cross-linking in conjunction with mass spectrometry, and higher resolution cryo-EM methods.
Reviewer #2 (Recommendations For The Authors):
I would suggest more frankly discussing the weaknesses mentioned in my public review, as well as approaches that could be used in the future to address these.
In the cryo-ET analysis, INP mutations of the R-coils that lead to loss of ice-nucleating activity fail to show fibres in the bacteria (Fig. S4), which is consistent with the loss of self-assembly. We are working on physical methods that can assess the degree of assembly of the different INP constructs and mutations. We are working to validate and improve the working model of INP multimers.
Reviewer #3 (Recommendations For The Authors):
Abstract
Line 18. Below 0 Celsius should be < 0 {degree sign}C.
Done
Line 25. E. coli should be Escherichia coli
Done
Line 29. E. coli should be in italics.
Done
Introduction
The introduction is weak and not suitable for non-expert readers. Moreover, in some parts it is cryptic and it is not clear whether the authors are describing INP in general or PbINP. The introduction should be reorganized to highlight the novelty of this paper compared to Forbes et al. 2022.
The changes we have made to the Introduction can be seen in the ‘documents compared’ version where the changes are tracked.
Line 45. It is unclear whether this paragraph is a result reported in the literature or the result of this work. Please clarify.
These are results reported in the literature as indicated by the references cited in the paragraph.
Line 54. It is not clear whether this paragraph describes PbINP or INP in general.
This paragraph begins with INPs in general and then focuses on PbINP.
Results
Line 109. This section would benefit from a paragraph in which the authors describe the rationale for this bioinformatic analysis.
We added the following Statement: “A bioinformatic analysis of bacterial INPs was undertaken to identify their variations in size and sequence to understand what is common to all that could guide experiments to probe higher order structure and help develop a collective model of the INP multimer.”
Some information is needed on the selected sequences such as sequence identity, what do the authors mean by nr database?
The abbreviation nr has been replaced by ‘non-redundant’. As explained in that same paragraph the sequences selected were those from long-read sequences that could be relied on to accurately count the number of solenoid coils.
Line 144. The standard deviation is necessary to understand whether these differences are statistically significant.
These have been added as p values.
Figure 2. I noticed that the authors used GFP-tagged PbINP. Why? In addition, panel C is never mentioned in the manuscript.
The GFP tag was used to confirm expression of the PbINP in E. coli. We have added this sentence: “As previously described these constructs were tagged with GFP as an internal control for INP production, and its addition had no measured effect on ice nucleation activity (8).”The GFP tag was also useful as in internal control for the heat denaturation experiments featured in Fig. 6, where it lost its fluorescence between 65 and 75 °C.
Fig. 2C is now cited alongside Fig. 2B.
Figure 3. In my opinion, the results of the R-coil deletion should also be shown in Figure 2. Line 171. This section is cryptic. A logo sequence or an alignment of WO-coils and R-coils of PbINP could be helpful for the reader. Instead of the architecture of the whole protein, it would be useful to have the sequence of the R-coils with the residues that the authors mutagenised.
The logo sequences are available in Fig. 1.
Line 202. Here, the authors describe a new experimental setup. As the Materials and Methods section follows the Discussion, the authors should state in the first paragraph of the Results section that IN activity was measured on whole cells.
We have now modified the introductory sentence to read: “Ice nucleation assays were performed on intact E. coli expressing PbINP to assess the activity of the incremental replacement mutants.”
Line 202. The authors investigated the effects of pH and temperature (Line 223) on the IN activity. The authors should better introduce the rationale for these experiments and how they fit within the work.
We have now modified the following sentence to provide the rationale: “To see how important electrostatic interactions were in the multimerization of PbINP as reflected by its ice nucleation activity, it was necessary to lyse the E. coli to change the pH surrounding the INP multimers.”
Line 245. This work is supported by a model provided by Alphafold. I wonder how reliable this model is; the authors should indicate the quality of the model and provide the accuracy values of the residuals.
This information is now provided in Figure S1.
Line 259. Typically in mutagenesis studies, a key residue is substituted with alanine to create a loss of function variant. In this case, the authors have made the following substitutions F1204D, D1208L, and Y1230D, it is not clear to me why the authors have replaced an aromatic residue with one of aspartic acid that is negatively charged.
We have justified these more extreme changes as follows: “For an enhanced effect of the mutations hydrophobic residues were replaced with charged ones and vice versa.”
Line 269. This paragraph seems completely unrelated to the section entitled: The β-solenoid of INPs is stabilized by a capping structure at the C terminus, but not at the N terminus.
We had omitted the sub-heading “Cryo-electron tomography reveals INPs multimers form bundled fibres in recombinant cells”, which is now in place.
Discussion
Overall, the discussion is too long and some parts appear cryptic, this section should be reorganized.
The changes we have made to the Discussion can be seen in the ‘documents compared’ version where the changes are tracked.
Line 354. It is not clear what experimental evidence supports this model. In the results, this model is never mentioned and it is not clear whether it was obtained by computational analysis or not.
The model is presented in the Discussion because it was not arrived at by experimentation but is an attempt to integrate the observations made in the Results section. The experimental evidence that supports this model is reviewed in the Discussion section: “Working model of the INP multimer is consistent with the properties of INPs and their multimers.”
Line 354. The authors used GFP-tagged PbINP. The Authors should discuss the role of GFP in this model and IN activity. A measurement of IN activity on PbINP without GFP would be useful.
We have previously shown in Ref 8 that the GFP tag has no detrimental effect on ice nucleation activity. Our model for the INP multimer can accommodate this C-terminal tag without any steric hindrance.
Line 364. The Authors hypothesize that electrostatic interactions stabilize end-to-end dimer associations. To test this hypothesis, the authors should measure the activity of IN at increasing concentrations of NaCl. It is known that high salt concentrations shield charges by preventing the formation of electrostatic intermolecular interactions.
We have added this sentence to the Discussion: “Another useful test of the electrostatic component to the multimer model would be to study the effects of increasing salt concentration on ice nucleation activity of the E. coli extracts.”
Line 439. Conclusions should be useful for the reader.
Material and Methods
In several sections, the authors refer to what has already been published in Forbes et al. However, the minimum information should also be described in this work. In addition, the Authors should indicate the number of replicates.
The ice nucleation assays on whole cells were done on the WISDOM apparatus, which integrates 100’s of individual measurements to obtain a T50 value. These T50 values were confirmed by assays on the nanoliter osmometer apparatus. The numbers of replicates used on the nanoliter osmometer apparatus are indicated by box and whisker plots in Figs. 5 & 6 with boxes and bars showing quartiles, with medians indicated by a centre line.
Line 500. This paragraph should be removed as the results are not described in the manuscript.
This is a Methods section that describes how that INPs were expression in E. coli. It has details that are important for researchers who want to repeat our findings, such as the use of the Arctic Express strain for producing INP.
Author Response
The following is the authors’ response to the original reviews.
Reviewer #1 (Public Review):
"MAGIC" was introduced by the Rong Li lab in a Nature letters article in 2017. This manuscript is an extension of this original work and uses a genome wide screen the Baker's yeast to decipher which cellular pathways influence MAGIC. Overall, this manuscript is a logical extension of the 2017 study, however the manuscript is challenging to follow, complicated by the data often being discussed out of sequence. Although the manuscripts make claims of a mechanism being pinpointed, there are many gaps and the true mechanisms of how the factors identified in the screen influence MAGIC is not clear. A key issue is that there are many assumptions drawn on previous literature, but central aspects of the mechanisms being proposed are not adequately shown.
Key comments:
- Reasoning and pipelines presented in the first two sections of the results are disordered and do not follow figure order. In some instances, the background to experimental analyses such as detailing the generation of spGFP constructs in the YKO mutant library, or validation of Snf1 activation are mentioned after respective results are discussed. This needs to be fixed.
We thank the reviewer for pointing out potential confusion to readers. We have revised the first two sections according to reviewer’s suggestion. (Page 4-6)
- In general there is a lack of data to support microscopy data and supporting quantification analysis. The validity of this data could be significantly strengthened with accompanying western blots showing accumulation of a given constructs in mitochondrial sub compartments (as was the case in the lab’s original paper in 2017).
We appreciate the reviewer’s suggestion on biochemical validations. However, the validity of this imaging-based assay for detecting import of cytosolic misfolded proteins into mitochondria, including the use of FlucSM as a model misfolding-prone protein, was carefully established in our previous study by using appropriate controls, super resolution imaging, APEX-based proximity labeling, and classical biochemical fractionation and protease protection assay (Ruan et al., 2017 Nature, ref. 10). We have reminded readers of these validation experiments in the previous study on Page 4, line 14-17.
In recent years, advancements in imaging-based tools have allowed many protein interactions and dynamic processes, which were previously examined by using biochemical assays in lysates of populations of cells, to be observed with various level of quantitation in live cells with intact cellular compartments. Many of these assays, e.g., the RUSH assay for ER to Golgi transport, FRAP-based analysis for nuclear/cytoplasmic shuttling of proteins, or FRET-based assays for protein-protein interactions, have been well accepted and even embraced by the respective fields of study once validated with genetic and biochemical approaches. The advantages for live-cell imaging-based assays are often their unique ability to report dynamic processes or unstable molecular species with spatiotemporal sensitivity. Respectfully, it is our view, based on our own experience, that the traditional protease protection assay is not adequate or sufficiently quantitative for examining the presence of unstable misfolded proteins in mitochondrial sub-compartments, given the obligatorily lengthy in vitro cell lysis and mitochondrial isolation process, during which the unstable proteins are continuously being degraded. This likely explains our previous biochemical fractionation result that only weak protein signals were detected in the matrix fraction (Ruan et al., 2017 Nature, ref. 10). In addition, unlike stably folded, native mitochondrial matrix proteins, misfolded/unfolded proteins such as Lsg1 or FlucSM are highly susceptible to protease treatment. This sensitivity makes the assay unreliable for detecting such proteins if trace amount of the protease penetrates mitochondrial membranes during cell lysis even without detergent treatment.
While we agree that protease protection assay is highly valuable for qualitative detection of the presence of a protein in certain mitochondrial compartments or determining its topology on membranes, this assay (regrettably in our hands) does not allow quantitative comparisons that were necessary for this study, because of inherent sample to sample variation, yet the laborious and low throughput nature of this assay makes it difficult for adequate statistical analysis. Furthermore, the level of protein detection in various fractions is highly sensitive to how the sample is treated with protease and detergent. Our imaging-based quantification, on the other hand, allows us to compare increased or decreased presence of GFP11-tagged proteins in mitochondria under different metabolic conditions or in different mutant or wild-type strains. Data from hundreds of cells and at least three independent biological replicates allowed us to apply adequate statistical analysis to aid our conclusion.
- Much of the mechanisms proposed relies on the Snf1 activation. This is however not shown but assumed to be taking place. Given that this activation is central to the mechanism proposed, this should be explicitly shown here - for example survey the phosphorylation status of the protein.
Both REG1 deletion and low glucose conditions have been demonstrated extensively for Snf1 phosphorylation and activation in yeast (e.g., many seminal papers from Marian Carlson’s and other lab, such as ref. 24-28). In our study, we have indeed corroborated this by showing that Mig1 was exported from the nucleus in Δreg1 mutant and in low glucose conditions (Figure 1—figure supplement 2H and I. The mechanism of Snf1-mediated nuclear export of Mig1 has been characterized in detail as well (e.g., ref. 29-31).
Recommendations for the authors: please note that you control which, if any, revisions, to undertake
Reviewer #1 (Recommendations For The Authors):
SPECIFIC COMMENTS
Genetic Screen o Line 20 - the narrative moves to SNF1, but the reasoning for the selection of this Class I substrate is not defined. What was the basis for this selection - what happened to the other Class I substrates. It is stated in the text that the other Class I proteins show the same increase in spGFP signal. The data showing this should be included in the Supp Figure 1 for transparency.
We have moved the narratives of Snf1 function to the second section and clarified that we were interested in this gene due to its central role in metabolism and mitochondrial functions that may influence MAGIC (Page 5: line 16-20). Other genes in class 1 were shown in Table S1. Detailed discussion of other genes in this category is beyond the scope of this study.
Snf1/AMPK prevents MP accumulation in mitochondria:
The FlucDM data in human RPE-1 mitochondria seems to be added to only increase the significance of the work. The mechanisms suggested here with Hap4 would not be possible in human cells as there is no homologue of this protein in human cells. Making generalisations that these pathways are conserved based on this one experiment is not appropriate.
We appreciate this feedback. Although the focus of this study is the regulation of MAGIC by the yeast AMPK Snf1, we would like to share our initial observation that suggests a similar role of AMPK in human RPE-1 cells. We acknowledge that the underlying mechanisms regarding the downstream transcription factors and pathway for misfolded protein import could be different in mammalian cells, but the overall effect of AMPK in mitochondrial biogenesis is well known to resemble that of Snf1. To avoid making over-generalization, we changed our statement of conclusion to: ‘These results suggest that AMPK in human cells regulates MP accumulation in mitochondria following a similar trend as in yeast, although the underlying mechanisms might differ between these organisms.’ (Page 7: line 2-4)
Mechanisms of MAGIC regulation by Snf1:
While the lysosome is ruled out here the authors have not considered the proteasomes. Is there a reason for this? Given accumulation of aggregates outside of mitochondria, and previous connections of the proteasome to mitochondrial quality control this would be an obvious thing to check. We examined the role of lysosomal degradation here because it is known to be activated under Snf1active condition (ref. 37). We appreciate this feedback and have included a new analysis on MG132treated FlucSM spGFP strains in which PDR5 gene was deleted to avoid drug efflux.
This result suggests that the proteosome inhibitor did not ablate the difference in FlucSM accumulation between these conditions. That MG132 promoted mitochondrial accumulation of FlucSM in both high glucose and low glucose conditions was not surprising, as FlucSM is also degraded by proteasome in the cytosol (Ruan et al., 2017 Nature, ref. 10), and preventing this pathway could divert more of such protein molecules toward MAGIC. (Page 7: line 26-29).
Line 13 "we hypothesized that elevated expression of mitochondrial preproteins induced by the activation of Snf1-Hap4 axis (REF) may outcompete MPs for import channels". This statement has some assumptions. The authors have not shown that Snf1 is activated in thier models and more importantly that they have an accumulation of mitochondrial preproteins. The data that follows using the cytosolic domains of the receptors is hard to rationalise without seeing evidence that there is in fact pre-protein accumulation or impacts on the mitochondrial proteome in this system.
As stated in our response to main point [3], Snf1 activation in reg1 mutant or in low glucose is evidenced by our data showing Mig1 export from nucleus to cytoplasm and had also been shown in many previous publications. A recent study (Tsuboi et al., 2020 eLife) also showed a dramatic increase in mitochondrial volume fraction in Δreg1 cells and wild-type cells in respiratory conditions, further supporting the role of Snf1 in mitochondrial biogenesis. We have provided relevant references in the manuscript (ref. 24-28).
The ability of Tom70 cytosolic domain (Tom70cd), which can bind mitochondrial preproteins but not localize to mitochondria due to lack of N-terminal targeting sequence, to compete with endogenous Tom70 for mitochondrial preproteins has been well documented (ref. 47-49). However, we agree with the reviewer that a future quantitative proteomics study to measure changes in mitochondrial proteome under Tom70cd over-expression could allow more accurate interpretation of our experimental result.
AMPK protects cellular fitness during proteotoxic stress:
The inhibition of preprotein import by overexpressing the cytosolic domains of receptors is not supported with some proof of principle data. If this was working as the authors assume, it is not clear why only an effect with Tom70 is observed. The majority of the mitochondrial proteome is imported via Tom20/Tom22 so this does not align with what the authors are suggesting. Is the Tom70CD and any associated Hsp proteins facilitating the observed changes to the MPs?
We thank the reviewer for raising this point. We expressed different TOM receptor cytosolic domains but found that Tom70cd had the strongest rescue on MAGIC under AMPK activation conditions. It is possible that certain Tom70 substrates or Tom70-assoicated heat shock proteins inhibit the import of MAGIC substrates. We admit that a clear explanation of this unexpected observation necessitates a better understanding of how native and MAGIC substrates are selected and imported by the outer-membrane channel. We can only offer our best interpretation based on the current state of the understanding, and we feel that we have been careful to acknowledge such in the manuscript.
While the effect of AMPK inactivation reducing FUS accumulation was striking, this was all in the context of overexpression and may not be physiologically relevant - or may occur very transiently under basal conditions. Is GST an appropriate control here, why not use WT FUS? Likewise, one representative image is shown in Figure 5 - can the authors show western blotting that mitochondrial accumulation of FUS can be reduced with AMPK activation?
We thank the reviewer for this suggestion, however, overexpressed FUS WT is also aggregation prone (Zhihui Sun et al., 2011, PloS Biology; Shulin Ju, 2011, PloS Biology; Jacqueline C. Mitchell et., 2013, Acta Neuro). We believe that GST, as a well-folded protein, is an appropriate control (Ruan et al., 2017 Nature, ref. 10). As we discussed in response to main point [1], the in vitro assay involving protease protection and western blots do not allow reliable quantitative comparison in our hands.
In text changes.
The analysis pipeline of the YKO mutant library should be introduced at the very start of the first paragraph, not the end.
Addressed on Page 4, second paragraph
"Fluc" should be introduced as "Firefly luciferase" within the first paragraph of the first section, also need to define SM and DM in FlucSM/FlucDM - these appear to be missing.
Addressed in both Introduction (Page 2: line 29; Page 3: line 8-9) and re-clarified in Result (Page 5: line 27-29)
The role of Reg1 should be explicitly stated in the text, not just in the figure.
Addressed on Page 6: line 3-6
Figure 1H legend states Reg1 (WT) is Snf1-inactive and Reg1 KO is Snf1-active. This wording is confusing and is not supported by data, but by assumption. If the authors want to use this wording then evidence needs to be provided - as suggested above.
We have changed this and other legends to only show genotypes and medium conditions.
"Tom70cd overexpression also exacerbated growth rate reduction due to FlucSM expression in HG medium (Figure 4A; Figure 4 - figure supplement 1A)" should be figure supplement 1B.
Fixed on Page 10: line 10
"These results suggest that glucose limitation protects mitochondria and cellular fitness during FlucSM induced proteotoxic stress through Snf1-dependent inhibition of MP import into mitochondria". The phrase "Snf1-dependent inhibition of MP import into mitochondria" may be misleading, as Snf1 isn't modulating import directly but is acting on transcriptional regulators to modulate mitochondrial import under stress.
We restated the conclusion as follows: ‘These results suggest that Snf1 activation under glucose limitation protects mitochondrial and cellular fitness under FlucSM-associated proteotoxic stress.’ (Page 10: line 20- 21)
"... Significantly increased the fraction of spGFP-positive and MMP-low cells in both HG and LG medium (Figure 4G-K)" should be (Figure 4J-K).
Fixed on Page 11: line 3
Reviewer #2 (Public Review):
Work of Rong Li´s lab, published in Nature 2017 (Ruan et al, 2017), led the authors to suggest that the mitochondrial protein import machinery removes misfolded/aggregated proteins from the cytosol and transports them to the mitochondrial matrix, where they are degraded by Pim1, the yeast Lon protease. The process was named mitochondria as guardian in cytosol (MAGIC).
The mechanism by which MAGIC selects proteins lacking mitochondrial targeting information, and the mechanism which allows misfolded proteins to cross the mitochondrial membranes remained, however, enigmatic. Up to my knowledge, additional support of MAGIC has not been published. Due to that, MAGIC is briefly mentioned in relevant reviews (it is a very interesting possibility!), however, the process is mentioned as a "proposal" (Andreasson et al, 2019) or is referred to require "further investigation to define its relevance for cellular protein homeostasis (proteostasis)" (Pfanner et al, 2019).
Rong Li´s lab now presents a follow-up story. As in the original Nature paper, the major findings are based on in vivo localization studies in yeast. The authors employ an aggregation prone, artificial luciferase construct (FlucSM), in a classical split-GFP assay: GFP1-10 is targeted to the matrix of mitochondria by fusion with the mitochondrial protein Grx5, while GFP11 is fused to FlucSM, lacking mitochondrial targeting information. In addition the authors perform a genetic screen, based on a similar assay, however, using the cytosolic misfolding-prone protein Lsg1 as a read-out.
My major concern about the manuscript is that it does not provide additional information which helps to understand how specifically aggregated cytosolic proteins, lacking a mitochondrial targeting signal could be imported into mitochondria. As it stands, I am not convinced that the observed FlucSM-/Lsg1-GFP signals presented in this study originate from FlucSM-/Lsg1-GFP localized inside of the mitochondrial matrix. The conclusions drawn by the authors in the current manuscript, however, rely on this single approach.
In the 2017 paper the authors state: "... we speculate that protein aggregates engaged with mitochondria via interaction with import receptors such as Tom70, leading to import of aggregate proteins followed by degradation by mitochondrial proteases such as Pim1." Based on the new data shown in this manuscript the authors now conclude "that MP (misfolded protein) import does not use Tom70/Tom71 as obligatory receptors." The new data presented do not provide a conclusive alternative. More experiments are required to draw a conclusion.
In my view: to confirm that MAGIC does indeed result in import of aggregated cytosolic proteins into the mitochondrial matrix, a second, independent approach is needed. My suggestion is to isolate mitochondria from a strain expressing FlucSM-GFP and perform protease protection assays, which are well established to demonstrate matrix localization of mitochondrial proteins. In case the authors are not equipped to do these experiments I feel that a collaboration with one of the excellent mitochondrial labs in the US might help the MAGIC pathway to become established.
We thank Reviewer 2 for these suggestions, but we would like to respectfully offer our difference in opinion:
a. Regarding the suggestion “to isolate mitochondria from a strain expressing FlucSM-GFP and perform protease protection assays”, in our previous study (Ruan et al., 2017 Nature, ref. 10), we have indeed applied two independent biochemical approaches: APEX-mitochondrial matrix proximity labeling and classic protease protection assay using non-spGFP strains, both consistently confirmed the entry of misfolded proteins into mitochondria under proteotoxic stress. Our super-resolution imaging further confirmed the import of the split GFP-labeled proteins to be inside mitochondria. Moreover, as we discussed in response to Reviewer 1’s main point [2], while the suggested biochemical assay is useful for validating topology within mitochondria, it is not quantitative and may not reliably report the in vivo accumulation of misfolded proteins in mitochondria due to the isolation process that takes hours, during which the unstable proteins could be continuously degraded within mitochondria.
While we agree with the reviewer that we do not yet understand how misfolded proteins are imported into mitochondria, it would be unfair to state “as it stands, I am not convinced..” simply because the underlying mechanism remains to be elucidated. We would like to point out that targeting sequences for many well-established mitochondrial proteins are still not well defined. It is well known that mitochondrial targeting sequences are not as uniformly predictable as, for example, nuclear targeting sequences. Our finding that deletion of TOM6 enhances the import of misfolded proteins suggest that their import may involve the TOM channel in a more promiscuous conformation, which may reduce the requirement for a specific sequence-based targeting signal associated with the substrate.
b. Regarding the role of Tom70, in our 2017 study, using proteomics and subsequently immunoprecipitation we validated the binding, albeit not necessarily direct, between misfolded protein FlucSM and Tom70. Therefore, “we speculate that protein aggregates engaged with mitochondria via interaction with import receptors such as Tom70”. Recent studies from different labs confirmed the interactions between Tom70 and aggregation prone proteins (Backes et al., 2021, Cell Reports; Liu et al., 2023, PNAS). In the current study, surprisingly, knockout of TOM70 did not block MAGIC, suggesting redundant components of mitochondria import system may facilitate the recruitment of misfolded proteins in the absence of Tom70, and this does not contradict the notion that Tom70 helps tether protein aggregates to mitochondria.
c. Regarding other studies also showing the import of misfolding or aggregation-prone cytosolic proteins into mitochondria, there have been at least several recent studies in the literature for mammalian cells involving either model substrates or disease proteins (e.g., ref. 12-15; 56-58; Vicario, M. et al. 2019 Cell Death Dis.). The studies are briefly mentioned in Introduction (Page 3, paragraph 2). The present manuscript documents a major effort from our group using whole genome screen in yeast to understand the mechanism and regulation of MAGIC. Many of the screen hits have yet to be studied in detail. We full agree that much remains to be understood about whether and how this pathway affects proteostasis and what might be the evolutionary origin for such a mechanism.
Additional comments:
The genetic screen:
The genetic screen identified five class 1 deletion strains, which lead to enhanced accumulation of Lsg1GFP and a larger set of class 2 mutants, which lead to reduced accumulation. Please note, in my opinion it is not clear that accumulation of the reporters occurs inside the mitochondria. In any case, the authors selected one single protein for further analysis: Snf1, the catalytic subunit of the yeast SNF complex, which is required for respiratory growth of yeast.
The results of the screen are not discussed in any detail. The authors mention that ribosome biogenesis factors are abundant among class 2 mutants. Noteworthy, Lsg1 is involved in 60S ribosomal subunit biogenesis. As Lsg1-GFP11 is overexpressed in the screen this should be discussed. Class 2 mutants also .include several 40S ribosomal subunit proteins (only one of the 60S subunit). What does this imply for the MAGIC model? Also, it should be discussed that the screen did not identify reg1 and hap4, which I had expected as hits based on the data shown in later parts of the manuscript.
We apologize for the confusion, but the GFP11 tag was in fact knocked into the C-terminus of Lsg1 in the endogenous LSG1 locus, and so Lsg1 was not overexpressed in the screen. We have made sure that this information is clearly conveyed in the revised manuscript (Page 4: line 20-22). How the ribosome small subunit affects MAGIC is beyond the focus of the current study and will be pursued in the future.
Regarding why certain mutants did not come out of our initial screen, this is not unexpected as the YKO collection, although extremely valuable to the community, is known to be potentially affected by false knockouts, suppressor accumulation and cross contamination (for references, e.g., Puddu et al., 2019 Nature). Additionally, high-through screens can also miss real hits. In our experience using this collection in several studies, we often found additional hits from analysis of genes implicated by known genetic or biochemical interactions.
Mutant yeast strains and growth assays:
The Δreg1 strain grows poorly in all growth conditions and frequently accumulates extragenic suppressor mutations (Barrett et al, 2012). It would be good to make sure that this is not the case in the strains employed in this study. My suggestion is to do (and show) standard yeast plating assays with the relevant mutant strains including Δreg1, snf1, hap4, Δreg1Δhap4 without the split GFP constructs and also with them (i.e. the strains that were used in the assays).
We thank the reviewer for the suggestion. We were indeed aware of potential accumulation of suppressor mutations from the YKO library. Therefore, deletion mutants like Δreg1 and loss of TFs downstream of Snf1 that we used in the study after the initial screen were all freshly made and validated. At least 3 independent colonies were analyzed for each mutant (mentioned in Methods & Materials; Page 33, line 57). Moreover, the plating assay suggested here may not reveal additional information other than growth, which was taken into consideration during our experiments.
Activation of Snf1 in the relevant strains should be tested with the commercially available antibody recognizing active Snf1, which is phosphorylated at Snf1-T210.
Snf1 activation was validated by the Mig1 exporting from the nucleus. We also noted above that many studies have clearly demonstrated Snf1 activation in reg1 mutant and under low glucose growth (e.g., ref. 24-28).
Effects of Snf1, Reg1, Hap4 and respiratory growth conditions:
The authors show that split GFP reporters show enhanced accumulation during fermentative growth, in Δsnf1, and Δreg1Δhap4 and fail to accumulate during respiratory growth, in Δreg1 and upon overexpression of HAP4. Analysis of Δhap4 should be included in Fig. 2. The suggestion that upon activation of Snf1 enhanced Hap4-dependent expression "outcompetes" misfolded protein import seems unlikely as only a fraction of mitochondrial genes is under control of Hap4. Without further experimental evidence I do not find that a valid assumption. More likely, the membrane potential plays a role: it is low during fermentative growth, in Δsnf1 and Δreg1Δhap4, and high during respiratory growth and in Δreg1 (Hübscher et al, 2016). Such an effect of the membrane potential seems to contradict the findings in the 2017 paper and the issue should be clarified and discussed. In any case, these data do not reveal that GFP reporters accumulate inside of the mitochondria. Based on the currently available evidence they may accumulate in close proximity/attached to the mitochondria. This has to be tested directly (see above).
We have included our analysis of Δhap4 in Page 8: line 14-15 and Figure 2—figure supplement 1H. Consistent with our result for Δreg1Δhap4 in glucose-rich medium, HAP4 deletion also resulted in a significant increase in mitochondrial accumulation of FlucSM in low glucose medium compared to WT. It did not have effect in high glucose condition in which Snf1 is largely inactive.
It is our view that the importance of Hap4 should not be judged by the number of nuclear encoded mitochondrial proteins they regulate. Still, this sub-group comprises a considerable number of proteins (at least 55 genes upregulated by Hap4 overexpression, ref. 43), and certain substrates may be more competitive with misfolded cytosolic proteins for import. Our genetic data strongly suggest that the inhibitory effect of active Snf1 on MAGIC is through Hap4, although we agree with the reviewer that detailed mechanism on how Hap4 substrates may compete with misfolded proteins need to be addressed in future studies.
Membrane potential is important for mitochondrial import. During respiratory growth and in Δreg1, membrane potential is well known to be elevated comparing to fermentative condition (e.g., Figure 4C). Our observation that the import of misfolded proteins into mitochondria is reduced under these conditions simply suggests that this reduction is not due to a lack of membrane potential. This is not in any way contradictory to our 2017 finding that misfolded protein import requires membrane potential (ref. 10).
Again, the accumulation of misfolded proteins in mitochondria, especially the model protein FlucSM, has been validated by using super resolution imaging (Figure 1—figure supplement 1A) in addition to the protease protection assay in our 2017 study.
Introduction and Discussion:
Both are really short, too short in my view. Please provide some background of the general principals of mitochondrial protein import and information of how exactly translocation of cytosolic, aggregated proteins (lacking targeting information) is supposed to work. I do not understand exactly how the authors actually envisage the process.
We thank the reviewer for the suggestion. In the revised manuscript, we have extended both Introduction (Page 2-3) and Discussion section (Page 11-13)
The results from the 2022 eLife paper (Liu et al, 2022), which suggests that Tom70 may "regulate both the transcription/biogenesis and import of mitochondrial proteins so the nascent mitochondrial proteins do not compromise cytosolic proteostasis or cause cytosolic protein aggregation" should be discussed with regard to the data obtained with overexpression of the Tom70 soluble domain.
We thank the reviewer for pointing out that study and we have included a brief comment in Discussion section (Page 12: line 13-16). As the function of Tom70 appears to be complex, we cannot exclude the possibility that overexpression of the cytosolic domain has additional or indirect effects in addition to that due to preprotein binding.
Andreasson, C., Ott, M., and Buttner, S. (2019). Mitochondria orchestrate proteostatic and metabolic stress responses. EMBO Rep 20, e47865.
Barrett, L., Orlova, M., Maziarz, M., and Kuchin, S. (2012). Protein kinase A contributes to the negative control of Snf1 protein kinase in Saccharomyces cerevisiae. Eukaryot Cell 11, 119-128.
Hubscher, V., Mudholkar, K., Chiabudini, M., Fitzke, E., Wolfle, T., Pfeifer, D., Drepper, F., Warscheid, B., and Rospert, S. (2016). The Hsp70 homolog Ssb and the 14-3-3 protein Bmh1 jointly regulate transcription of glucose repressed genes in Saccharomyces cerevisiae. Nucleic Acids Res. 44, 5629-5645.
Liu, Q., Chang, C.E., Wooldredge, A.C., Fong, B., Kennedy, B.K., and Zhou, C. (2022). Tom70-based transcriptional regulation of mitochondrial biogenesis and aging. Elife 11
Pfanner, N., Warscheid, B., and Wiedemann, N. (2019). Mitochondrial proteins: from biogenesis to functional networks. Nat Rev Mol Cell Biol 20, 267-284.
Ruan, L., Zhou, C., Jin, E., Kucharavy, A., Zhang, Y., Wen, Z., Florens, L., and Li, R. (2017). Cytosolic proteostasis through importing of misfolded proteins into mitochondria. Nature 543, 443-446.
I prefer to have "all in one", also due to time limitation.
It would be great to be able to upload the review file as otherwise formatting and symbols get lost.
Reviewer #3 (Public Review):
In this study, Wang et al extend on their previous finding of a novel quality control pathway, the MAGIC pathway. This pathway allows misfolded cytosolic proteins to become imported into mitochondria and there they are degraded by the LON protease. Using a screen, they identify Snf1 as a player that regulates MAGIC. Snf1 inhibits mitochondrial protein import via the transcription factor Hap4 via an unknown pathway. This allows cells to adapt to metabolic changes, upon high glucose levels, misfolded proteins an become imported and degraded, while during low glucose growth conditions, import of these proteins is prevented, and instead import of mitochondrial proteins is preferred.
This is a nice and well-structured manuscript reporting on important findings about a regulatory mechanism of a quality control pathway. The findings are obtained by a combination of mostly fluorescent protein-based assays. Findings from these assays support the claims well.
While this study convincingly describes the mechanisms of a mitochondria-associated import pathway using mainly model substrates, my major concern is that the physiological relevance of this pathway remains unclear: what are endogenous substrates of the pathway, to which extend are they imported and degraded, i.e. how much does MAGIC contribute to overall misfolded protein removal (none of the experiments reports quantitative "flux" information). Lastly, it remains unclear by which mechanism Snf1 impacts on MAGIC or whether it is "only" about being outcompeted by mitochondrial precursors.
We thank Reviewer 3 for the positive and encouraging comments on our manuscript. We agree with the reviewer that identifying MAGIC endogenous substrates and understanding what percentage of them are degraded in mitochondria are very important issues to be addressed. We are indeed carrying out projects to address these questions. We also agree with Reviewer 3 that the effect of Snf1 on MAGIC may have additional mechanisms in addition to precursors competition, such as Tom6 mediated conformational changes of TOM pores. In the revised manuscript, we had added a discussion to address these comments (Page 12: line 21-28).
Reviewer #3 (Recommendations For The Authors):
- In their screen, the authors utilize differences in GFP intensity as a measure for import efficiency. However, reconstitution of the GFP from GFP1-10 and GFP11 in the matrix might also be affected (folding factors, differential degradation).
Upon Snf1 activation, the protein abundance of mitochondrial chaperones such as Hsp10, Hsp60, and Mdj1, and mitochondrial proteases such as Pim1 are not significantly changed (ref. 35). Therefore, it is unlikely that the folding and degradation capacity of mitochondrial matrix is drastically affected by Snf1 activation.
To examine the effect of Snf1 activation on spGFP reconstitution, Grx5 spGFP strain was constructed in which the endogenous mitochondrial matrix protein Grx5 was C-terminally tagged with GFP11 at its genomic locus, and GFP1-10 was targeted to mitochondria through cleavable Su9 MTS (MTS-mCherryGFP1-10) (ref. 10). Only modest reduction in Grx5 spGFP intensity was observed in LG compared to HG, and no significant difference after adjusting the GFP1-10 abundance (spGFP/mCherry ratio) (Figure 1— figure supplement 3A-D). These data suggest that any effect on spGFP reconstitution is insufficient to explain the drastic reduction of MP accumulation in mitochondria under Snf1 activation. Overall, our results demonstrate that Snf1 activation primarily prevents mitochondrial accumulation of MPs, but not that of normal mitochondrial proteins. (Page 6: line 17-25).
We admit, however, that to fully rule out these factors, specific intra-mitochondrial folding or degradation reporter assays would be needed.
- Scoring of protein import always takes place using fluorescence-based assays. These always require folding of the "sensors" in the matrix. An additional convincing approach that would not rely on matrix folding could be pulse chase approaches coupled to fractionation assays and immunoprecipitation.
We thank reviewer 3 for this suggestion. In our previous study, we applied two different biochemical assays: APEX proximity labeling, and mitochondrial fractionation followed by protease protection. Both confirmed the entry of misfolded proteins into mitochondria as observed by using split GFP. As we discussed in response to Reviewer 1’s main point [3], the fractionation assays are not quantitative enough for the comparisons made in our study. In particular, during the over 2-hour assay, misfolded proteins continue to be degraded within mitochondria. By using proper controls, our spGFP system provides quantitative comparisons for mitochondrial accumulation of misfolded proteins in non-disturbed physiological conditions.
- Could the pathway be reconstituted in vitro with isolated mitochondria to test for the "competition hypothesis"
This is an excellent suggestion, but setting up such a reconstituted system is a project on its own. The study documented in this manuscript already encompasses a large amount of work that we feel should be published timely.
- Fluorescence figures are not colour blind friendly (red-green). This should be improved by changing the color scheme.
We thank reviewer 3 for pointing this out and sincerely apologize for any inconvenience. However, we are unfortunately unable to change all images within a limited time. We will adopt another color scheme in future work.
- spGFP in human cells appears to form "spot-like" structures. What are these granules?
We indeed observed granule-like structures by spGFP labeled FUS in mitochondria, which is interesting, but we did not investigate this further because it is a not a focus of this study.
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
Overall, we were pleased that the reviewers found our study carefully designed and interesting. We have addressed their comments below.
Reviewer #1 (Evidence, reproducibility and clarity (Required)):
The manuscript by Kern, et al., demonstrates that phagocytosis in macrophages is regulated in part by the intermolecular distance of phagocytosis-promoting receptors engaging phagocytic targets. Cells expressing chimeric receptors containing cytosolic domains of Fc receptors (FcR) and defined ligand-binding DNA domains were used to drive phagocytosis of opsonized glass beads coated with complementary DNA ligands of defined spacing and number. These so-called origami ligands allowed manipulation of receptor spacing following engagement, which allowed the demonstration that tight spacing of ligands (7 nm or 3.5 nm) optimized signaling for phagocytosis. The study is carefully performed and convincing. I have a few technical concerns and minor suggestions.
We did not use any shade correction. Instead we only collected data from a central ROI in our TIRF field. To check for uniformity of illumination, we plotted the origami pegboard fluorescent intensity along the x and y axis. We observed very modest drop off in signal - the average signal intensity of origamis within 100 pixels of the edge is 76 ± 6% the intensity of origamis in a 100 pixel square in the center of the ROI. Fitting this data with a Gaussian model resulted in very poor R values. While this may account for some of the variation in signal intensity at individual points, we expect the normalized averages of each condition to be unaffected. We have amended the methods to describe this strategy (Lines 851-854).
(Image could not be uploaded)
__ Likewise, how much variation was there in the expression of the chimeric receptors? Large variation in receptor numbers per cell could significantly alter the quantitative studies. Aside from the flow sorting for cells expressing two different molecules, how were cells selected for analysis?__
We thank the reviewer for bringing up this point. We confirmed comparable receptor expression levels at the cell cortex of the DNA CAR-𝛾 and the DNA CAR-adhesion used throughout the paper. We also have confirmed that receptor levels at the cell cortex were similar for the large DNA CAR constructs used in Figure 6C-D. This data is now included in Figures S5 and S7. We have also altered the text to include this (lines 169-172):
Expression of the various DNA CARs at the cell cortex was comparable, and engulfment of beads functionalized with both the 4T and the 4S origami platforms was dependent on the Fc𝛾R signaling domain (Figure S5).
When quantifying bead engulfment, cells were selected for analysis based on a threshold of GFP fluorescence, which was held constant throughout analysis for each individual experiment. We have amended the “Quantification of engulfment” methods section to convey this (lines 921-923).
__ The scale of the origami relative to the cells is difficult to discern in Figures 2C and D. Additional text would be helpful to indicate, for example, that the spots on the Fig. 2D inset indicate entire origami rather than ligand spots on individual origami particles.__
Thank you for pointing this out, we see how the legend was unclear and have corrected it (lines 453-454), including specifically noting “Each diffraction limited magenta spot represents an origami pegboard.” We have also outlined the cell boundary in yellow to make the cell size more clear.
__ Figure 5 legend, line 482: How was macrophage membrane visualized for these measurements?__
We have added the following clarification (line 535-536): “The macrophage membrane was visualized using the DNA CAR𝛾, which was present throughout the cell cortex.”
__ line 265: "our data suggest that there may be a local density-dependent trigger for receptor phosphorylation and downstream signaling". This threshold-dependent trigger response was also indicated in the study of Zhang, et al. 2010. PNAS.__
The Zhang et al. study was influential in our study design, and we wish to give the appropriate credit. Zhang et al. found that a sufficient amount of IgG is necessary to activate late (but not early) steps in the phagocytic signaling pathway. In contrast, our study addresses IgG concentration in small nanoclusters. We find that this nanoscale density affects receptor phosphorylation. Thus, we think these two studies are distinct and complementary.
Lines 283-287 now read:
While this model has largely fallen out of favor, more recent studies have found that a critical IgG threshold is needed to activate the final stages of phagocytosis (Zhang et al., 2010). Our data suggest that there may also be a nanoscale density-dependent trigger for receptor phosphorylation and downstream signaling.
__ line 55: Rephrase, “we found that a minimum threshold of 8 ligands per cluster maximized FcgR-driven engulfment.” It is difficult to picture how a minimum threshold maximizes something.__
We now state “we found that 8 or more ligands per cluster maximized FcgR-driven engulfment.”
__ line 184: Rephrase, "we created... pegboards with very high-affinity DNA ligands that are predicted not to dissociate on a time scale of >7 hr". Remove "not".__
Thank you for pointing this out, it is now correct.
Reviewer #1 (Significance (Required)):
This study provides a significant advance in understanding about the molecular mechanisms of signaling for particle ingestion by phagocytosis.
--
Reviewer #2 (Evidence, reproducibility and clarity (Required)):
The manuscript on “Tight nanoscale clustering of Fcg-receptors using DNA origami promotes phagocytosis" studies how clustering and nanoscale spacing of ligand molecules for a chimeric Fcg-receptors influence the phagocytosis of functionalized silicon beads by macrophage cell lines. The basis of this study is the design of a chimeric Fc-receptor (DNA-CARg) comprising an extracellular SNAP-tag domain that can be loaded with single-stranded (ss) DNA, the transmembrane part of CD86 and the cytosolic part of the Fc-receptor g-chain containing an immunoreceptor tyrosine-based activation motif (ITAM) as well as a C-terminal green fluorescent protein (GFP). As control the authors used a similar designed DNA-CAR that is lacking the intracellular ITAM-containing FCg tail. The chosen target for this chimeric DNA-CAR, are silicon beads covered by a lipid bilayer that contains biotin-labelled lipids that, via Neutravidin, can be loaded with a biotinylated DNA origami pegboard displaying complimentary ss-DNA as ligand for the DNA-CAR. The DNA origami pegboard contains four ATTO647N fluorescence for visualization and the ssDNA ligand in different quantities and spacing. Using these principles, the authors study how ligand affinity, concentration and spacing influence the activation of the DNA-CARg and the engulfment of the loaded beads.
The authors show that bead engulfment is increased between 2 till 8 ssDNA ligands on the pegboard. After this, ligand numbers do not play a role anymore in the engulfment. They then study the role of the ligand spacing using pegboards that either contain 4 single strand DNA ligands in close (7nm/3,5nm) proximity or a more spaced version using 21/17,5 nm or 35/38,5 nm. The authors find that the bead engulfment is maximally and positively affected by the close spacing of the ssDNA ligands. In their final experiments the authors vary the design of the DNA-CARs by tetramerization of the ITAM-containing Fcg-signaling subunit. In their discussion the authors mention different possibilities for the effect of spacing on the engulfment process.
I think that, in general, this is an interesting study. However, it has some caveats and open issues that should be clarified before its publication.
**Major comments**
We agree with the reviewer completely. We have repeated experiments shown in Figure 4A with a DNA-CAR containing the Fc𝛾 transmembrane domain instead of CD86 as the reviewer suggests. We also included a DNA-CAR version of the Fc𝛾R1 alpha chain, although this construct was not expressed as well as the others. These data are now included in Figure S5, and referenced in lines 167-168.
__ An important issue that is discussed by the authors but not addressed in this manuscript is whether the different amount and spacing of the ligand is only impacting on signaling or also on the mechanical stress of the cells. Indeed, mechanical stress on the cytoskeleton arrangement could influence the engulfment process. For this, it would be very important to test that the different bead engulfment, for example, those shown in Fig. 4, is strictly dependent on signaling kinases. The authors should repeat the experiment of Fig. 4 a and b in the presence or absence of kinase inhibitors such as the Syk inhibitor R406 or the Src inhibitor PP2 to show whether the different phase of engulfment is dependent on the signaling function of these kinases. This crucial experiment is clearly missing from their study.__
We agree this is an interesting point. We find that ligand spacing affects receptor phosphorylation; however this does not preclude effects on downstream aspects of the signaling pathway. We will clarify this by adding the following comment to the manuscript (line 299-301):
While our data pinpoints a role for ligand spacing in regulating receptor phosphorylation, it is possible that later steps in the phagocytic signaling pathway are also directly affected by ligand spacing.
The DNA-CAR-adhesion in Figure 1 strongly suggests that intracellular signaling is essential for phagocytosis. We have now included additional controls using this construct as detailed in our response to point 3 below. Unfortunately, Src and Syk inhibitors or knockout abrogate Fc𝛾R mediated phagocytosis (for example, PMIDs 11698501, 9632805, 12176909, 15136586) and thus would eliminate phagocytosis in both the 4T and 4S conditions. This precludes analysis of downstream steps in the phagocytic signaling pathway.
__ Another problem of this study is that the authors show in Fig. 1A the control DNA-CAR-adhesion but then hardly use it in their study. For example, the crucial experiments shown in Fig. 4 should be conducted in parallel with DNA-CAR-adhesion expressing macrophage cells. This study could provide another indication whether or not ITAM signaling is important for the engulfment process.__
We have added this control. It is now included in Figure S5 and S7. Figure 3D also shows that the DNA-CAR-adhesion combined with the 4T origami pegboards does not activate phagocytosis and we have amended the text to make this more clear (line 152).
__ Another important aspect is how the concentration of the loaded origami pegboard is influencing the engulfment process. In particular, it would be interesting to show the padlocks with different spacings such as the 4T closed spacing versus 4s large spacing show a different dependency on the concentration of this padlock loading on the beads. This would be another important experiment to add to their study.__
We agree that this is an interesting question. We suspect that at a very high origami density, 4S signaling would improve, and potentially approach the 4T. However, we are currently coating the beads in saturating levels of origami pegboards. Thus we cannot increase origami pegboard density and address this directly.
**Minor comments:**
__ The authors discuss that it is the segregation of the inhibitory phosphatase CD45 from the clustered Fc receptors is the major mechanism explaining their finding that 4T closed spacing is more effective than 4s large spacing. With the event of the CRISPR/Cas9 technology it is trivial to delete the CD45 gene in the genome of the RAW264.7 macrophage cell line used in this study and I am puzzled why they author are not conducting such a simple but for their study very important experiment (it takes only 1-2 month to get the results).__
This experiment may be informative but we have two concerns about its feasibility. First, CD45 is a phosphatase with many different roles in macrophage biology, including activating Src family kinases by dephosphorylating inhibitory phosphorylation sites (PMID 8175795, 18249142, 12414720). Second, CD45 is not the only bulky phosphatase segregated from receptor nanoclusters. For example, CD148 is also excluded from the phagocytic synapse (PMID 21525931). CD45 and CD148 double knockout macrophages show hyperphosphorylation of the inhibitory tyrosine on Src family kinases, severe inhibition of phagocytosis, and an overall decrease in tyrosine phosphorylation (PMID 18249142). CD45 knockout alone showed mild phenotypes in macrophages. We anticipate that knocking out CD45 alone would have little effect, and knocking out both of these phosphatases would preclude analysis of phagocytosis. Because of our feasibility concerns and the lengthy timeline for this experiment, we believe this is outside of the scope of our study.
In our discussion, we simplistically described our possible models in terms of CD45 exclusion, as the mechanisms of CD45 exclusion have been well characterized. This was an error and we have amended our discussion to read (lines 335-343):
As an alternative model, a denser cluster of ligated receptors may enhance the steric exclusion of the bulky transmembrane proteins like the phosphatases CD45 and CD148 (Bakalar et al., 2018; Goodridge et al., 2012; Zhu, Brdicka, Katsumoto, Lin, & Weiss, 2008).
Reviewer #2 (Significance (Required)):
The innovative part of this study is the combination of SNAP-tag attached, chimeric Fc-receptor with the DNA origami pegboard technology to address important open question on receptor function.
**Referees cross-commenting**
I find most of my three reviewing colleagues reasonable
I also agrée to Reviewer #1 comments 2
Likewise, how much variation was there in the expression of the chimeric receptors? Large variation in receptor numbers per cell could significantly alter the quantitative studies. Aside from the flow sorting for cells expressing two different molecules, how were cells selected for analysis?
But I want to add it is not only the amount of receptors but ils the nanoscale location that is key to receptor function
We have ensured that all receptors are trafficked to the cell surface. We have also measured their intensity at the cell cortex as discussed in response to Reviewer 1.
Reviewer #3 (Evidence, reproducibility and clarity (Required)):
This is a very nicely done synthetic biology/biophysics study on the effect of ligands spacing on phagocytosis. They use a DNA based recognition system that the group has previously use to investigate T cell signaling, but express the SNAP tag linked transmembrane receptor in a macrophage cell line and present the ligands using DNA origami mats to control the number and spacing of complementary ligands that are designed to be in the typical range for low or high affinity FcR, a receptor that can trigger phagocytosis. The study offers some very nice quantitative data sets that will be of immediate interest to groups working in this area and, in the future, for design of synthetic receptors for immunotherapy applications. Other groups are working on similar platform for TCR. I don't feel there is any need for more experiments, but I have some questions and suggestions. Answering and considering these could clarify the new biological knowledge gained.
We thank the reviewer for their support of our manuscript. Given the reviewer’s statement that no new experiments are required, we have answered their questions to the best of our ability given the current data. Should the editor decide that any of these topics require experimental data to enhance the significance of the paper, we are happy to discuss new experiments.
Reviewer #3 (Significance (Required)):
I think the significance would be increased by addressing these questions, that would help understand how the synthesis system described related to other system directed as similar questions and more natural settings.
We saw robust phagocytosis at 300 molecules/micron of ssDNA, which is similar to the IgG density used on supported lipid bilayer-coated beads in other phagocytosis studies (PMID 29958103, 32768386). As the reviewer noticed, this is significantly higher than ligand density necessary to activate T cells (PMID 28340336). We have added a comment on ligand density to lines 96-97.
__ Are the origami mats generally laterally mobile on the bilayers. If so, what is the diffusion coefficient? Can one detect the mats accumulating in the initial interface between the bead and cell, particularly in cased where there is no phagocytosis? Would immobility of the mats make them more efficient at mediating phagocytosis compared to the monodispersed ligands, which I assume are highly mobile and might even be "slippery".__
We have confirmed that our bead protocol generally produces mobile bilayers, where his-tagged proteins can freely diffuse to the cell-bead interface (see accumulation of a his-tagged FRB binding to a transmembrane FKBP receptor at the cell-bead synapse below). We can qualitatively say that the origamis appear mobile on a planar lipid bilayer (see Dong et. al 2021 and images below). Directly measuring the diffusion coefficient on the beads is extremely difficult because the beads themselves are mobile (both diffusing and rotating), and cannot be imaged via TIRF. We do not see much accumulation of the origami at cell-bead synapses. This could reflect lower mobility of the origamis, or could be because the relative enrichment of origamis is difficult to detect over the signal from unligated origamis.
Overall, we expect the origami pegboards (tethered by 12 neutravidins) are less mobile than single strand DNA (tethered by a single neutravidin, supported by qualitative images below). We are uncertain whether this promotes phagocytosis. At least one study suggests that increased IgG mobility promotes phagocytosis (PMID 25771017). However, the zipper model would suggest that tethered ligands may provide a better foothold for the macrophage as it zippers the phagosome closed (PMID 14732161). Hypothetically, ligand mobility could affect signaling in two ways - first by promoting nanocluster formation, and second by serving as a stable platform for signaling as the phagosome closes. Since our system has pre-formed nanoclusters, the effect of ligand mobility may be quite different than in the endogenous setting.
(Image could not be uploaded)
In the above images, a 10xHis-FRB labeled with AlexaFluor647 was conjugated to Ni-chelating lipids in the bead supported lipid bilayer. The macrophages express a synthetic receptor containing an extracellular FKBP and an intracellular GFP. Upon addition of rapamycin, FRB and FKBP form a high affinity dimer, and FRB accumulates at the bead-macrophage contact sites.
(Image could not be uploaded)
In the above images, single molecules were imaged for 3 sec. The tracks of each molecule are depicted by lines, colored to distinguish between individual molecules. The scale bar represents 5 microns in both panels.
__ Breaking down the analysis into initiation and completion is interesting. When using the non-signalling adhesion constructs, would they get to the initiation stage or would that attachment be less extensive than the initiation phase.__
This is an interesting question. While we did not include the DNA-CAR-adhesion in our kinetic experiments, we have now quantified the frequency of cups that would match our ‘initiation’ criteria in 3 representative data sets where macrophages were fixed after 45 minutes of interaction with origami pegboard-coated beads. We found that an average of 16/125 of 4T beads touching DNA-CAR-adhesion macrophages met the ‘initiation’ criteria and an average of 2/125 were eaten (14% total). In comparison, we examined 4T beads touching DNA CAR𝛾 macrophages and found that on average 23/125 met the ‘initiation’ criteria, and 45/125 were already engulfed (54%). This suggests that the DNA-CAR-adhesion alone may induce enough interaction to meet our initiation criteria, but without active signaling from the FcR this extensive interaction is rare. We have added this data in a new Figure S6 and commented on this in lines 213-215.
__ It would be interesting to put these results in perspective of earier work on spacing with planar nanoarrays, although these can't be applied to beads. For integrin mediated adhesion there was a very distinct threshold for RGD ligand spacing that could be related to the size of some integrin-cytoskeletal linkers (PMID: 15067875). On the other hand, T cell activation seemed more continuous with changes in spacing over a wide range with no discrete threshold (PMID: 24117051, 24125583) unless the spacing was increased to allow access to CD45, in which case a more discrete threshold was generated (PMID: 29713075). The results here for phagocytosis with the very small ligands that would likely exclude CD45 seems to be more of a continuum without a discrete threshold, although high densities of ligand are needed. This issue of continuous sensing vs sharp threshold is biologically interesting so would be good assess this by as consistent standards are possible across systems.__
We agree that this is an interesting body of literature worth adding to our discussion. We have added a paragraph that puts our study in the context of prior work on related systems, including these nanolithography studies (Line 364-382):
How does the spacing requirements for Fc𝛾R nanoclusters compare to other signaling systems? Engineered multivalent Fc oligomers revealed that IgE ligand geometry alters Fcε receptor signaling in mast cells (Sil, Lee, Luo, Holowka, & Baird, 2007). DNA origami nanoparticles and planar nanolithography arrays have previously examined optimal inter-ligand distance for the T cell receptor, B cell receptor, NK cell receptor CD16, death receptor Fas, and integrins (Arnold et al., 2004; Berger et al., 2020; Cai et al., 2018; Deeg et al., 2013; Delcassian et al., 2013; Dong et al., 2021; Veneziano et al., 2020). Some systems, like integrin-mediated cell adhesion, appear to have very discrete threshold requirements for ligand spacing while others, like T cell activation, appear to continuously improve with reduced intermolecular spacing (Arnold et al., 2004; Cai et al., 2018). Our system may be more similar to the continuous improvement observed in T cell activation, as our most spaced ligands (36.5 nm) are capable of activating some phagocytosis, albeit not as potently as the 4T. Interestingly, as the intermembrane distance between T cell and target increases, the requirement for tight ligand spacing becomes more stringent (Cai et al., 2018). This suggests that IgG bound to tall antigens may be more dependent on tight nanocluster spacing than short antigens. Planar arrays have also been used to vary inter-cluster spacing, in addition to inter-ligand spacing (Cai et al., 2018; Freeman et al., 2016). Examining the optimal inter-cluster spacing during phagosome closure may be an interesting direction for future studies.
--
Additional experiments performed in revision
In addition to these reviewer comments, we have added additional controls validating the DNA-CAR-4x𝛾 used in Figure 6c,d. We compared the DNA-CAR-4x𝛾 to versions of the DNA-CAR-1x𝛾-3x𝛥ITAM construct with the functional ITAM in the second and fourth positions (see the schematics now included Figure S7). We found that four individual receptors with a single ITAM each were able to induce phagocytosis regardless of which position the ITAM was in. However the DNA-CAR-4x𝛾 construct, which also contains 4 ITAMs, was not. This further validates the experiment presented in 6c,d. We also fixed minor errors we discovered in the presentation of data for Figures 1C and S1A.
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
Manuscript number: RC-2023-02157
Corresponding author(s): Satish, Mishra
We thank the editor and reviewers for their helpful comments. We have successfully addressed most of the comments. We are performing some additional experiments as suggested by the reviewers and will be included if considered further. We attempted to pulldown the S14 interacting partner using anti-mCherry antibody from S14-3XHA-mCherry transgenic sporozoites and then further tried to identify interactome using mass spectrometry but failed. So, accordingly, we have toned down the conclusion.
The point-by-point response to the reviewer’s comments is given as follows.
Reviewer #1:
Figure 1F You have not formally shown that this signal corresponds to palmitoylated S14. Could be heavy chain. Response: The possibility of a heavy chain is negligible because we have used sporozoite samples and probed it with an anti-rabbit antibody conjugated to HRP. Also, the size of the S14 bands does not correspond to heavy chain. However, we have toned down the conclusion. Currently, we are performing the depalmitoylation experiment, and data will be included in the next round of revision.
Reviewer #2
Line 149: To definitively state S14 is a membrane protein, biochemical assays proving such should be performed. (or perhaps genetic mutation of the predicted palmitoylation site?) Otherwise, this should be rephrased. Response: We are performing the depalmitoylation assay, and the data will be included during the second round of revision. However, we have rephrased the sentence in the current version of the manuscript.
Lines 257-258: for yeast 2-hybrid, the controls of expressing S14, GAP45 and MTIP together with control proteins where no interaction would be predicted are absent. Response: We are performing experiments with additional controls, and data will be included in the next round of revision.
Reviewer #3
Conclusions that S14 knockout does not impact the expression and organization of two surface proteins, CSP and TRAP, and two IMC rely on a qualitative analysis only. However, quantitative analysis to support their observations is missing. Response: We are quantifying the IFA images and data will be included in the next round of revision.
Reviewer #1 (Evidence, reproducibility and clarity (Required)):
Summary: The authors have identified a sporozoite gliding motility protein through bioinformatic analysis. From the main text I do not know how, or what bioinformatic analysis was performed, in order to focus on this protein which is called S14. The authors then go on to tag the protein, produce a KO and show its involvement in gliding motility. The KO shows that parasites lacking S14 fail to invade the mosquito salivary glands. This is due to a motility defect. Y2H and docking studies are used to define an interaction with MTIP and GAP45, two known components of the glideosome. Response: We identified this gene from the Kaiser et al., 2004 paper (DOI: 10.1046/j.1365-2958.2003.03909.x). The S14 was found to be highly upregulated in salivary gland sporozoites but lacked signal sequence and transmembrane domain. Next, we looked into other sporozoite proteins lacking signal sequence and transmembrane domain and found several gliding-associated proteins with similar properties. By using the guilt-by-association principle (DOI: 10.1186/gb-2009-10-4-104), we studied the following properties of existing glideosome components along with S14: (1) Classical pathway secretion using the signal peptide (SignalP, https://services.healthtech.dtu.dk/services/SignalP-5.0) (http://dx.doi.org/10.1016/j.jmb.2004.05.028). (2) Nonclassical pathway secretion (SecretomeP , https://services.healthtech.dtu.dk/services/SecretomeP-1.0/) (10.1093/protein/gzh037). (3) Presence of transmembrane domains (TMHMM , https://services.healthtech.dtu.dk/services/TMHMM-2.0/) (10.1006/jmbi.2000.4315). (4) Presence of a potential palmitoylation site (CSS-Palm, http://bioinformatics.lcd-ustc.org/css_palm) (Ren et al, 2008). This is a similar association prediction method as employed by the STRING database. However, mentioning that we identified a gliding motility protein by bioinformatic analysis was wrong, and we modified the sentence.
Major comments: The paper is sometimes hard to follow and lacks clarity. The reason: important information is omitted, or explained at the end of a section rather than at first mention; experimental details that are of essence need to be mentioned or explained in the main text; there is ample use of the word 'bioinformatic' without explaining what kind of analysis was performed in the main text. I cite from the abstract: 'In silico analysis of a novel protein, S14, which is uniquely upregulated in salivary gland sporozoites, suggested its association with glideosome-associated proteins.' I cite from the introduction: 'A study comparing transcriptome differences between sporozoites and merozoites using suppressive subtraction hybridization found several genes highly upregulated in sporozoites and named them 'S' genes (Kaiser et al, 2004). We narrowed it down to a candidate named S14, which lacked signal peptide and transmembrane domains.' From reading the main text, I do not know why Plasmodium berghei S14 was chosen in this manuscript. S14 is one of 25 transcripts identified by Kappe et al in Plasmodium yoelii (https://doi.org/10.1046/j.1365-2958.2003.03909.x) to be upregulated in sporozoites. The material and methods section does not explain either why S14 was chosen. Perhaps the authors could update Figure 2 from Kappe et al with the most recent annotations from plasmodb. Response: We have edited the manuscript for clarity and mentioned the name of the bioinformatic analysis performed. We chose S14 from Kaiser et al., 2004 (https://doi.org/10.1046/j.1365-2958.2003.03909.x) that identified transcripts in P. yoelii. We work on the rodent malaria parasite P .berghei and validated S14 transcripts by qPCR which showed its upregulation in sporozoites.
Rodent malaria parasites P. berghei and P. yoelii have been used extensively as models of human malaria. Both species have been widely used in studies on the biology of Plasmodium sporozoites and liver stages due to the availability of efficient reverse genetics technologies, and the ability to analyze these parasites throughout the life cycle stages have made these two species the preferred models for the analysis of Plasmodium gene function. A genetic screen and phenotype analysis were performed in P. berghei (DOI: 10.1016/j.cell.2017.06.030 and DOI: 10.1016/j.cell.2019.10.030) that makes in-depth characterization easier due to the availability of reagents and preliminary gene-phenotype like its dispensability in the blood. As suggested by this reviewer, we have updated the most recent annotations from PlasmoDB.
Reproducibility: None of the main Figures or Figure legends define ' N = '. For example I cite: 'The S14 KO clonal lines were first analyzed for asexual blood-stage propagation, and for this, 200 µl of iRBCs with 0.2% parasitemia was intravenously injected into a group of mice.' There are 2 mentions of 'N=' in the supplementary figures. I have not found any others.
I'm not sure what the convention is. Should unpublished data for this gene (PBANKA_0605900) found in pberghei.eu (a database for mutant berghei parasites) be cited? After all it confirms their findings.
The authors need to use more recent references for some of their statements; see some comments below. __Response: __We have mentioned N in the figures legends of the revised manuscript and also mentioned the unpublished data of RMGM. We have also added recent references in the revised manuscript.
Minor comments:
line 1-2 Add the Plasmodium species of this study.
Response: Added.
abstract Which species do you work with?
Response: We have mentioned P. berghei in the abstract of the revised manuscript.
29 mosquito salivary glands and human host hepatocytes
Response: Corrected.
30 to the glideosome, a protein complex containing [...]
Response: Corrected.
32-33 What kind of in silico analysis suggested S14 is part of the glideosome? S14 is not uniquely upregulated; there are other S-type genes identified by Kappe and Matuschewski. 25 I believe.
Response: Mentioning that in silico analysis suggested S14 is part of the glideosome was a wrong statement, and we have modified the sentence for clarity in the revised manuscript.
32 Please point out he species were S genes were identified. SGS of which species?
Response: The S genes were identified in the transcriptomic study of Plasmodium yoelii.
34 expression: change to transcription
Response: Changed.
39 What kind of in silico analysis was used here? and therefore malaria transmission
__Response: __In silico, protein-protein docking interaction analysis was used.
55 A single zygote transforms into a single ookinete, which establishes a single oocyst, which in turn can produce thousands of midgut sporozoites. Please correct the life cycle passage.
Response: Corrected. located or anchored in the IMC? And located between the IMC and plasma membrane?
Response: Glideosome is located between the plasma membrane and IMC, and the same has been corrected in the revised manuscript.
61-63 Refer to Table S1 and its contents here 64 Name the known GAPs. Response: Done.
65-67 Which transmembrane domain proteins? Please add more recent references than King 1988.
Response: We have added TRAP as a transmembrane domain protein and updated the reference.
71-72 TRAP was the first protein found to be ...
Response: Corrected.
74-76 Add additional, more recent references: for example search Frischknecht and TRAP
Response: Added.
76 S6 (TREP) is also [...]
Response: Done.
88 Some of these proteins are also expressed in ookinetes.
Response: Corrected.
89-91 The sentence needs a verb.
Response: Added.
88-96 Please add some more recent glideosome papers. After 2013.
Response: Added.
91 Why do you call it a peripheral protein?
Response: Because the GAP45 was detected at the periphery of the merozoite in P. falciparum. As there are no such reports in sporozoites hence we have removed peripheral in the revised manuscript.
91-93 There are more recent citations for GAP45 and GAP50. Response: We have added recent citations.
96 Insert a reference here.
Response: Added.
99 Please define the gliding-associated proteins. What are they? Aren't there papers on GAP40, 45 and 50? DOI: 10.1016/j.chom.2010.09.002
Response: Done.
99 .... What prompted you to identify a novel GAP? And why is S14 classified as a GAP?
Response: This was a wrong statement, which we removed in the revised manuscript.
99-102 What kind of bioinformatic study? Why was S14 chosen? Please outline how you ended up with S14. Any other proteins that came out of the bioinformatic screen from the list of S genes?
Response: We identified S14 from the Kaiser et al., 2004 paper and analyzed its properties using the “guilt-by-association” principle. The analysis showed that S14 had properties similar to GAP45 and MTIP. The S14 upregulation in sporozoites and its properties similar to known GAPs, we were prompted to study this gene's function.
How many proteins were identified in the screen for sporozoite upregulated proteins by Kappe and Matuschewski?
Response: 25 genes were identified in that paper, including the two characterized genes CSP and TRAP during that study.
102-103 Define the nonclassical secretion pathway. Please reference GAP45 and GAP50 data for the nonclassical pathway.
Response: When proteins are secreted out of the cytosol without predictable or known signal sequences or secretory motifs are classified as non-classically secreted proteins, and the pathway is called a non-classical protein secretory pathway. References: https://doi.org/10.1371/journal.pone.0125191; https://doi.org/10.1016/S0171-9335(99)80097-1; doi: 10.3389/fmicb.2016.00194
105 Please add P. berghei to the title, the abstract, the introduction.
Response: Added.
111 The results section does not outline what bioinformatic analysis was used
Response: The guilt-by-association principle was used, and it is outlined in the revised manuscript.
112-114 Please specify the exact number of upregulated in sporozoites genes. I think it was 25. And add the species the study was performed in. Why did you choose the Kappe study but not the uis genes from berghei?
Response: It was 25, and the species was P. yoellli. The domains of all 25 proteins are shown in Figure 2 of Kappe study. It intrigued us after having a glance at it. Later, we confirmed the upregulation of S14 transcripts in P. berghei sporozoites and chose to study the function of this gene.
114-115 How did you narrow it down to S14? The Kappe paper lists 25 S-type genes from P. yoelii.
Response: The domains of all 25 proteins are shown in the Kappe study. Two proteins, S14 and S15, lack signal sequence and transmembrane domain, which intrigued us after glancing at them. We chose S14 because its microarray induction is higher compared to S15.
118 Plasmodia is not the plural for a group of different Plasmodium species. Use: [...] conserved among Plasmodium spp.
Response: Corrected.
118-119 Which proteins did you analyze? And how did you analyze them? Where is the data for this analysis? Outline the amino acids that predict palmitoylation? The nonclassical pathway?
Response: The proteins we analyzed are given in Table S1. We analyzed them by the guilt-by-association principle. The data for this analysis is shown in Table S1. The amino acids predicted to be palmitoylated are C59 and C228 (S14), C5 (GAP45), C8 and C5 (MTIP). Non-classical pathway secretion was predicted by SecretomeP ( 10.1093/protein/gzh037).
119-122 Here: do you mean S14 has similar properties as GAP 45 and GAP50? Define the nonclassical pathway? How do you know S14 is in the IMC?
Response: The similar properties of S14 and GAP45 are Signal Peptide Prediction, Prediction of Non-classical pathway secretion, number of predicted transmembrane domains and prediction of Palmitoylation signal. GAP50 was wrongly mentioned here and has been removed from the revised manuscript.
When proteins are secreted out of the cytosol without predictable or known signal sequences or secretory motifs are classified as non-classically secreted proteins. The pathway is called a non-classical protein secretory pathway.
Our colocalization data of S14 with GAP45 and MTIP indicated that S14 is in the IMC.
122-123 Please reference the bioinformatic analysis plus URL that allows targeting to the IMC to be analyzed.
Response: All the URLs with references are given in the method section, lines 348-358 in the revised manuscript.
123-124 Please reference the URLs for TM, palmitoylation, and interactions analyses.
Response: All URLs with references are given in the method section, lines 348-358 in the revised manuscript.
125-127 How did you predict that S14 is secreted via the nonclassical pathway?
Response: We predicted non-classical pathway secretion of S14 using - SecretomeP (https://services.healthtech.dtu.dk/services/SecretomeP-1.0/) (10.1093/protein/gzh037).
128-130 Define the nonclassical pathway when it first appears in your manuscript.
The citation Moskes 2004 is not in the reference list
Response: The nonclassical pathway is defined in lines 105-107. The citation Moskes 2004 has been included in the revised manuscript.
132 Which membrane?
Response: Live S14-mCherry localization on the membrane does not differentiate between the outer membrane or IMC. Hence, only membrane is mentioned. Next, in Figure 4A, we confirmed S14 localization on IMC by treating sporozoites with Triton X-100 and colocalizing with IMC proteins GAP45 and MTIP.
134-135 In which species?
Response: We have mentioned P. berghei in the text in the revised manuscript.
141-142 Please include images of blood stage and liver stage parasites.
Response: Blood and liver stage images are included in the revised manuscript as Figure S2.
142-143 Which membrane?
Response: Live S14-mCherry localization on the membrane does not differentiate between the outer membrane or IMC. Hence, only membrane is mentioned. Next, in Figure 4A, we confirmed S14 localization on IMC by treating sporozoites with Triton X-100 and colocalizing with IMC proteins GAP45 and MTIP.
148-149 I cannot find the specific figure you refer to; I checked the online version of the Frenal 2010 paper.
Response: Electromobility shifts of GAP45 due to the palmitoylation have been reported in (Rees-Channer et al, 2006; DOI: 10.1016/j.molbiopara.2006.04.008). Frenal 2010 paper has stated about two bands but experimentally, it was shown in Rees-Channer et al, 2006 in Figures 1 and 2B.
175 gland, we counted [...]
Response: Corrected.
177 Compared to the
Response: Corrected.
177-179 Failed to invade (absolutely)? Or invaded in highly reduced numbers?
Response: Corrected.
182-186 Please be precise: I think you mean you let all types of mosquitoes take a blood meal; s14 knockout-infected mosquitoes did not infect mice.
Response: Corrected.
181-202 Perhaps use paragraphs to indicate the different types of experiments performed here.
Response: Done.
204 Please introduce paragraphs to identify the different experiments in this section
Response: Done.
208 Outer or inner membrane of what? IMC, the plasma membrane?
Response: We treated sporozoites with Triton X-100 to analyze whether S14 is present on the outer membrane (plasma membrane) or IMC.
228 onwards Structural models were obtained from whom? Which species did you use for the docking study? Could you use in one approach 3 berghei proteins, and confirm your docking studies with the falciparum proteins? That would strengthen your model. Should you include a negative control protein in the approach? Response: The structural models were obtained using the trROSETTA server. We used P. berghei for the docking study. In the old annotation and RMGM, the ortholog of P. berghei (PBANKA_0605900) in P.falciparum (PF3D7_1207400) was indicated. However, the updated PlasmodDB does not show PBANKA_0605900 ortholog in P. falciparum. We did try to generate structure models of P. falciparum MTIP, GAP45 and S14 using the trROSETTA server. We successfully reproduced the structure of MTIP, and GAP45 but the quality of S14 structure was unsuitable for the interaction studies. The negative control cannot be included in this kind of study because it gives a false interface, and none of the previous studies have used negative control.
250-251 Was all of the gene cloned? Please define amino acid range. discussion
Response: Full-length gene of S14, MTIP and GAP45 was cloned and the same has been mentioned in materials and methods in the revised manuscript.
Please discuss data from https://elifesciences.org/articles/77447 in relation to your protein Response: Discussed.
298-300 More recent glideosome papers exist. For example https://doi.org/10.1038/s42003-020-01283-8
Response: Included.
340 List the proteins you analysed. Add URL (websites) to the analyses tools.
Response: They are listed in Table S1. The method section gives all the URLs with references, lines 348-358 in the revised manuscript.
343 Known association from the literature: how was this done?
Response: The interactions demonstrated by different groups have been summarized in the review by Boucher & Bosch, 2015 (doi: 10.1016/j.jsb.2015.02.008).
346-349 A few glideosome components? On what basis were they selected and which are they? Response: The analysis showed that S14 had properties similar to GAP45 and MTIP. Additionally, S14 localized with GAP45 and MTIP, hence selected for interaction studies.
471 Can AlphaFold Structure Predictions be used in the docking studies?
Response: Even the Alphafold AI is trained on existing sequence/structure information despite being advertised as a de novo prediction system. That's why it can't produce good quality structures of evolutionarily unique proteins such as S14. We initially started our protein model generation by alphafold2, but the quality of the structure was very low; then we further used the trRosetta server (https://yanglab.nankai.edu.cn/trRosetta/), which shows the quality of all three protein structures above 95 after validation by using UCLA-DOE LAB-SAVES V6.0 (https://saves.mbi.ucla.edu/).
tr-Rosetta includes inter-residue distance, orientation distribution by a deep-neural network, and homologous template to improve the accuracy of models (DOI: 10.1038/s41596-021-00628-9).
We have given the model structure generated using alphafold2 for your reference.
Model generated by using AlphaFold2.ipynb (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2.ipynb#scrollTo=kOblAo-xetgx).
Structure quality assessment by __http://saves.mbi.ucla.edu/.__
GAP 45
__S14 __
MTIP
487 What parts of theses genes was cloned? Define the amino acid range.
Response: The full-length protein-encoding gene was cloned.
714 Please split the table into A Mosquito bite and B haemolymph Sporozoites Response: Done.
Figure 1 For clarity, maybe write S14::mCherry
Response: Done.
Figure 1 It would be useful to show blood stage parasite images.
Response: Blood stage parasite image is included in the revised manuscript as Figure S2.
Figure 2G Haemolymph sporozoites ?
Response: Done.
Figure 8 You argued that S14 is a membrane-bound protein through palmitoylation. Here the protein is shown to be cytoplasmic. Please update our model with more recent ones. Response: We have shown that S14 colocalizes with GAP45 and MTIP, suggesting its IMC localization. We have updated our model in Figure 8.
Figure S2B It would be good to include a positive control for these PCRs.
Response: We have replaced the figure's new gel with a positive control.
Figure S3 It would be good to include a positive control for these PCRs. Response: We already have positive controls in Figure S3C and S3F for all the primer pairs used.
Tabel S1 Table S1 is only mentioned twice in the text: lines 124 and 128. There is no mention that the table contains all (??) known gliding motility proteins.
Response: The table does not contain all the gliding proteins; however, most of the proteins mentioned in the Boucher & Bosch, 2015 paper (doi: 10.1016/j.jsb.2015.02.008) were included.
Table S1 The algorithms / websites used for bioinformatic prediction need to be listed here.
Response: Included.
Table S2 Add the plasmodb gene identifiers here. The table does not show all Plasmodium spp. but a selection. Response: All the orthologs mentioned in Figure S1 and Table S2 are not shown in the updated PlasmoDB. Accordingly, we have removed the Figure S1 and Table S2 in the revised manuscript__.__
Reviewer #1 (Significance (Required)):
General assessment: The authors provide an in-depth analyses of the Plasmodium berghei protein S14 and its involvement in gliding motility. Response: Thank you.
Advance: This paper is the first analysis of the S14 protein. The authors suggest a bridging function for the protein between MTIP and GAP45. Response: Thank you.
Audience: Gliding motility is of interest to the apicomplexan field. I think this particular proteins is specific to Plasmodium spp. Response: Thank you.
Reviewer #2
Summary:
The authors tag the sporozoite protein S14 in P. berghei and show localization near the sporozoite plasma membrane. They also convincingly show, through the generation of S14 knockout lines, that S14 is required for sporozoite motility and thereby also salivary gland and hepatocyte invasion. Their bioinformatic results support possible interactions between S14 and the inner membrane complex proteins MTIP and GAP45. These analyses were performed with these specific candidate proteins rather than being unbiased searches for potential interaction partners. The yeast 2-hybrid data to support these possible protein interactions need further controls.
Lines 143-144: Unless the sporozoites were not permeablized prior to staining, it is not clear if the protein is "on" the plasma membrane or just under the plasma membrane. Furthermore, this statement anyway seems contradictory to the authors' interpretation of Figure 4A. Response: Live S14-mCherry localization on the membrane does not differentiate between the outer membrane or IMC. Next, in Figure 4A, we confirmed S14 localization on IMC by treating sporozoites with Triton X-100 and colocalizing with IMC proteins GAP45 and MTIP. Further, we ensured that mCherrey signals were bleached post-fixation and performed IFA with and without permeabilization. We revealed the mCherry and CSP signals using Alexa 488 and Alexa 594, respectively. We observed the mCherrey signal with permeablized sporozoites, not without permeabilization.
Line 218: "This result indicates that S14 is present within the inner membrane of sporozoites." While this data shows that S14 is not in the plasma membrane of the parasite, how can the authors be sure it is at the IMC? Response: S14 colocalization with MTIP and GAP45 suggested its localization on IMC.
Line 225-226: This sentence overreaches in its conclusion. There is no indication that this protein provides the power or force behind the sporozoites forward movement. Several proteins are known to be required for gliding motility, but they are not all force-providing factors. Response: We have modified the sentence, and now it states, ‘These data demonstrate that S14 is an IMC protein, essential for the sporozoite's gliding motility.
Minor comments:
Line 99: "the role of gliding-associated proteins is unexplored" There are several publications on GAP40, GAP45 and GAP50 (some of which are referenced in the previous paragraph). Response: We have included the reference for studied proteins and modified the sentence for clarity.
Line 114: "We narrowed it down to a candidate" Narrowed down how? Or rephrase. Response: We identified the S14 gene from the Kaiser et al., 2004 paper (DOI: 10.1046/j.1365-2958.2003.03909.x) and rephrased the sentence in the revised manuscript.
Lines 120-123 are strangely written, and I don't follow the logic. What "similar properties" do GAP45 and GAP50 have with S14 and are they really indicative of function? Also if palmitoylation and myristylation and nonclassical secretion are present in most eukaryotes, why would they necessarily be evidence of IMC targeting? Response: It was wrongly written, we have modified the sentence for clarity.
Line 148-149. I did not see examples of this electromobility shift of GAP45 in this publication (although I may have overlooked it).
Response: Electromobility shifts of GAP45 due to the palmitoylation have been reported in (Rees-Channer et al, 2006; DOI: 10.1016/j.molbiopara.2006.04.008). Frenal 2010 paper has stated about two bands, but experimentally it was shown in Rees-Channer et al, 2006 in Figure 1 and 2B.
Table 1 legend should preferably specify that hemolymph sporozoites were used for IV infections. Response: Done.
Line 228: Should be rephrased for accuracy. "revealed the" should be replaced with "suggests" Response: Replaced.
Lines 305-307: I don't entirely understand the logic laid out here.
Response: This was written about GAP45 and MTIP coordination; however, it has been removed in the revised manuscript.
Lines 320-322: "We hypothesize that S14 possibly plays a structural role and maintains the stability of IMC required for the activity of motors during gliding and invasion." The data about the IMC structure shown is fluorescence microscopy - and there no change is observed in the IMC in the knockout line. I suggest removing or rephrasing this point if no extra data is provided to show this. Response: We have removed this sentence in the revised manuscript.
Reviewer #2 (Significance (Required)):
The work gives insights into an unstudied, conserved Plasmodium protein, S14, which the authors show is critical for Plasmodium transmission from mosquitoes. The parasite genetics and phenotyping demonstrating this are strong. The conclusions about interactions with glideosome/inner membrane complex components need further experimental support. The work is of interest to the Plasmodium field and may be also of interest to people interested in other protozoan parasites or in cellular motility.
Reviewer #3 (Evidence, reproducibility and clarity (Required)):
The manuscript by Gosh and colleagues demonstrates that S14 is a glideosome-associated protein in sporozoites. S14 knockout sporozoites fail to infect mosquito salivary glands and liver cells in the mammalian host. These sporozoites are also defective in gliding motility as S14 localizes to the inner membrane. S14 was shown to interact with the glideosome-associated proteins GAP45 and MTIP using the yeast two-hybrid system. The authors also provide an in-silico prediction on the S14, GAP45 and MTIP interaction.
Major issues:
Overall, there is information lacking in the manuscript, including on the figure legends, regarding experiments replication and n analyzed.
For complementation, the authors engineered an independent S14 knockout line. For this line is clear that parasites failed to infect salivary glands contrarily to the knockout line. Despite not showing it, did the authors confirm that this knockout line has no defects in infecting mosquito midguts and producing sporozoites? Response: We analyzed the midgut for sporozoite formation, which was comparable to the original KO line, and included the data (Figure 2D) in the revised manuscript.
Did the authors conduct IV injections in mice with a higher number of sporozoites? Hemolymph sporozoites are less infectious than sporozoites collected from the salivary glands and I was wondering whether patent infections with S14 ko sporozoites can be obtained by injecting a higher inoculum. The same applies to the infectivity experiments with HepG2cells. Response: We increased the sporozoites dose and infected mice with 10,000 hemolymph sporozoites, but no infection was observed (Table 1). No EEFs were observed in HepG2 cells infected with 10,000 S14 KO hemolymph sporozoites.
Please provide information on the number of sporozoites that were analyzed in the trails experiment. Response: We analyzed 210, 225, and 212 sporozoites for WT GFP, S14 KO c1, and S14 KO c2, respectively.
Minor issues:
In Figure 1. F) WB on S14-3xHA-mCherry tagged sporozoites showing two bands on the WB. The Palm-band is only inferred thus I suggest correcting the figure to S14-3xHA-mcherry. On 1D all the mcherry signal is detected on the membrane but then on WB, a smaller fraction is palm? What is the explanation for the ratio between the two bands? Why so distinct CSP intensity bands between wt and tagged line? Were very distinct amounts of protein loaded?
Response: We have corrected the Palm-S14-3xHA-mcherry to S14-3xHA-mcherry.
This reviewer raises a valid point regarding the discrepancy between IFA and Western blot. The non-palmitoylated S14-mCherrey signal was possibly corrected after deconvolution in image 1D and mainly the membrane signal was prominent. In Figure 1C, many sporozoites show some cytosolic signal, perhaps representing non-palmitoylated S14. Western blot concentrates the protein of interest as a single band, allowing more accurate visualization of protein.
The distinct CSP intensity bands between wt and the tagged line are due to the loading of a higher amount of parasite lysate in WT lane. To ensure that the western blot signal is specific to S14, we loaded a higher amount of protein in WT.
Figure 1. A) Statistical analysis is missing. Not clear if the bars represent mean values +/- standard deviation. No information on the material and methods of how the relative expression was calculated. Response: No error bars are shown in Figure 1 because it was performed once.
In the introduction lines 54 and 58 I suggest replacing humans with mammalian host. Response: Replaced.
Line 58. Not clear why the ref Ripp et al., 2021 is used for a general sentence to introduce the Plasmodium life cycle. Response: Removed.
Line 72: I suggest replacing "TRAP mutant" with "TRAP knockouts" (Sultan et al., 1997). More recently there are TRAP mutants with impaired motility and normal invasion of mosquito salivary glands (Klug et al., 2020) Response: Replaced.
Lines 78 to 86: In this paragraph, authors refer to several proteins involved in sporozoite gliding motility and host cell invasion, however for most of the studies this conclusion comes from the characterization of knockouts defective phenotype and actually a direct role for some of these molecules in the process awaits clear demonstration. Response: We have replaced involved with implicated.
Line 78: Authors do not consider that maebl knockout sporozoites display reduced adhesion, including to cultured hepatocytes, which could contribute to the defects in multiple biological processes, such as in gliding motility, hepatocyte wounding, and invasion. Response: We have corrected maebl role in the revised manuscript.
Line 80: I suggest authors reconcile the contradictory reports in the literature on the role of TRSP in sporozoites invasion. Response: We have removed this reference in the revised manuscript.
Line 82-83: Please revise it. Response: Revised.
Table 1. Correct table as when sporozoites were transmitted by mosquito bite the term "number of sporozoites injected" does not apply. Please give more details on the bite experiments. Is this the number of mosquitoes for all four animals? For how long the mosquitoes were allowed to bite? Response: For clarity, we have split the table into A Mosquito bite and B haemolymph Sporozoites. We used ten mosquitoes/mice in the bite experiment. Mosquitos were allowed to probe for blood meal for 20 minutes, and the feeding was ensured by observing mosquitoes post-blood meal; approximately 70% of mosquitoes received the blood meal in all the cages.
Line 288 and 289. There are several publications showing that maebl knockout sporozoites are impaired at invading the mosquito salivary glands and at infecting the vertebrate host contradicting Kariu et al., 2002 findings in the vertebrate host. Response: We have removed maebl from this line.
Line 290. I suggest "was most likely due to" instead of " due to" as sporozoite adhesion to cells was not evaluated. Response: Corrected.
Line 291: "Cellular transmigration and host cell invasion are prerequisites for gliding motility" please revise. Response: Revised.
Line 437: indicate which clone was used.
Response: Indicated (3D11).
Line: 463: indicate the % of the gel in the SDS-PAGE Response: We have used 10% SDS-PAGE gel and it is indicated in the revised manuscript.
Line 499: indicate the version of the GraphPad Prism software. Response: GraphPad Prism version 9.
Figure S3 legend needs to be corrected. Panels in the figure are from A to F while in legend G and H are included. Response: Corrected.
Reviewer #2
Line 39-41: "Using in silico and the yeast two-hybrid system, we showed the interaction of S14 with the glideosome-associated proteins GAP45 and MTIP. Together, our data show that S14 is a glideosome-associated protein" Although these interactions can be speculated based on the results shown, these interactions were not confirmed in this study. Response: We attempted to pulldown the S14 interacting partner using anti-mCherry antibody from S14-3XHA-mCherry transgenic sporozoites and then further tried to identify interactome using mass spectrometry but failed. Hence, we selected two known IMC localized gliding proteins MTIP and GAP45. Performing pull-down from sporozoites is challenging, so we checked this interaction using yeast 2-hybrid assay and bioinformatic analysis for protein-protein interaction.
In order to claim interaction between S14 and IMC proteins, interaction needs to be shown experimentally. Well-controlled yeast 2-hybrid would be a start - then interaction would be more than just speculative. But immunoprecipitation from sporozoites or other biochemical interactions would give more support to this idea. Response: We attempted to pulldown the S14 interacting partner using an anti-mCherry antibody from S14-3XHA-mCherry transgenic sporozoites and then further tried to identify interactome using mass spectrometry but failed. Hence, we selected two known IMC localized gliding proteins MTIP and GAP45. Performing pull-down from sporozoites is challenging, so we checked this interaction using yeast 2-hybrid assay and bioinformatic analysis for protein-protein interaction.
Reviewer #3
The authors provide convincing data on the S14 localization in the inner membrane of sporozoites and interaction with GAP45 and MTIP using the yeast model. Did the authors consider conducting co-IP followed by MS analysis to pull down S14 in the complex with GAP45 and MTIP? Response: We attempted to pulldown the S14 interacting partner using an anti-mCherry antibody from S14-3XHA-mCherry transgenic sporozoites and then further tried to identify the interactome using mass spectrometry but failed. Hence, we selected two known IMC localized gliding proteins, MTIP and GAP45. Performing pull-down from sporozoites is challenging, so we checked this interaction using yeast 2-hybrid assay and bioinformatic analysis for protein-protein interaction.
__Reviewer #3 (Significance (Required)):____ __ Sporozoite gliding motility is a critical feature of parasite infectivity. Impairment of this important feature has been described for several mutant/knockout parasite lines. This study goes beyond the phenotypic analysis of mutant parasites to infer the role of S14 by providing more mechanistic evidence to show S14 interaction with other glideosome-associated proteins. However, this interaction was investigated using the two-hybrid system in yeast. Still, in sporozoites, no experiments were conducted to evaluate the interaction between these proteins.
Response: We attempted to pulldown the S14 interacting partner using an anti-mCherry antibody from S14-3XHA-mCherry transgenic sporozoites and then further tried to identify interactome using mass spectrometry but failed. Hence, we selected two known IMC localized gliding proteins, MTIP and GAP45. Performing pull-down from sporozoites is challenging, so we checked this interaction using yeast 2-hybrid assay and bioinformatic analysis for protein-protein interaction.
Please consider I'm not an expert on the in-silico interaction studies.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary:
The authors tag the sporozoite protein S14 in P. berghei and show localization near the sporozoite plasma membrane. They also convincingly show, through the generation of S14 knockout lines, that S14 is required for sporozoite motility and thereby also salivary gland and hepatocyte invasion. Their bioinformatic results support possible interactions between S14 and the inner membrane complex proteins MTIP and GAP45. These analyses were performed with these specific candidate proteins rather than being unbiased searches for potential interaction partners. The yeast 2-hybrid data to support these possible protein interactions need further controls.
Major comments:
Line 39-41: "Using in silico and the yeast two-hybrid system, we showed the interaction of S14 with the glideosome-associated proteins GAP45 and MTIP. Together, our data show that S14 is a glideosome-associated protein" Although these interactions can be speculated based on the results shown, these interactions were not confirmed in this study.
Lines 143-144: Unless the sporozoites were not permeablized prior to staining, it is not clear if the protein is "on" the plasma membrane or just under the plasma membrane. Furthermore, this statement anyway seems contradictory to the authors' interpretation of Figure 4A.
Line 218: "This result indicates that S14 is present within the inner membrane of sporozoites." While this data shows that S14 is not in the plasma membrane of the parasite, how can the authors be sure it is at the IMC?
Line 149: To definitively state S14 is a membrane protein, biochemical assays proving such should be performed. (or perhaps genetic mutation of the predicted palmitoylation site?) Otherwise, this should be rephrased.
Line 225-226: This sentence overreaches in its conclusion. There is no indication that this protein provides the power or force behind the sporozoites forward movement. Several proteins are known to be required for gliding motility, but they are not all force-providing factors.
Lines 257-258: for yeast 2-hybrid, the controls of expressing S14, GAP45 and MTIP together with control proteins where no interaction would be predicted are absent.
In order to claim interaction between S14 and IMC proteins, interaction needs to be shown experimentally. Well-controlled yeast 2-hybrid would be a start - then interaction would be more than just speculative. But immunoprecipitation from sporozoites or other biochemical interactions would give more support to this idea.
Minor comments:
Line 99: "the role of gliding-associated proteins is unexplored" There are several publications on GAP40, GAP45 and GAP50 (some of which are referenced in the previous paragraph).
Line 114: "We narrowed it down to a candidate" Narrowed down how? Or rephrase.
Lines 120-123 are strangely written, and I don't follow the logic. What "similar properties" do GAP45 and GAP50 have with S14 and are they really indicative of function? Also if palmitoylation and myristylation and nonclassical secretion are present in most eukaryotes, why would they necessarily be evidence of IMC targeting?
Line 148-149. I did not see examples of this electromobility shift of GAP45 in this publication (although I may have overlooked it).
Table 1 legend should preferably specify that hemolymph sporozoites were used for IV infections.
Line 228: Should be rephrased for accuracy. "revealed the" should be replaced with "suggests"
Lines 305-307: I don't entirely understand the logic laid out here.
Lines 320-322: "We hypothesize that S14 possibly plays a structural role and maintains the stability of IMC required for the activity of motors during gliding and invasion." The data about the IMC structure shown is fluorescence microscopy - and there no change is observed in the IMC in the knockout line. I suggest removing or rephrasing this point if no extra data is provided to show this.
The work gives insights into an unstudied, conserved Plasmodium protein, S14, which the authors show is critical for Plasmodium transmission from mosquitoes. The parasite genetics and phenotyping demonstrating this are strong. The conclusions about interactions with glideosome/inner membrane complex components need further experimental support. The work is of interest to the Plasmodium field and may be also of interest to people interested in other protozoan parasites or in cellular motility.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary: The authors have identified a sporozoite gliding motility protein through bioinformatic analysis. From the main text I do not know how, or what bioinformatic analysis was performed, in order to focus on this protein which is called S14. The authors then go on to tag the protein, produce a KO and show its involvement in gliding motility. The KO shows that parasites lacking S14 fail to invade the mosquito salivary glands. This is due to a motility defect. Y2H and docking studies are used to define an interaction with MTIP and GAP45, two known components of the glideosome.
Major comments: The paper is sometimes hard to follow and lacks clarity. The reason: important information is omitted, or explained at the end of a section rather than at first mention; experimental details that are of essence need to be mentioned or explained in the main text; there is ample use of the word 'bioinformatic' without explaining what kind of analysis was performed in the main text. I cite from the abstract: 'In silico analysis of a novel protein, S14, which is uniquely upregulated in salivary gland sporozoites, suggested its association with glideosome-associated proteins.' I cite from the introduction: 'A study comparing transcriptome differences between sporozoites and merozoites using suppressive subtraction hybridization found several genes highly upregulated in sporozoites and named them 'S' genes (Kaiser et al, 2004). We narrowed it down to a candidate named S14, which lacked signal peptide and transmembrane domains.' From reading the main text, I do not know why Plasmodium berghei S14 was chosen in this manuscript. S14 is one of 25 transcripts identified by Kappe et al in Plasmodium yoelii (https://doi.org/10.1046/j.1365-2958.2003.03909.x) to be upregulated in sporozoites. The material and methods section does not explain either why S14 was chosen. Perhaps the authors could update Figure 2 from Kappe et al with the most recent annotations from plasmodb.
Reproducibility: None of the main Figures or Figure legends define ' N = '. For example I cite: 'The S14 KO clonal lines were first analyzed for asexual blood-stage propagation, and for this, 200 µl of iRBCs with 0.2% parasitemia was intravenously injected into a group of mice.' There are 2 mentions of 'N=' in the supplementary figures. I have not found any others.
I'm not sure what the convention is. Should unpublished data for this gene (PBANKA_0605900) found in pberghei.eu (a database for mutant berghei parasites) be cited? After all it confirms their findings.
The authors need to use more recent references for some of their statements; see some comments below.
Minor comments:
line
1-2 Add the Plasmodium species of this study. abstract Which species do you work with? 29 mosquito salivary glands and human host hepatocytes 30 to the glideosome, a protein complex containing [...] 32-33 What kind of in silico analysis suggested S14 is part of the glideosome? S14 is not uniquely upregulated; there are other S-type genes identified by Kappe and Matuschewski. 25 I believe. 32 Please point out he species were S genes were identified. SGS of which species? 34 expression: change to transcription 39 What kind of in silico analysis was used here? and therefore malaria transmission 55 A single zygote transforms into a single ookinete, which establishes a single oocyst, which in turn can produce thousands of midgut sporozoites. Please correct the life cycle passage. located or anchored in the IMC? And located between the IMC and plasma membrane? 61-63 Refer to Table S1 and its contents here 64 Name the known GAPs.
65-67 Which transmembrane domain proteins? Please add more recent references than King 1988. 71-72 TRAP was the first protein found to be ... 74-76 Add additional, more recent references: for example search Frischknecht and TRAP 76 S6 (TREP) is also [...] 88 Some of these proteins are also expressed in ookinetes. 89-91 The sentence needs a verb. 88-96 Please add some more recent glideosome papers. After 2013. 91 Why do you call it a peripheral protein? 91-93 There are more recent citations for GAP45 andGAP50. 96 Insert a reference here. 99 Please define the gliding-associated proteins. What are they? Aren't there papers on GAP40, 45 and 50? DOI: 10.1016/j.chom.2010.09.002 99 .... What prompted you to identify a novel GAP? And why is S14 classified as a GAP? 99-102 What kind of bioinformatic study? Why was S14 chosen? Please outline how you ended up with S14. Any other proteins that came out of the bioinformatic screen from the list of S genes? How many proteins were identified in the screen for sporozoite upregulated proteins by Kappe and Matuschewski? 102-103 Define the nonclassical secretion pathway. Please reference GAP45 and GAP50 data for the nonclassical pathway. 105 Please add P. berghei to the title, the abstract, the introduction. 111 The results section does not outline what bioinformatic analysis was used 112-114 Please specify the exact number of upregulated in sporozoites genes. I think it was 25. And add the species the study was performed in. Why did you choose the Kappe study but not the uis genes from berghei? 114-115 How did you narrow it down to S14? The Kappe paper lists 25 S-type genes from P. yoelii. 118 Plasmodia is not the plural for a group of different Plasmodium species. Use: [...] conserved among Plasmodium spp. 118-119 Which proteins did you analyze? And how did you analyze them? Where is the data for this analysis? Outline the amino acids that predict palmitoylation? The nonclassical pathway? 119-122 Here: do you mean S14 has similar properties as GAP 45 and GAP50? Define the nonclassical pathway? How do you know S14 is in the IMC? 122-123 Please reference the bioinformatic analysis plus URL that allows targeting to the IMC to be analyzed. 123-124 Please reference the URLs for TM, palmitoylation, and interactions analyses. 125-127 How did you predict that S14 is secreted via the nonclassical pathway? 128-130 Define the nonclassical pathway when it first appears in your manuscript. The citation Moskes 2004 is not in the reference list 132 Which membrane? 134-135 In which species? 141-142 Please include images of blood stage and liver stage parasites. 142-143 Which membrane? 148-149 I cannot find the specific figure you refer to; I checked the online version of the Frenal 2010 paper. 175 gland, we counted [...] 177 Compared to the 177-179 Failed to invade (absolutely)? Or invaded in highly reduced numbers? 182-186 Please be precise: I think you mean you let all types of mosquitoes take a blood meal; s14 knockout-infected mosquitoes did not infect mice. 181-202 Perhaps use paragraphs to indicate the different types of experiments performed here. 204 Please introduce paragraphs to identify the different experiments in this section 208 Outer or inner membrane of what? IMC, the plasma membrane? 228 onwards Structural models were obtained from whom? Which species did you use for the docking study? Could you use in one approach 3 berghei proteins, and confirm your docking studies with the falciparum proteins? That would strengthen your model. Should you include a negative control protein in the approach? 250-251 Was all of the gene cloned? Please define amino acid range. discussion Please discuss data from https://elifesciences.org/articles/77447 in relation to your protein
298-300 More recent glideosome papers exist. For example https://doi.org/10.1038/s42003-020-01283-8 340 List the proteins you analysed. Add URL (websites) to the analyses tools. 343 Known association from the literature: how was this done? 346-349 A few glideosome components? On what basis were they selected and which are they?
471 Can AlphaFold Structure Predictions be used in the docking studies? 487 What parts of theses genes was cloned? Define the amino acid range. 714 Please split the table into A Mosquito bite and B haemolymph Sporozoites Figure 1 For clarity, maybe write S14::mCherry Figure 1 It would be useful to show blood stage parasite images. Figure 1F You have not formally shown that this signal corresponds to palmitoylated S14. Could be heavy chain. Figure 2G Haemolymph sporozoites ? Figure 8 You argued that S14 is a membrane-bound protein through palmitoylation. Here the protein is shown to be cytoplasmic. Please update our model with more recent ones.
Figure S2B It would be good to include a positive control for these PCRs. Figure S3 It would be good to include a positive control for these PCRs.
Tabel S1 Table S1 is only mentioned twice in the text: lines 124 and 128. There is no mention that the table contains all (??) known gliding motility proteins. Table S1 The algorithms / websites used for bioinformatic prediction need to be listed here. Table S2 Add the plasmodb gene identifiers here. The table does not show all Plasmodium spp. but a selection.
General assessment: The authors provide an in-depth analyses of the Plasmodium berghei protein S14 and its involvement in gliding motility.
Advance: This paper is the first analysis of the S14 protein. The authors suggest a bridging function for the protein between MTIP and GAP45.
Audience: Gliding motility is of interest to the apicomplexan field. I think this particular proteins is specific to Plasmodium spp.
I think we are a victim of behavioural norms and so many of the apps that I use have this pattern. That's not to say it's the right behaviour, but it may be hard to break the pattern for users.
Wenige Tage nach dem „2023 State of the Climate report“ listet der „Interconnected Disaster Report“ der UN University sechs Bedrohungen für die lebenserhaltenden Systeme des Planeten auf, darunter Gletscherschmelze, Grundwasserverlust, Artensterben und Erhitzung. Nur ein transformativer Wandel statt aufschiebender Lösungen kann die wachsenden Risiken miteinander verbundener Katastrophen verringern. https://www.liberation.fr/environnement/climat/climat-la-vie-sur-la-planete-terre-est-en-etat-de-siege-alertent-des-scientifiques-20231025_2MHSFZZWUNHZPPBONAUFLKKUZM/
Studie: https://interconnectedrisks.org/
Merh zum 2023 State of the Climate report: https://hypothes.is/search?q=tag%3A%22report%3A%202023%20state%20of%20the%20climate%20report%22
Note: This response was posted by the corresponding author to Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Dear Editor,
Herewith we submit our fully revised peer-reviewed preprint that had been reviewed by Review Commons. We thank the Review Commons team and reviewers for thoroughly commenting on our preprint and providing very useful additional points for consideration and discussion.
You will find - the revised manuscript (third preprint version uploaded on biorxiv)<br /> - two reviewer letters (through Review Commons), - our rebuttal letter<br /> - a revised manuscript version with highlighted changes.
Our manuscript reports that an active form of FIT, an essential transcription factor for root iron acquisition in plants, forms dynamic nuclear condensates in response to a blue light stimulus.<br /> A hallmark of our work is the thorough investigation of the nature of the FIT nuclear bodies in plant cells, that we were able to characterize as highly dynamic condensates in which active FIT homo- and heteromeric protein complexes can accumulate preferentially. Through co-localization with nuclear body markers, we found that these FIT condensates are related to speckles, which are a sub-type of nuclear bodies connected with splicing activities. This suggests that FIT condensates are linked with post-transcriptional regulation mechanisms.
The reviewers highlight that an “impressive set of microscopic techniques” has been combined to study in a unique manner the characteristics and functionalities of FIT nuclear bodies in living plant cells. We show that FIT nuclear bodies can be formed in roots of Arabidopsis thaliana. The microscopic imaging techniques we used to characterize the nature and functionalities of FIT nuclear bodies in plant cells have several constraints related to sensitivity and a required strength of fluorescent protein signal. For technical reasons to be able to apply qualitative and quantitative imaging techniques, we conducted the investigation of FIT condensates in Nicotiana benthamiana, a classical and widely used plant protein expression system.
As stated in the reviews, the connection between plant nutrition and nuclear bodies is an “unprecedented” new mode of regulation. The significance of our work is underlined by the fact that we report a “very precise cellular and molecular mechanism in nutrition” that is as yet “still largely unexplored in this context”. Therefore, our study “sheds light on the functional role of this membrane-less compartment and will be appreciated by a large audience.”
We propose that condensate formation is a mechanism that may steer iron nutrition responses by providing a link between iron and light signaling. For sessile plants, it is absolutely essential that environmental signals are sensed and integrated with developmental and physiological programs so that plants can rapidly adjust to a changing environment and potential stress situations. Since iron is a micronutrient that may be toxic when present in excess, e.g. through catalyzing oxidative stress, plants strictly control the acquisition and allocation of iron. Hence, FIT nuclear bodies may be regulatory hubs that integrate at the sub-nuclear level environmental signaling inputs in the control of micronutrient uptake, possibly connected with splicing.
Our work lays the ground for future studies that can address the proof of concept in more detailed manner in plants exposed to varying environmental conditions to reveal the interconnection of environmental and nutritional signaling.
We prepared a revised preprint in which we address all reviewer comments. Please find our revision and our detailed response to all reviewer comments.
With these changes, we hope that our peer-reviewed preprint can receive a positive vote,
We are looking forward to your response,
Sincerely
Petra Bauer and Ksenia Trofimov on behalf of all authors
Reviewer #1 (Evidence, reproducibility and clarity):
In this paper entitled " FER-LIKE IRON DEFICIENCY-INDUCED TRANSCRIPTION FACTOR (FIT) accumulates in homo- and heterodimeric complexes in dynamic and inducible nuclear condensates associated with speckle components", Trofimov and colleagues describe for the first time the function of FIT in nuclear bodies. By an impressive set of microscopies technics they assess FIT localization in nuclear bodies and its dynamics. Finally, they reveal their importance in controlling iron deficiency pathway. The manuscript is well written and fully understandable. Nonetheless, at it stands the manuscript present some weakness by the lack of quantification for co-localization and absence controls making hard to follow authors claim. Moreover, to substantially improve the manuscript the authors need to provide more proof of concepts in A. thaliana as all the nice molecular and cellular mechanism is only provided in N. bentamiana. Finally, some key conclusions in the paper are not fully supported by the data.<br /> Please see below:
Main comments:
1) For colocalization analysis, the author should provide semi-quantitative data counting the number of times by eyes they observed no, partial or full co-localization and indicate on how many nucleus they used.
Authors:
We have added the information in the Materials and Method section, lines 731-734:
In total, 3-4 differently aged leaves of 2 plants were infiltrated and used for imaging. One infiltrated leaf with homogenous presence of one or two fluorescence proteins was selected, depending on the aim of the experiment, and ca. 30 cells were observed. Images are taken from 3-4 cells, one representative image is shown.
In all analyzed cases, except in the case of colocalization of FIT and PIF4 fusion proteins, the ca. 30 cells had the same localization and/or colocalization patterns. This information has also been added in the figure legends. Each experiment was repeated at least 2-3 times, or as indicated in the figure legend.
2) Do semi-quantitative co-localization analysis by eyes, on FIT NB with known NB makers in the A. thaliana root. For now, all the nicely described molecular mechanism is shown in N. benthamiana which makes this story a bit weak since all the iron transcriptional machinery is localized in the root to activate IRT1.
Authors:
The described approach has been very optimal, and we were able to screen co-localizing marker proteins in FIT NBs in N. benthamiana to better identify the nature of FIT NBs. This has been successful as we were able to associate FIT NBs with speckles. The N. benthamiana system allowed optimal microscopic observation of fluorescence proteins and quantification of FIT NB characteristics in contrast to the root hair zone of Arabidopsis where Fe uptake takes place. FIT is expressed at a low level in roots and also in leaves, whereby fluorescence protein expression levels are insufficient for the here-presented microscopic studies. The tobacco infiltration system is also well established to study FIT-bHLH039 protein interaction and nuclear body markers. We discuss this point in the discussion, see line 489-500.
3) The authors need to provide data clearly showing that the blue light induce NB in A. thaliana and N. benthamiana.
Authors:
For tobacco, see Figure 1B (t = 0, 5 min) and Supplemental Movies S1. For Arabidopsis, please see Figure 1A (t = 0, 90 and 120 min) and Supplemental Figure S1A. We provide an additional image of pFIT:cFIT-GFP Arabidopsis control plants, showing that NB formation is not detected in plants that were grown in white light and not exposed to blue light before inspection (Supplemental Figure S1B). We state, that upon blue light exposure, plants had FIT NBs in at least 3-10 nuclei of 20 examined nuclei in the root epidermis in the root hair zone (in three independent experiments with three independent plants). White-light-treated plants showed no NB formation unless an additional exposure to blue light was provided (in three independent experiments, three independent plants per experiment and with 15 examined nuclei per plant).
4) Direct conclusion in the manuscript:
- Line 170: At this point of the paper the author cannot claim that the formation of FIT condensates in the nucleus is due to the light as it might be indirectly linked to cell death induced by photodamaging the cell using a 488 lasers for several minutes. This is true especially with the ELYRA PS which has strong lasers made for super resolution and that Cell death is now liked to iron homeostasis. The same experiment might be done using a spinning disc or if the authors present the data of the blue light experiment mentioned above this assumption might be discarded. Alternatively, the author can use PI staining to assess cell viability after several minutes under 488nm laser.
Authors:
As stated in our response to comment 3, we have included now a white light control to show that FIT NB formation is not occurring under the normal white light conditions. Since the formation of FIT NBs is a dynamic and reversible process (Figure 1A), it indicates that the cells are still viable, and that cell death is not the reason for FIT NB formation.
- Line 273: I don't agree with the first part of the authors conclusion, saying that "wild-type FIT had better capacities to localize to NBs than mutant FITmSS271AA, presumably due its IDRSer271/272 at the C-terminus. This is not supported by the data. In order to make such a claim the author need to compare the FA of FIT WT with FITmSS271AA by statistical analysis. Nonetheless, the value seems to be identical on the graphs. The main differences that I observed here are, 1) NP value for FITmSS271AA seems to be lower compared to FIT-WT, suggesting that the Serine might be important to regulate protein homedimerization partitioning between the NP and the NB. 2) To me, something very interesting that the author did not mention is the way the FA of FITmSS271AA in the NB and NP is behaving with high variability. The FA of those is widely spread ranging from 0.30 to 0.13 compared to the FIT-WT. To me it seems that according to the results that the Serine 271/272 are required to stabilize FIT homodimerization. This would not only explain the delay to form the condensate but also the decreased number and size observed for FITmSS271AA compared to FIT-WT. As the homodimerization occurs with high variability in FITmSS271AA, there is less chance that the protein will meet therefore decreasing the time to homodimerize and form/aggregate NB.
Authors:
We fully agree. We meant to describe this result it in a similar way and thank you for help in formulating this point even better. Rephrasing might make it better clear that the IDRSer271/272 is important for a proper NB localization, lines 272-278:
“Also, the FA values did not differ between NBs and NP for the mutant protein and did not show a clear separation in homodimerizing/non-dimerizing regions (Figure 3D) as seen for FIT-GFP (Figure 3C). Both NB and NP regions showed that homodimers occurred very variably in FITmSS271AA-GFP.
In summary, wild-type FIT could be partitioned properly between NBs and NP compared to FITmSS271AA mutant and rather form homodimers, presumably due its IDRSer271/272 at the C-terminus.”
- Line 301: According to my previous comment (line 273), here it seems that the Serine 271/272 are required only for proper partitioning of the heterodimer FIT/BHLH039 between the NP and NB but not for the stability of the heterodimer formation. However, it might be great if the author would count the number of BHLH039 condensates in both version FITmSS271AA and FIT-WT. To my opinion, they would observe less BHLH039 condensate because the homodimer of FITmSS271AA is less likely to occur because of instability.
Authors:
bHLH039 alone localizes primarily to the cytoplasm and not the nucleus, and the presence of FIT is crucial for bHLH039 nuclear localization (Trofimov et al., 2019). Moreover, bHLH039 interaction with FIT depends on SS271AA (Gratz et al., 2019). We therefore did not consider this experiment for the manuscript and did not acquire such data, as we did not expect to achieve major new information.
5) To wrap up the story about the requirements of NB in mediating iron acquisition under different light regimes, provide data for IRT1/FRO2 expression levels in fit background complemented with FITmSS271AA plants. I know that this experiment is particularly lengthy, but it would provide much more to this nice story.
Authors:
Data for expression of IRT1 and FRO2 in FITmSS271AA/fit-3 transgenic Arabidopsis plants are provided in Gratz et al. (2019). To address the comment, we did here a NEW experiment. We provide gene expression data on FIT, BHLH039, IRT1 and FRO2 splicing variants (previously reported intron retention) to explore the possibility of differential splicing alterations under blue light (NEW Supplemental Figure S6 and S7, lines 454-466). Very interestingly, this experiment confirms that blue light affects gene expression differently from white light in the short-term NB-inducing condition and that blue light can enhance the expression of Fe deficiency genes despite of the short 1.5 to 2 h treatment. Another interesting aspect was that the published intron retention was also detected. A significant difference in intron retention depending on iron supply versus deficiency and blue/white light was not observed, as the pattern of expression of transcripts with respective intron retentions sites was the same as the one of total transcripts mostly spliced.
Minor comments
In general, I would suggest the author to avoid abbreviation, it gets really confusing especially with small abbreviation as NB, NP, PB, FA.
Authors:
We would like to keep the used abbreviations as they are utilized very often in our work and, in our eyes, facilitate the understanding.
Line 106: What does IDR mean?
Authors:
Explanation of the abbreviation was added to the text, lines 105-108:
“Intrinsically disordered regions (IDRs) are flexible protein regions that allow conformational changes, and thus various interactions, leading to the required multivalency of a protein for condensate formation (Tarczewska and Greb-Markiewicz, 2019; Emenecker et al., 2020).”
Line 163-164: provide data or cite a figure properly for blue light induction.
Authors:
We have removed this statement from the description, as we provide a white light control now, lines 157-158:
“When whole seedlings were exposed to 488 nm laser light for several minutes, FIT became re-localized at the subnuclear level.”
Line 188: Provide Figure ref.
Authors:
Figure reference was added to the text, lines 184-185:
“As in Arabidopsis, FIT-GFP localized initially in uniform manner to the entire nucleus (t=0) of N. benthamiana leaf epidermis cells (Figure 1B).”
Line 194: the conclusion is too strong. The authors conclude that the condensate they observed are NB based on the fact the same procedure to induce NB has been used in other study which is not convincing. Co-localization analysis with NB markers need to be done to support such a claim. At this step of the study, the author may want to talk about condensate in the nucleus which might correspond to NB. Please do so for the following paragraph in the manuscript until colocalization analysis has not been provided. Alternatively provide the co-localization analysis at this step in the paper.
Authors:
We agree. We changed the text in two positions.
Lines 176-178__: “__Since we had previously established a reliable plant cell assay for studying FIT functionality, we adapted it to study the characteristics of the prospective FIT NBs (Gratz et al., 2019, 2020; Trofimov et al., 2019).”
Lines 192-193: “__We deduced that the spots of FIT-GFP signal were indeed very likely NBs (for this reason hereafter termed FIT NBs).”
Line 214: In order to assess the photo bleaching due to the FRAP experiment the quantification of the "recovery" needs to be provided in an unbleached area. This might explain why FIT recover up to 80% in the condensate. Moreover, the author conclude that the recovery is high however it's tricky to assess since no comparison is made with a negative/positive control.
Authors:
In the FRAP analysis, an unbleached area is taken into account and used for normalization.
We reformulated the description of Figure 1F, lines 212-214:
“According to relative fluorescence intensity the fluorescence signal recovered rapidly within FIT NBs (Figure 1F), and the calculated mobile fraction of the NB protein was on average 80% (Figure 1G).”
Line 220-227: The conclusion it's too strong as I mentioned previously the author cannot claim that the condensate are NBs at this step of the study. They observed nuclear condensates that behave like NB when looking at the way to induce them, their shape, and the recovery. And please include a control.
Authors:
Please see the reformulated sentences and our response above.
Lines 176-178: “Since we had previously established a reliable plant cell assay for studying FIT functionality, we adapted it to study the characteristics of the prospective FIT NBs (Gratz et al., 2019, 2020; Trofimov et al., 2019).”
Lines 192-193: “__We deduced that the spots of FIT-GFP signal were indeed very likely NBs (for this reason hereafter termed FIT NBs).”
Line 239: It's unappropriated to give the conclusion before the evidence.
Authors:
Thank you. We removed the conclusion.
Line 240: Figure 2A, provide images of FIT-G at 15min in order to compare. And the quantification needs to be provided at 5 minutes and 15 minutes for both FIT-G WT and FIT-mSS271AA-G counting the number of condensates in the nucleus. Especially because the rest of the study is depending on these time points.
Authors:
This information is provided in the Supplemental Movie S1C.
Line 241: the author say that the formation of condensate starts after 5 minutes (line 190) here (line 241) the author claim that it starts after 1 minutes. Please clarify.
Authors:
In line 190 we described that FIT NB formation occurs after the excitation and is fully visible after 5 min. In line 241 we stated that the formation starts in the first minutes after excitation, which describes the same time frame. We rephrased the respective sentences.
Lines 185-188: “A short duration of 1 min 488 nm laser light excitation induced the formation of FIT-GFP signals in discrete spots inside the nucleus, which became fully visible after only five minutes (t=5; Figure 1B and Supplemental Movie S1A).”
Lines 239-242: “While FIT-GFP NB formation started in the first minutes after excitation and was fully present after 5 min (Supplemental Movie S1A), FITmSS271AA-GFP NB formation occurred earliest 10 min after excitation and was fully visible after 15 min (Supplemental Movie S1C).”
Line 254: Not sure what the authors claim "not only for interaction but also for FIT NB formation ". To me, the IDR is predicted to be perturbed by modeling when the serines are mutated therefore the IDR might be important to form condensates in the nucleus. Please clarify.
Authors:
The formation of nuclear bodies is slow for FITmSS271AA as seen in Figure 2. Previously, we showed that FITmSS271AA homodimerizes less (Gratz et al., 2019.) Therefore, the said IDR is important for both processes, NB formation and homodimerization. We have added this information to make the point clear, lines 253-255:
“This underlined the significance of the Ser271/272 site, not only for interaction (Gratz et al., 2019) but also for FIT NB formation (Figure 2).”
Line 255: It's not clear why the author test if the FIT homodimerization is preferentially associated with condensate in the nucleus.
Authors:
We test this because both homo- and heterodimerization of bHLH TFs are generally important for the activity of TFs, and we unraveled the connection between protein interaction and NB formation. We state this in lines 228-232.
Line 269-272: It's not clear to what the authors are referring to.
Authors:
We are describing the homodimeric behavior of FIT and FITmSS271AA assessed by homo-FRET measurements that are introduced in the previous paragraph, lines 256-268.
Line 309: This colocalization part should be presented before line 194.
Authors:
We find it convincing to first examine and characterize the process underlying FIT NB formation, then studying a possible function of NBs. The colocalization analysis is part of a functional analysis of NBs. We thank the reviewer for the hint that colocalization also confirms that indeed the nuclear FIT spots are NBs. We will take this point and discuss it, lines 516-522:
“Additionally, the partial and full colocalization of FIT NBs with various previously reported NB markers confirm that FIT indeed accumulates in and forms NBs. Since several of NB body markers are also behaving in a dynamic manner, this corroborates the formation of dynamic FIT NBs affected by environmental signals.”
“In conclusion, the properties of liquid condensation and colocalization with NB markers, along with the findings that it occurred irrespective of the fluorescence protein tag preferentially with wild-type FIT, allowed us to coin the term of ‘FIT NBs’.”
Line 328: add the ref to figure, please.
Authors:
Figure reference was added to the text, lines 330-332:
“The second type (type II) of NB markers were partially colocalized with FIT-GFP. This included the speckle components ARGININE/SERINE-RICH45-mRFP (SR45) and the serine/arginine-rich matrix protein SRm102-mRFP (Figure 5).”
Line 334: It seems that the size of the SR45 has an anormal very large diameter between 4 and 6 µm. In general a speckle measure about 2-3µm in diameter. Can the author make sure that this structure is not due to overexpression in N. benthamiana or make sure to not oversaturate the image.
Authors:
Thank you for this hint. Indeed, there are reports that SR45 is a dynamic component inside cells. It can redistribute depending on environmental conditions and associate into larger speckles depending on the nuclear activity status (Ali et al., 2003). We include this reference and refer to it in the discussion, lines 557-564:
“Interestingly, typical FIT NB formation did not occur in the presence of PB markers, indicating that they must have had a strong effect on recruiting FIT. This is interesting because the partially colocalizing SR45, PIF3 and PIF4 are also dynamic NB components. Active transcription processes and environmental stimuli affect the sizes and numbers of SR45 speckles and PB (Ali et al., 2003; Legris et al., 2016; Meyer, 2020). This may indicate that, similarly, environmental signals might have affected the colocalization with FIT and resulting NB structures in our experiments. Another factor of interference might also be the level of expression.”
Line 335: It seems that the colocalization is partial only partial after induction of NB. The FIT NB colocalize around SR45. But it's hard to tell because the images are saturated therefore creating some false overlapping region.
Authors:
The localization of FIT with SR45 is partial and occurs only after FIT has undergone condensation, see lines 335-338.
Line 344-345: It's unappropriated to give the conclusion before the evidence.
Authors:
We explain at an earlier paragraph that we will show three different types of colocalization and introduce the respective colocalization types within separate paragraphs accordingly, see lines 314-321.
Line 353: increase the contrast in the image of t=5 for UAP56H2 since it's hard to assess the colocalization.
Authors:
This is done as noted in the figure legend of Figure 6.
Line 381-382: "In general" does not sound scientific avoid this kind of wording and describe precisely your findings.
Authors:
We rephrased the sentence, line 387-388:
Localization of single expressed PIF3-mCherry remained unchanged at t=0 and t=15 (Supplemental Figure S5A).
Line 384-385: Provide the data and the reference to the figure.
Authors:
We apologize for the misunderstanding and rephrased the sentence, line 389-391:
After 488 nm excitation, FIT-GFP accumulated and finally colocalized with the large PIF3-mCherry PB at t=15, while the typical FIT NBs did not appear (Figure 7A)
Line 386: The structure in which FIT-G is present in the Figure 7A t=15 is not alike the once already observed along the paper. This could be explained by over-expression in N. benthamiana. Please explain.
Authors:
Thank you for the hint. We discuss this in the discussion part, see lines 555-568.
Line 393: Explain and provide data why the morphology of PIF4/FIT NB do not correspond to the normal morphology.
Authors:
Thank you for the valuable hints. Several reasons may account for this and we provide explanations in the discussion, see lines 555-568.
Line 396-398: It seems also from the data that co-expression of PIF4 of PIF3 will affect the portioning of FIT between the NP and the NB.
Authors:
We can assume that residual nucleoplasm is depleted from protein during NB formation. This is likely true for all assessed colocalization experiments. We discuss this in lines 492-494.
The discussion is particularly lengthy it might be great to reduce the size and focus on the main findings.
Authors:
We shortened the discussion.
Referees cross-commenting
All good for me, I think that the comments/suggestions from Reviewer #2 are valid and fair. If they are addressed they will improve considerably the manuscript.
Reviewer #1 (Significance):
This manuscript is describing an unprecedent very precise cellular and molecular mechanism in nutrition throughout a large set of microscopies technics. Formation of nuclear bodies and their role are still largely unexplored in this context. Therefore, this study sheds light on the functional role of this membrane less compartment and will be appreciated by a large audience. However, the fine characterization is only made using transient expression in N. Bentamiana and only few proofs of concept are provided in A. thaliana stable line.
Reviewer #2 (Evidence, reproducibility and clarity):
The manuscript of Trofimov et al shows that FIT undergoes light-induced, reversible condensation and localizes to nuclear bodies (NBs), likely via liquid-liquid phase separation and light conditions plays important role in activity of FIT. Overall, manuscript is well written, authors have done a great job by doing many detailed and in-depth experiments to support their findings and conclusions.
However, I have a number of questions/comments regarding the data presented and there are still some issues that authors should take into account.
Major points/comments:
1) Authors only focused on blue light conditions. Is there any specific reason for selecting only blue light and not others (red light or far red)?
Authors:
There are two main reasons: First, in a preliminary study (not shown) blue light resulted in the formation of the highest numbers of NBs. Second, iron reductase activity assays and gene expression analysis under different light conditions showed a promoting effect under blue light, but not red light or dark red light (Figure 9). This indicated to us, that blue light might activate FIT, and that active FIT may be related to FIT NBs.
2) Fig. 3C and D: as GFP and GFP-GFP constructs are used as a reference, why not taking the measurements for them at two different time points for example t=0 and t=5 0r t=15???
Authors:
Free GFP and GFP-GFP dimers are standard controls for homo-FRET that serve to delimit the range for the measurements.
3) Line 27-271: Acc to the figure 3d, for the Fluorescence anisotropy measurement of NBs appears to be less. Please explain.
Authors:
FA in NBs with FITmSS271AA is variable and the value is lower than that of whole nucleus but not significantly different compared with that in nucleoplasm. We describe the results of Figure 3D in lines 272-275.
4) Figure 4: For the negative controls, data is shown at only t=0, data should be shown at t=5 also to prove that there is no decrease in fluorescence in these negative controls when they are expressed alone without bhlh39 as there is no acceptor in this case.
Authors:
Neither for FIT/bHLH039 nor the FITmSS271AA/bHLH039 pair, there is a significant decrease in the fluorescence lifetime values between t=0 and t=5/15. FIT-G is a control to delimit the range. The interesting experiment is to compare the protein pairs of interest between the different nuclear locations at t=5/15.
5) Line 300-301: In Figure 4D and 4E. Fluorescence lifetime of G measurement at t=0 seems very similar for both FIT-G as well as FITmSS but if we look at the values of t=0 for FIT-G+bhlh039 it is greater than 2.5 and for FITmSS271AA-G+bhlh039 it is less which suggests more heterodimeric complexes to be formed in FITmSS271AA-G+bhlh039. Similar pattern is observed for NBs and NPs, according to the figure 4d and E.
Therefore, heterodimeric complexes accumulated more in case of FITmSS271AA-G+bhlh039 as compared to FIT-G+bhlh039 (if we compare measurement values of Fluorescence lifetime of G of FITmSS271AA-G+bhlh039 with FIT-G+bhlh039).
Please comment and elaborate about this further.
Authors:
These conclusions are not valid as the experiments cannot be conducted in parallel. Since the experiments had to be performed on different days due to the duration of measurements including new calibrations of the system, we cannot compare the absolute fluorescence lifetimes between the two sets.
6) Figure 4: For the negative controls, data is shown at only t=0, data should be shown at t=5 also to prove that there is no decrease in fluorescence in these negative controls when they are expressed alone without bhlh39 as there is no acceptor in this case.
Authors:
Please see our response to your comment 4).
7) Line 439-400: As iron uptake genes (FRO2 and IRT1) are more induced in WT under blue light conditions and FRO2 is less induced in case of red-light conditions. So, what happens to Fe content of WT grown under blue light or red light as compared to WT grown under white light. Perls/PerlsDAb staining of WT roots under different light conditions will add more information to this.
Authors:
We focused on the relatively short-term effects of blue light on signaling of nuclear events that could be related to FIT activity directly, particularly gene expression and iron reductase activity as consequence of FRO2 expression. These are both rapid changes that occur in the roots and can be measured. We suspect that iron re-localization and Fe uptake also occur, however, in our experience differences in metal contents will not be directly significant when applying the standard methods like ICP-MS or PERLs staining.
Minor comments:
Line 75-76: Rephrase the sentence
Authors:
We rephrased the sentence, lines 73-74:
“As sessile organisms, plants adjust to an ever-changing environment and acclimate rapidly. They also control the amount of micronutrients they take up.”
Line 119: Rephrase the sentence
Authors:
We rephrased the sentence, line 118-119:
“Various NBs are found. Plants and animals share several of them, e.g. the nucleolus, Cajal bodies, and speckles.”
Line 235-236: rephrase the sentence
Authors:
We rephrased the sentence, line 232-234:
“In the work of Gratz et al. (2019), the hosphor-mimicking FITmS272E protein did not show significant changes in its behavior compared to wild-type FIT.”
Line 444: Correct the sentence “Fe deficiency versus sufficiency”
Authors:
We corrected that, line 449-451:
“In both, the far-red light and darkness situations, FIT was induced under iron deficiency versus sufficiency, while on the other side, BHLH039, FRO2 and IRT1 were not induced at all in these light conditions (Figure 9I-P).”
Referees cross-commenting
I agree with R1 suggestions/comments and i think manuscript quality will be much better if authors carry out the experiments suggested by R1. I believe this will also strengthen their conclusions.
Reviewer #2 (Significance):
Overall, manuscript is well written, authors have done a nice job by doing several key experiments to support their findings and conclusions. However, the results and manuscript can be improved further by addressing some question raised here. This study is interesting for basic scientists which unravels the crosstalk of light signaling in nutrient signaling pathways.
Note: This response was posted by the corresponding author to Review Commons. The content has not been altered except for formatting.
Learn more at Review Commons
*Reviewer #1 (Evidence, reproducibility and clarity (Required)): *
*In their study, Yamano et al. dissect the mechanism of TBK1 activation and downstream effects, especially in its relation to mitophagy adaptor OPTN. The authors find that OPTN's interaction with ubiquitin and the autophagy machinery, forming contact sites between mitochondria and autophagic membranes, results in TBK1 accumulation and subsequent autophosphorylation. Based on these findings, the authors propose a self-propagating feedback loop wherein OPTN phosphorylation by TBK1 promotes recruitment and accumulation of OPTN to damaged mitochondria and specifically the autophagosome formation site. This formation site is then involved in TBK1 autophosphorylation, and the activated TBK1 can then further phosphorylate other pairs of OPTN and TBK1. A OPTN monobody investigation strengthens their findings. *
*Critique: *
It would be helpful if the authors could more clearly highlight the previous findings in OPTN-TBK1 relationship and which gaps in the understanding their study addresses.* We thank the reviewer for this comment. As suggested, we have highlighted previous findings and detailed in the Discussion how the study advances our understanding of TBK1 activation.
It is not always clear whether experiments have been replicated sufficiently; this should be indicated in the figure descriptions.* In the original manuscript, most of the data shown was derived from duplicated experiments. For the revised version, we repeated experiments as needed to generate the replication necessary (i.e, N = 3) for determining statistical significance. Error bars and statistical significance have been added to the graphs and figure legends accordingly.
During the discussion, references to the figures that indicate conclusions should be added where appropriate.* We thank the reviewer for the suggestion. References to figures have been added were appropriate to the Discussion.
*Figure 1 / Result "OPTN is required for TBK1 phosphorylation and subsequent autophagic Degradation": *
*o In a) the TBK1 and TOMM20 blots feature an image artefact that makes it appear like the blots are stitched together or there was a problem with the digital imager. The quantification in b) seems to be missing replications. *
We found that the artifact came from an automatic pixel interpolation process in Adobe Photoshop when the image was rotated by a small angle. We have provided the original immunoblotting data below as evidence that the data were not stitched from separate images. More accurate representations of the images without the artifact are now shown in Fig1 A of the revised manuscript.
For Fig 1b, the experiment was independently replicated three times with error bars added to each plot on the graph.
*o g) should feature the wt cell line on the same blot for better comparability as well as quantification and replication like done in f) *
As suggested, we have included the WT cell line in the immunoblot (See Fig 1g). In addition, Reviewer 2 asked that we provide data for Penta KO cells without exogenous expression of the autophagy adaptors and expressed concern regarding the lower expression of NDP52 relative to OPTN. To address these issues, we repeated the mitophagy experiments and detected phosphorylated TBK1 in six different cell lines: WT, Penta KO, Penta KO stably expressing OPTN at both low and high expression levels, and Penta KO stably expressing NDP52 at low and high expression levels. Immunoblots of phos-TBK1(pS172), TBK1, OPTN, NDP52, TOMM20, and actin were generated under four different conditions (DMSO, valinomycin for 1 hr, valinomycin for 3 hrs, and valinomycin in the presence of bafilomycin for 3 hrs). In addition, phos-TBK1 abundance in the six cell lines was determined in response to val and baf for 3 hrs and the expression levels of NDP52 and OPTN were similarly determined in response to DMSO. Error bars based on three independent experiments have been incorporated into the data, which are shown in Figure 1g and 1h of the revised manuscript.
*o h) is missing the blots for controls actin and TOMM20 *
Immunoblots for actin and TOMM20 have been added, please see Fig 1i in the revised manuscript.
*o In the text to e/f), the authors write that NDP52 KO effect on pS172 are comparable to controls, though the quantitation in f) indicates that pS172 signal is indeed significantly reduced compared to wt *
The reviewer is correct, the phos-TBK1 (pS172) signal in NDP52 KO cells is reduced compared to that in WT cells, but is only moderately lower in NDP52 KO cells relative to OPTN KO. We regret the error, which has been corrected in the revised manuscript.
*o In the text to h/i), the authors write "there was a significant increase in the TBK1 pS172 signal in cells overexpressing OPTN", though the quantification in i) does not indicate significance levels *
We performed statistical analyses on the phos-TBK1 (pS172) levels between cells with or without OPTN overexpression and have added the degree of significance to Fig 1j. As indicated in the original manuscript, there was a significant increase in phos-TBK1 (pS172) levels when OPTN was overexpressed.
*Figure 2 / Result "OPTN association with the autophagy machinery is required for TBK1 activation": ** o In b), pTBK1 at val 1 hr only features one dot/experiment per cell line *
Three independent replicates of the experiment (val 1 hr) were performed. The levels of phos-TBK1 (pS172), total TBK1, and actin were quantified, and the graph was remade with error bars and statistical significance incorporated. Please see Fig 2b in the revised manuscript.
*o In the text to c), the authors claim that the mutants reduce/abolish the recruitment of OPTN to the autophagosome site. A costain for LC3, as done for SupFig 1b, would be necessary to support that specific claim. *
To address the reviewer’s concern regarding the recruitment of OPTN mutants to the autophagosomal formation site, we performed two different experiments. First, when OPTN WT is recruited to the contact site between the autophagosomal formation site and damaged mitochondria, it should be heterogeneously distributed across mitochondria. In contrast, OPTN mutants that are unable to associate with the autophagosome formation sites should be largely localized to damaged mitochondria since the mutants are still capable of binding ubiquitin. When we examined the mitochondrial distribution of OPTN WT following valinomycin treatment for 1 hr, more than 80% of the Penta KO cells exhibited a heterogeneous distribution, whereas only 10% of the cells showed a similar distribution for OPTN 4LA or OPTN 4LA/F178A (please see Fig 2g in the revised manuscript). Although the OPTN F178A mutant exhibited 50% heterogeneous distribution (Fig 2g), this may be because OPTN F178A retains the ability to interact with ATG9A vesicles. In fact, our previous mitophagy analyses (Keima-based FACS analysis, Yamano et al 2020 JCB), which are strongly correlated with OPTN mitochondrial distribution, showed that the OPTN F178A mutant moderately (~ 60%) induced mitochondrial degradation. This degradation effect was slightly higher (80%) with OPTN WT but significantly lower (9%) with the 4LA/F178A mutant. In the second experiment, Penta KO cells expressing either OPTN WT or the OPTN mutants were immunostained for exogenous FLAG-tagged OPTN, endogenous WIPI2, and HAP60 (a mitochondrial marker) after valinomycin treatment for 1 hr (see Fig 2e and 2f in the revised manuscript). Because LC3B is assembled on the autophagosomal formation site as well as completed autophagosomes, we detected endogenous WIPI2 because WIPI2 is only recruited to autophagosomal formation sites (Dooley et al. 2014 Mol Cell). Confocal microscopy images and their associated quantification data indicate that WIPI2 foci formation during mitophagy was reduced in Penta KO cells expressing the OPTN mutants (4LA, F178A and 4LA/F178A) as compared to Penta KO cells expressing OPTN WT.
*o d) and g) as simple confirmations of KO/KD efficiency might be better suited for the supplemental part, or blots for FIP/ATG be included with the blots in e) and h) *
Based on the reviewer comments, we performed additional experiments related to Figure 2 and have incorporated the new data into the revised figure. The original Figure 2d, e, f, g, h, and I have been moved to supplemental Figure 5.
*o In the text to e), the authors claim that the levels of pS172 in the KO cell lines did not increase during mitophagy, though the blot and quantification in f) seem to indicate an increase. The results therefore don't seem to align completely with the claims that pS172 generation in response to mitophagy requires the autophagy machinery, or that FIP200 and ATG9A rather than ATG5 are critical for TBK1 phosphorylation. *
Although newly generated pS172 TBK1 was reduced in FIP200 KO and ATG9A KO cells relative to WT cells, the signals gradually increased. In the autophagy KO cell lines (FIP200 KO and ATG9A KO), phos-TBK1 accumulates prior to mitophagy stimulation. Although suggesting it is mitophagy-independent, phos-TBK1 accumulation prior to mitophagy stimulation in autophagy KO cell lines complicated interpretation of the results. To avoid this issue, we used siRNA to transiently knock down FIP200 and ATG9A. As shown in the original manuscript (Fig 2g, h, I in the original manuscript, supplementary Fig 5d, e, f in the revised manuscript), knockdown of FIP200 and ATG9A prior to mitophagy induction allowed us to observe mitophagy-dependent phosphorylation of TBK1. This result strongly suggests that the autophagy machinery does induce TBK1 phosphorylation in response to Parkin-mediated mitophagy. However, TBK1 phosphorylation still increases, albeit very slightly, in the FIP200 and ATG9A knock down cells. Thus, it may be reasonable to assume that OPTN-dependent phosphorylation of TBK1 can occur to a certain degree even in the absence of autophagy components. We have noted this in the Discussion.
While conducting experiments for the revised manuscript, we determined that TAX1BP1 is responsible for the accumulation of phos-TBK1 in the autophagy KO cell lines under basal conditions. When TAX1BP1 is knocked down in FIP200 KO or ATG9A KO cells, the basal accumulation of phos-TBK1 was eliminated and then we could observe mitophagy-specific TBK1 phosphorylation (please see Fig 2h, i, j, k in the revised manuscript). These results showed that mitophagy-dependent phos-TBK1 is largely attenuated in FIP200KO and was almost completely eliminated in ATG9A KO cells (Fig 2k in the revised manuscript).
*o f) is missing significance indications. Its description has a typo: "bad" instead of "baf" *
Newly synthesized pTBK1 (pS172) during mitophagy was quantified and statistical significance incorporated into the figure (please see supplementary Fig 5c). The identified typo has been corrected.
*Figure 3 / Result "TBK1 activation does not require OPTN under basal autophagy conditions": *
*o In the text to SupFig2, the authors claim that pS172 levels are significantly elevated, but no significance levels are indicated *
Statistical significance was determined for all proteins shown in original supplementary Fig 2 and the results have been incorporated into the relevant figure. The original supplementary Fig 2 is now supplementary Fig 6.
*o In the text to a), NBR1 is claimed to colocalize with Ub, but no costaining with Ub is shown. The claimed lacking colocalization of OPTN with Ub is not obvious from the images; a quantification might be appropriate. *
Since the anti-NBR1 antibody used in the original manuscript is derived from mouse, we were unable to use it in conjunction with the mouse ubiquitin antibody. Because ubiquitin-positive foci and NBR1-positive foci contain p62 (original Fig 3a) and NBR1 and p62 are known to tightly interact each other (Kirkin et al. 2009 Mol Cell and Sanchez-Martin et al. 2020 EMBO Rep), we stated that "NBR1 colocalizes with Ub". However, the reviewer is correct. To remedy this confusion, we obtained a rabbit anti-NBR1 antibody (a gift from the Masaaki Komatsu group) and used it to co-immunostain with anti-Ub antibodies (please see supplementary Fig 7a of the revised manuscript). NBR1 foci colocalize with both ubiquitin and p62 in FIP200 KO and ATG9A KO cells. Further, based on comments from Reviewer 2, we purchased several anti-TBK1 antibodies and identified one that was able to detect endogenous TBK1 by immunostaining (see Figure 1 for reviewers in our response to Reviewer 2 below). Using this anti-TBK1 antibody, we showed that a part of TBK1 also associates with ubiquitin and p62-positive aggregates.
*o In the text to b), the authors make reference to significant changes, but replication/ quantification/ significance testing is missing. *
We independently performed the same experiments three times. The levels of TBK1, phos-TBK1 (pS172), all five autophagy adaptors, and TOMM20 in both the supernatants and pellets have been quantified with error bars and statistical significance indicated. These results have been incorporated into Figure 3c in the revised manuscript.
*Figure 4b) is missing the pTBK1 data that is referenced in the text. In the text to figure 5 c/d), the authors claim that certain mutants have no significant effect on mitophagy, though d) is missing significance testing *
*Figure 6 c/d/i) appear to be missing replication. *
For Figure 4b, phos-TBK1 was immunoblotted (See Fig 4b of the revised manuscript). For Figure 5b and d, statistical significance was determined for the effect of TBK1 mutations on autophosphorylation and OPTN phosphorylation and the effect of the TBK1 mutants on Parkin-mediated mitophagy. For Figure 6 c/d/I, the experiment was repeated; error bars and statistical significance have been added to the associated graphs.
*Reviewer #1 (Significance (Required)): Removal of damaged mitochondria by the mitophagy pathway provides an important safeguarding mechanism for cells. The Pink1/Parkin mechanism linked to numerous modulators and adaptor proteins ensures an efficient targeting of damaged mitochondria to the phagophore. The Ser/Thr kinase TBK1, in addition of multiple roles in innate immunity, is a major mitophagy regulator as has been revealed by the Dikic and Youle groups in 2016 (Richter et al., PNAS). The mechanistic insights provided by this manuscript add to a growing body of studies of how the autophagy machinery interconnects with cellular signalling networks. Although parts of the results need to be further validated, the data shown is of high quality, revealing an important conceptual advance. The paper is interesting and of general relevance beyond the signalling and autophagy community. *
We would like to thank Reviewer 1 for the comments and suggestions, many of which improved our manuscript. We hope that the reviewer’s comments have been adequately addressed in the revised manuscript.
*Reviewer #2 (Evidence, reproducibility and clarity (Required)): Summary In this manuscript, Yamano and colleagues show that as for Sting-mediated TBK1 activation, Optn provides a platform for TBK1 activation by autophosphorylation and that TBK1 is activated after the interaction of Optn with the autophagy machinery and ubiquitin and not before. They show that TBK1 phosphorylation is blocked by bafilomycine A1, an inhibitor of vacuolar ATPases that blocks the late phase of autophagy. Furthermore, they demonstrate that Optn is require for TBK1 phosphorylation since variation of Optn expression regulates TBK1 phosphorylation in response to PINK/Parkin-mediated autophagy. Interestingly, using immunofluorescence microscopy, they show that Optn forms sphere like structures at the surface of damage mitochondria which are more dispersed in the absence of TBK1. In addition, TBK1 is also recruited at the surface of damage mitochondria and as Optn and NDP52 (but not p62) colocalize with LC3B in response to PINK/Parkin-mediated mitophagy. Next, it is demonstrated that the Leucin zipper and LIR domains of Optn (which modulate Optn interaction with autophagosome) play an important role for TBK1 activation. Additionally, the autophagy core is shown to be required for TBK1 activation. Under basal conditions, depletion of the autophagosome machinery leads to an increase in autophagy receptors (except Optn) and TBK1 phosphorylation which colocalize with ubiquitin in insoluble moieties. In contrast, Optn remains cytosolic and is dispensable for TBK1 activation in these conditions. Then, using the fluoppi technic, the authors demonstrate that the generation of Optn-Ubiquitin condensates recruits and activates TBK1. They express in HCT116 TBK1-deficient cells engineered or pathological ALS mutations of TBK1 that affect ubiquitin interaction, structure, dimerization and kinase activity of TBK1. The expression level of TBK1 was only affected by the dimerization-deficient mutations. None of the mutations impaired Optn and TBK1 ubiquitination. Interestingly, some ALS-associated mutations affect TBK1 activity and it is said in the text that the dimerization-deficient mutations of TBK1 affect its activity proportionally to their level of expression, which is not really correct (the expression level of the mutants is very heterogenous and not always correlate to their activity). Regarding their effect on mitophagy, the authors claim that the phosphorylation of TBK1 correlate with mitophagy which is not really the case. By using TBK1 inhibitor or TBK1-depleted cells, the authors conclude that TBK1 is the only kinase phosphorylating Optn. However, BX-795 is not completely specific to TBK1. Finally, the authors use monobodies against Optn effective in inhibiting mitophagy in NDP52 KO cells. Some of the monobodies have been shown to form a ternary complex with Optn and TBK1, while others compete for the interaction between Optn and TBK1 which involves the amino-terminal region of Optn and the C-terminal region of TBK1. Monobodies that compete for the interaction of Optn with TBK1 could alter the cellular distribution of Optn and inactivate TBK1, but they do not alter the ubiquitination of Optn. Finally, these monobodies inhibit 50% of mitophagy. *
*Major and minor points: Introduction The first paragraph of the Introduction section is confused and difficult to read. First and second paragraphs (page 3 and top of page 4) are dedicated to macroautophagy processes but ended with one sentence on Parkin-mediated autophagy without further introduction, while all processes regarding mitophagy are detailed in the next paragraph. Links between ideas developed are also somewhat missing. For example, in page 6, the three last sequences detailed the phosphorylation of autophagosome component, the fact that Optn and TBK1 genes are involved in neurodegenerative diseases and autophosphorylation of TBK1 as a pre-requirement for TBK1 activation without evident links between them, except "interestingly". *
In response to the reviewer’s suggestion, we have rewritten the Introduction. The first paragraph focused on introducing the molecular mechanism underlying macroautophagy and the second paragraph focused on Parkin-mediated mitophagy. As the reviewer indicated, the ALS mutations and TBK1 phosphorylation during Parkin-mediated mitophagy are not well related, so we moved the background material on the relationship between OPTN and TBK1 in neurodegenerative diseases to the beginning of the section describing Figure 5. We believe these changes have made the Introduction easier to read and understand.
*Results *
*Major points: *
*1- Results are often over-interpreted regarding data obtained leading to inadequate conclusions (see below for details); *
We regret the reviewer’s concerns regarding over-interpretation. To address this issue, we have carefully considered the data, performed additional experiments where necessary, and rewritten the results accordingly. Please see our point-by-point responses below.
*2- Quantification of protein levels detected by western blot are provided as "relative intensities" without referring to specific loading control or to total protein when -phosphorylated forms are quantified (Fig. 1b, 1d, 1f, 1i, 2b, 2f, 2i, 5b, 7b, supplemental figures 2b). *
For the immunoblots, we loaded the same amount of total cell lysate and the phosphorylated forms were quantified relative to the total protein input. This has been mentioned in the Materials and Methods.
*3- In western blotting experiments, authors described slower migrating bands as "ubiquitinated" forms of detected proteins, but never provided experimental evidences that it could be the case. Use of non-specific deubiquitinase incubation of extracts prior to western blot could help to correctly identified ubiquitination versus other post-translational modifications such as phosphorylation, glycosylation, acetylation etc... *
We appreciate the reviewer’s suggestion. The cell lysates after mitophagy induction were incubated in vitro with a recombinant USP2 core domain (non-specific DUB), and then immunoblotted. As shown in supplemental Fig 1 of the revised manuscript, the slower migrating OPTN bands disappeared in a USP2-dependent manner. The slower migrating NDP52 and TOMM20 bands likewise disappeared. These results confirm that the slower migrating OPTN, NDP52, and TOMM20 bands are ubiquitinated.
*4- Conclusions from data obtained by immunofluorescent imaging are often drawn from only one image presented without further statistical analysis. *
Statistical significance was determined for the immunofluorescent data (original figures 1j, 2c and 3a). Please see Fig 1l, 2f, 2g, and 3a in the revised manuscript.
*Page 7: - authors referred to TBK1 phosphorylation induced by mitophagy induction as "TBK1 phosphorylation induced by Parkin-mediated ubiquitination" while mitophagy can be induced independently of Parkin (ex: via mitochondrial receptors) and without any evidence (according to referee's knowledge) of a link between ubiquitination by Parkin and TBK1 phosphorylation. *
As the reviewer indicated, Parkin-independent and ubiquitination-independent mitophagy pathways are also known (i.e. receptor-mediated mitophagy driven by NIX, BNIP3, BCL2L13, FKBP8, FUNDC1, or Atg32). Therefore, references to "mitophagy" in our manuscript were reworded as "Parkin-mediated mitophagy". Since TBK1 phosphorylation is observed before mitochondria are degraded and is dependent on Parkin-mediated ubiquitin (for example, see Fig 1c), we use the phrase "TBK1 phosphorylation triggered by Parkin-mediated OMM ubiquitination".
*Fig 1g: Western blots performed in Penta KO cells without exogene expression of any autophagy receptors should be provided as control. Furthermore, lower expression of NDP52 relative to that of Optn (using flag antibodies) should be discussed as it can explained the differential levels in TBK1 phosphorylation observed. *
As suggested, we repeated the experiment using Penta KO cells in the absence of exogeneous autophagy adaptor expression. Furthermore, we expressed different amounts of NDP52 and OPTN (indicated as low and high in the figure) in Penta KO cells to rule out the possibility that higher TBK1 phosphorylation is induced by simple overexpression of autophagy adaptor (please see Fig 1g and h in the revised manuscript). At high NDP52 expression (2.5-3.0-fold higher than endogenous NDP52), phosphorylated TBK1 was reduced to ~30% the level of that observed in WT cells after 3 hrs with val and baf. In contrast, Penta KO cells with higher OPTN expression (3.0-fold higher than endogenous OPTN) had phosphorylated TBK1 signals that were 2-fold higher than those in WT cells. Based on these results, we concluded that OPTN is an important adaptor for TBK1 activation during Parkin-mediated mitophagy.
*Page 8: Supplemental Fig 1a: - The inability of authors to observe TBK1 endogenous signal in HeLa cells using commercially available antibodies is surprising as many publications reported successful staining (see Figure 1 of Suzuki et al. 2013 Cell type-specific subcellular localization of phospho-TBK1 in response to cytoplasmic viral DNA. PLoS One. 8:e83639 among others) as well as commercial promotion (see Anti-NAK/TBK1 antibody from Abcam reference: ab235253). *
For the original manuscript, anti-TBK1 antibodies purchased from abcam (ab235253), CST (#3013S), Proteintech (28397-1-AP), and GeneTex (GTX12116) for immunostaining were unable to yield TBK1-positive signals (please see Fig 1 for reviewers below). WT and TBK1-/- HCT116 cells stably expressing Parkin were treated with valinomycin for 1 hr and immunostained with the indicated antibodies. Anti-phos-TBK1 antibody (CST, #5483) was used as a positive control. Based on these results, we stated in the original manuscript that the "endogenous TBK1 signal could not be observed using commercially available antibodies". At the reviewer’s suggestion, we purchased anti-TBK1 antibodies from abcam (ab40676) and CST (#38066). As shown in the figure below, the immunofluorescent signals generated by these antibodies were detected in WT, but not in TBK1-/- cells. The CST (#38066) antibody yielded a stronger signal, most of which was on damaged mitochondria. Thanks to this suggestion, we repeated the experiment using the new anti-TBK1 antibody. Furthermore, based on a suggestion from Reviewer 3, we detected mitochondrial recruitment of TBK1 during mitophagy stimulation (valinomycin for 30 min or 2 hrs in the presence and absence of bafilomycin; supplemental Fig 2 in the revised manuscript). We also detected association of endogenous TBK1 with ubiquitin-positive condensates in WT, FIP200KO, and ATG9A KO cells (Fig 3a and supplementary Fig 7a in the revised manuscript).
*- Conclusions of the localization of signal on mitochondria (dispersed, in the periphery or at contact sites) are clearly over-interpreted in the absence of other membrane or autophagosome specific labeling and statistical colocalization analyses of multiple images. It is particularly difficult to assess any difference between Tax1BP1, p62 and NBR1 localization on mitochondria subdomains. *
We previously expressed each FLAG-tagged autophagy adaptor in Penta KO cells and observed their localization during Parkin-mediated mitophagy and found that exogenous FLAG-tagged OPTN and NDP52, but not p62, colocalized with LC3B (Yamano et al 2020 JCB). No one has assessed and compared the localization of all five endogenous autophagy adaptors. Although we still believe that the results (supplemental Fig1 in the original manuscript) are informative for researchers in the autophagy field, we decided to remove that data from the revised manuscript since they are not the main focus of the study. We will consider publishing those data elsewhere in the future after co-staining with autophagosome markers and assessing the statistical significance of colocalization as the reviewer suggested.
*Page 9: *
*- First part of results ended without any conclusions. *
As detailed in the previous response, we have removed results for mitophagic recruitment of autophagy adaptors (supplementary Figure 1 in the original manuscript).
*- The observation that "TBK1 phosphorylation was not apparent in the Optn mutant cell lines, even after 3 hrs of valinomycin, ..." is inconsistent with detection of bands with anti-pS172-TBK1 antibodies in Fig 2a detected at 1hr (with F178A) and 3 hrs (4LA, F178A, and 4LA/F178A mutants) of treatment. *
We apologize for the confusion. This statement was clearly our mistake. We had intended to state when "all autophagy adaptors are deleted" no phosphorylated TBK1 was observed. We have rewritten this part as "TBK1 phosphorylation was not apparent in the Penta KO cells even after 3 hrs with valinomycin".
*- Similarly, decreased levels of phosphorylated TBK1 stated for F178A mutant was only observed at 1 but not 3hrs or at 3hrs in the presence of bafilomycin. *
Based on the mitophagy assay previously reported (Yamano et al 2020 JCB), the F178A mutant only moderately inhibited mitophagy (60% mitophagy with the F178A mutant vs 80% mitophagy with OPTN WT). Conversely, the 4LA mutant and 4LA/F178A double mutant had stronger inhibitory effects on mitophagy (35% for 4LA and 9% mitophagy for 4LA/F178A). Therefore, the levels of phos-TBK1 after 1 hr with valinomycin or 3 hrs with valinomycin in the presence of bafilomycin are consistent with mitophagy progression. When mitophagy proceeds efficiently, the amount of phos-TBK1 in the 1 hr val samples is reduced relative to the 3 hr val samples due to autophagic degradation.
To more clearly observe and compare the levels of mitophagy-dependent phos-TBK1 among Penta KO cells expressing OPTN WT and the mutants, we treated cells with valinomycin in the presence of bafilomycin for 0, 0.5, 1, and 2 hrs and quantified phos-TBK1. The results are shown in Fig 2c and d in the revised manuscript. The phos-TBK1 signal increased over time with val and baf treatment in all OPTN expressing cells. Cells with OPTN WT generated the most phos-TBK1, whereas the signal generated by the F178A mutant was 75% that of the OPTN WT-expressing cells and the 4LA and 4LA/F178A mutants were about 40%. The experiments were independently replicated three times and error bars and statistical significance were incorporated into the associated graph. These results indicate that OPTN association with the autophagy machinery, in particular ATG9A vesicles, is important for TBK1 activation.
*Page 10: *
*The results and their repartition between figure 2 d, e, f, g, h, I and figure 3 is a bit confusing. In these experiments, it is shown Figure 2 that the absence or depletion of the autophagy machinery increase the phosphorylation of TBK1 and in Figure 3 it is shown that not only the phosphorylation of TBK1 accumulate but also the expression of NDP52, Tax1BP1 and p62. Is it because their degradation by autophagy is blocked (like for phosphoTBK1)? *
The reviewer is correct that autophagy adaptors other than OPTN (especially TAX1BP1, p62 and NBR1) are constantly degraded by macro/micro autophagy (Mejlvang et al. 2018 J Cell Biol and Yamano et al. 2021 BBA Gen Subj). Therefore, these adaptors accumulate in autophagy deficient cell lines (original Fig 3). In this study, we found that in the absence of mitophagy stimulation phos-TBK1 accumulates in autophagy deficient cell lines. This suggests that the accumulated autophagy adaptors induce TBK1 phosphorylation under basal conditions. In the original manuscript, we claimed that TBK1 phosphorylation under basal conditions does not require OPTN since in FIP200 KO and ATG9A KO cells it did not accumulate and did not primarily colocalize with ubiquitin- and TBK1-positive foci (original Fig 3). To gain more direct evidence for the revised manuscript, we performed additional experiments and discovered that TAX1BP1 is the adaptor responsible for TBK1 autophosphorylation under basal autophagy. We treated FIP200KO and ATG9A KO cells with siRNAs against OPTN, NDP52, TAX1BP, p62, and NBR1, and immunoblotted total cell lysates with an anti-phos-TBK antibody. As shown in Fig 3f in the revised manuscript, TAX1BP1 siRNA treatment decreased phos-TBK1 levels without affecting total TBK1. This result indicates that the accumulation of TAX1BP1 in the FIP200 KO and ATG9A KO cells induced TBK1 autophosphorylation under basal conditions. Considering this result, we treated WT, FIP200 KO, and ATG9A KO cells with TAX1BP1 siRNA, and then induced Parkin-mediated mitophagy with valinomycin in the presence of bafilomycin. This strategy eliminated the basal accumulation of phos-TBK1 and allowed us to focus on mitophagy-dependent TBK1 phosphorylation. Please see revised Fig 2h, I, j, and k. The results showed that mitophagy-dependent phos-TBK1 is predominantly attenuated in FIP200 KO and ATG9A KO cells. In Figs 2 and 3, we would like to emphasize that OPTN is required for TBK1 phosphorylation in response to Parkin-mediated mitophagy, whereas TAX1BP1 is required for TBK1 phosphorylation in basal autophagy. Since Reviewer 3 commented that interpretation of the data in original Figs 2d, e, and f was challenging, we elected to move those results to the supplemental figures. We have incorporated the newly acquired data (mitophagy using FIP200 KO or ATG9A KO with TAX1BP1 siRNA cells) into the main figure. We believe that this makes the text easier for readers to understand.
*- Fig 2c: conclusions on *
*the reduction of recruitment of Optn mutants on autophagosome formation seem over-interpreted as: *
*1- no labeling with LC3 has been used to identified autophagsome, *
*2- immunofluorescent signals observed with mutants are dispersed throughout the entire mitochondria network (see the merged images) rendering impossible to distinguish between autophagosome-associated mitochondria and others. *
*The following conclusive sentence stating that association of Optn to damaged mitochondria is not sufficient for TBK1 activation based solely on IF of figure 2c seems therefore unrelated to the obtained data. *
To address concerns about the recruitment of OPTN mutants to the autophagosome formation site, we performed additional experiments. Penta KO cells and those expressing OPTN WT and mutants were treated with valinomycin for 1 hr, and FLAG-tagged OPTN, endogenous WIPI2, and HAP60 (mitochondrial marker) were detected by immunostaining. We detected endogenous WIPI2 because WIPI2 is recruited only to autophagosome formation sites (Dooley et al. 2014 Mol Cell), whereas LC3B assembles on autophagosome formation sites and is also associated with completed autophagosomes. Confocal microscopy images showed that cup-shaped OPTN WT that had been recruited to damaged mitochondria colocalized with WIPI2. Quantification further showed that during mitophagy the number of WIPI2 foci seen in cells expressing OPTN WT decreased in Penta KO cells and cells expressing OPTN mutants (4LA, F178A and 4LA/F178A). These data are shown in Fig 2e and f in the revised manuscript. In addition, we quantified the number of cells that either exhibited heterogeneous or homogeneous recruitment of OPTN to damaged mitochondria after treatment with valinomycin for 1 hr. More than 80% of Penta KO cells with OPTN WT had heterogeneous OPTN recruitment, whereas this distribution was only present in 10% of cells expressing either OPTN 4LA or OPTN 4LA/F178A. Although cells expressing the OPTN F178A mutant exhibited 50% heterogeneous recruitment, this may be because the mutant can interact with ATG9A. As mentioned above, our previous mitophagy analyses (Keima-based FACS analysis, Yamano et al 2020 JCB) showed that the OPTN F178A mutant induced ~60% mitochondrial degradation (which is correlated strongly with OPTN distribution), whereas it was 80% with OPTN WT and 9% with 4LA/F178A.
*- Fig 2d: authors should explain why ATG KO cells displayed lipidated LC3B in the absence of efficient autophagy processes. *
We thank the reviewer for the suggestion. We added the following sentence to explain the function of ATG5 in LC3B lipidation. "Since LC3B lipidation is catalyzed by ATG5, but not FIP200 and ATG9A, the lipidated form disappears only in ATG5 KO cells (Hanada et al 2007 J Biol Chem). "
*- Fig 2e: despite authors statement that TBK1 phosphorylation did not increase during mitophagy in ATG KO cells, increased pS172-TBK1 is visible in FIP200 and ATG5 KO cells especially between 1 and 3 hrs of stimulation, leading to inaccurate conclusions that TBK1 phosphorylation requires the autophagy machinery. Therefore, overall assumption that both ubiquitination and autophagy subunits are required for TBK1 autophosphorylation appears erroneous. *
As the reviewer indicated, phos-TBK1 levels gradually increased in ATG KO cells. The main text was rewritten to more accurately reflect this increase. Based on experiments using the monobodies and those conducted during the revision process, it is apparent that although the autophagy machinery may not be completely essential for TBK1 phosphorylation, it clearly facilitates TBK1 phosphorylation in response to Parkin-mediated mitophagy.
*Page 12: *
*- Fig 3a: conclusion that Optn signal is more cytosolic and did not localize with Ub condensates seems speculative as based on: *
*1- only one immunofluorescence image without statistical analysis *
*2- Optn and Ub signals are lower in images with Optn is analyzed compared to other images in which NDP52, TAX1BP1 and NBR1 are detected. *
To address these concerns, we compared and quantified the signal intensities of all endogenous autophagy adaptors in FIP200 KO and ATG9A KO cells. The quantification data are shown in Fig 3a and the immunofluorescence images are shown in supplementary Fig 6a of the revised manuscript.
*- Fig 3b: interpretation of western blot data is uncertain due to lack of appropriate loading control, especially with pellets (P) extracts. In addition, it is not clear how to conclude from the experiments in Fig 3b that autophagy adaptors other than Optn mediate TBK1 phosphorylation. *
When autophagy is inhibited, p62 accumulates in the cytosol as aggregates (Komatsu et al. 2007 Cell). Therefore, p62 should be a positive control. Indeed, Fig 3b in the original manuscript (Fig 3b and c in the revised manuscript) showed that the amount of p62 in the pellet fraction was elevated in FIP200 KO and ATG9A KO cells. Furthermore, these aggregates were also observed in the imaging data (Fig 3a and supplementary Fig 7 in the revised manuscript). As the reviewer indicated, the original manuscript did not clarify whether autophagy adaptors other than OPTN mediated TBK1 phosphorylation; however, our revised results clearly demonstrate that TAX1BP1 is the adaptor responsible inducing TBK1 autophosphorylation when basal autophagy is impaired (please see Fig 3f in the revised manuscript).
*Minor point: reference is missing in the last sentence of the paragraph stating that K48-linked chains dominate when autophagy pathways are impaired. *
While several autophagy adaptors preferentially interact with K48-linked ubiquitin chains (Donaldson et al. 2003 PNAS etc), TRAF6 is recruited to ubiquitin-condensates via p62-mediated K63-linked ubiquitination (Linares et al. 2013 Mol Cell). Furthermore, K33-linked ubiquitin chains are also present in p62-positive condensates (Nibe et al. 2018 Autophagy). Because it’s not clear which ubiquitin-linkage is dominant in the condensates, we decided to delete the sentence. We regret the confusion.
*Page 13: *
*Conversely to Optn, they find that the other autophagic receptors localize in insoluble fractions (what does it mean?) independently of TBK1 expression (experiments with DKO cells) and also independently of Optn (where is this shown?). Altogether, these experiments are far from the message of the manuscript. The title of the paragraph "TBK1 activation does not require Optn under basal autophagy conditions" is not correct because even if the level of expression of autophagic receptors and TBK1 phosphorylation are increase in response to the depletion of the autophagy machinery, it does not increase autophagy. *
According to the suggestion, we changed the title of the paragraph to "TAX1BP1, but not OPTN, mediates TBK1 phosphorylation when basal autophagy is impaired." In addition, we rewrote this section.
*- Fig 3d: authors should mention the nature of the upper band observed in Optn western blot and show the same experiment in since solely TBK1 depleted cells since they stated that "electrophoretic migration of Optn was not affected by TBK1 deletion". In addition, suggesting from these sole experiments that "NP52, TAX1BP1, p62, NBR1 and AZI2 form Ub-positive condensates where TBK1 is activated" seems over-interpretated. *
Reviewer 3 suggested we characterize the upper band in the OPTN blot (Fig 3d in the original manuscript). To determine if the band is genuine OPTN, we used phostag-PAGE to analyze cell lysates from cells treated with control siRNA or OPTN siRNA and found that both the lower and upper bands were OPTN species (please see "Figure 2 for reviewers" in our response to Reviewer 3). The same pattern was reported by the Wade Harper group (Heo et al. 2015 Mol Cell). They showed that the OPTN double band pattern on phos-tag PAGE was not affected by TBK1 deletion. We have cited this literature where appropriate in the revised manuscript. In WT cells, it is difficult to detect phosphorylation of autophagy adaptors by TBK1 because basal autophagy constantly degrades them. That’s why we used autophagy KO cell lines.
*Page 14: *
*- Fig 4: TBK1 phosphorylation was analyzed in Fig4d and not in Fig4b as stated. In addition, it is rather difficult to conclude from artificial multimerization experiments, as the authors have done, that interaction between Optn and autophagy components contributes to Optn multimerization in genuine conditions. *
Detection of phos-TBK1 has been corrected to Fig 4b. Although artificial, the fluoppi assay provides insights into how OPTN activates TBK1 and how the autophagy machinery contributes to TBK1 activation via OPTN. To determine if artificial OPTN multimerization could bypass the autophagy machinery requirement, we used the fluoppi assay. This assay was important for us to conclude that the autophagy machinery and Parkin-mediated ubiquitination allow OPTN to be assembled in close proximity to where TBK1 is activated. The main text was rewritten to better convey the benefits of the fluoppi assay.
*Page 15: *
*This work could have therapeutic consequences but the pathological mutants of TBK1 used affect ALS (Figure 5) while in the discussion it is proposed that monobodies could have a therapeutic interest in familial forms of glaucoma due to the E50K mutation of Optn. It should be better to target only one pathology. *
Both TBK1 and OPTN are causative genes for ALS and many pathogenic mutations are known to impact their function. In this study, we focused on ALS mutations in TBK1 that affect self-dimerization and investigated their impact in response to Parkin-mediated mitophagy. We created the monobodies as a tool to physically inhibit OPTN assembly at the contact site. Although our monobodies inhibit Parkin-mediated mitophagy, they would not be a useful therapeutic strategy for ALS due to the loss of function with the TBK1 mutations. However, because TBK1 E50K is a glaucomatous mutation that causes OPTN-TBK1 to bind more tightly, our monobodies might be applicable to glaucomatous pathology since they could disrupt this interaction. We thus feel that it is appropriate to mention the potential of the monobodies and their future utility in the Discussion.
*- Fig 5c, d: Authors stated that degree of TBK1 autophosphorylation correlated with OPTN phosphorylation at S177 whereas phosphorylated TBK1 is unaffected by L693Q and V700Q mutants that display decreased phosphorylated Optn In addition, authors interpretation of Figure 5 data is clearly problematic as they stated that: *
*1- neither 693Q and V700Q mutants had "significant effect on mitophagy", while decreasing efficiency from 78% to 37-51% *
*2- but conclude that 49.7% mitophagy levels of R357Q mutant is significant mitochondrial degradation. *
*Overall conclusion that mitophagy efficiency is correlated with phosphorylated TBK1 levels is therefore inaccurate. *
We regret that this section did not sufficiently describe the data. Reviewer 3 also noted that the text referencing Fig 5 was difficult to interpret. One of the reasons for the complicated data interpretation is the number of TBK1 mutants used. The L693Q and V700Q mutations used by Li et al. (2016 Nat Commun) were expected to inhibit Parkin-mediated mitophagy since those authors reported that the mutations prevented interactions with OPTN. However, our in-cell assay showed that the two mutations only moderately affected Parkin-mediated mitophagy. Furthermore, both the L693Q and V700Q mutations were engineered based on the X-ray structure, rather than being authentic pathogenic ALS mutations. To avoid any potential confusion, we decided to remove the L693Q and V700A data. We have re-evaluated the other data and have rewritten this section accordingly. Please see the revised main text.
*Discussion *
*Minor points: *
*page 20: - reference is missing in the sentence "Optn cannot oligomerize on its own on ubiquitin-decorated mitochondria". *
We have provided the appropriate reference.
*Major points: *
*Authors stated that they showed that Optn recruitment to damaged mitochondria, itself, is insufficient for TBK1 autophosphorylation, but did not show experiment of Optn recruitment to mitochondria and its consequences on TBK1 phosphorylation in the absence of mitophagy induction signal. Authors could for example target HA-Ash-6Ub to mitochondria in order to artificially recruit hAG-Optn to "ubiquitinated" mitochondria in the absence of mitophagy signal. *
We showed that the efficiency of TBK1 autophosphorylation was reduced in cells expressing the OPTN 4LA/F178A mutant, which cannot interact with the autophagy machinery (Fig 2c and d in the revised manuscript). Cells with FIP200 or ATG9A knockdown also have reduced phos-TBK1 (pS172) as shown in supplementary Fig 5e and f. The rate of phos-TBK1 (pS172) generation in ATG9AKO cells during Parkin-mediated mitophagy is reduced relative to that in WT cells (Fig 2j and k). Since a small amount of phos-TBK1 was generated in both ATG9A knockdown and KO cells (supplementary Fig 5e, f, Fig 2j and k), we concur that it would be premature to conclude that phosphorylation of TBK1 does not occur at all when autophagy core components are absent. A small amount of phos-TBK1 may be generated by OPTN that is freely distributed on the outer mitochondrial membrane. In the revised manuscript, we mention the possibility that TBK1 might be phosphorylated by OPTN independent of the autophagy machinery and were careful to avoid over-interpretation.
As shown in Fig 4, fusing OPTN with an Azami-Green tag can induce artificial multimerization and trigger the generation of phos-TBK1 (pS172). Therefore, we expect that mitochondria-targeted HA-Ash-6Ub would induce TBK1 phosphorylation in a hAG-OPTN-dependent manner as was observed with cytosolic HA-Ash-6Ub (Fig 4). The accumulation of OPTN at the contact site in Parkin-mediated mitophagy is important for TBK1 phosphorylation. Even if OPTN is forced to anchor to the mitochondria, this would induce isolation membrane formation and subsequent autophosphorylation of TBK1. Therefore, the utility of forcing OPTN to anchor to mitochondria is questionable.
*Similarly, experimental approaches used by authors lack dynamics parameters to conclude on formation and elongation of isolation membranes and contacts sites that could be probably obtained through video microscopy. *
Based on the reviewer’s comment, we performed time-lapse microscopy to observe OPTN recruitment to the contact site and followed its movement along with the elongation of isolation membranes during Parkin-mediated mitophagy. HeLa cells stably expressing GFP-OPTN and pSu9-mCherry (a mitochondrial marker) were treated with valinomycin (please see Fig 2l in the revised manuscript). Initial recruitment of GFP-OPTN near mitochondria was evident as small dot-like structures that then elongated over time to become cup-shaped structures and culminated in the formation of spherical structures. Considering the colocalization of OPTN with WIPI1/WIPI2 (markers of autophagosome formation site) in Fig 2e and supplementary Fig 2a, the time-lapse images strongly suggest that OPTN assembles at contact sites followed by elongation in tandem with isolation membranes during Parkin-mediated mitophagy.
*Finally, the model proposed by the authors does not take into account data showing that Optn basally interacts with ubiquitinated mitochondria and LC3 family members (see Wild et al., Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science. 2011 333:228-33), although at lower levels compared to induced conditions, relativizing the impact of the proposed model. *
According to the Reviewer 2 comment, we again read the Science paper (Wild et al. 2011) but could not find data showing that OPTN basally interacts with ubiquitinated mitochondria. At least, we think that under steady state conditions without mitophagy induction, mitochondrial ubiquitination and mitochondrial localization of OPTN are undetectable as shown in supplementary Figure 2 in our revised manuscript.
*In conclusion, this manuscript represents a lot of work but the experiments often lack controls and are over-interpretated. *
***Referees cross-commenting** *
*In my opinion, what emerges from these 3 reviews is that the results lack controls or have not been repeated enough to support the message that the interaction of Optn with ubiquitin and the ubiquitination machinery is sufficient to activate TBK1. In particular, as reviewer 1 says, the phosphorylation kinetics shown in Figure 1a are not consistent with TBK1 phosphorylation following the interaction of Optn with the ubiquitination machinery and ubiquitin. In Figure 1e, there is a decrease in TBK1 phosphorylation in contrast to WTcells as mentioned by Reviewer 1. In agreement with Reviewer 1, we believe that the WT cells are missing in Figure 1g. *
*With regard to Figure 2c, we agree with reviewer 1 that an LC3 label is missing in order to be able to interpret the data. In Figure 2e and f, we agree with reviewer 1 that it is difficult to understand why TBK1 phosphorylation increases in the absence of the autophagy machinery (FIP200 KO and ATG5KO). In Figure 3, loading controls are missing for 3b and c. The TBK1 KO cells alone are missing in Fig 2d. In Figure 2b, pTBK1 is missing. In agreement with reviewer 3, we believe that the data with fluoppi contradict the message of the manuscript since they show that TBK1 can be phosphorylated by ubiquitin in the absence of the ubiquitination machinery. In agreement with reviewer 3, we believe that the experiments in Figure 5 are very difficult to interpret. The first reviewer is right to ask the question of the replicates for figures 6c and d. *
We appreciate the summary of the reviewers’ comments. To address their concerns, we have included the appropriate controls and included the results of three independent experiments in the graphs, which now include appropriate error bars and statistical significance. Thus, we believe we have answered the most critical comments concerning the lack of controls.
In Fig 1a, phos-TBK1 was maximal following 30 min of valinomycin treatment. We confirmed using microscopy-based observations that recruitment of endogenous TBK1 and OPTN and the generation of phos-TBK1 and phos-OPTN at contact sites (marked by WIPI1) near damaged mitochondria was also maximal after 30 min of valinomycin treatment (supplementary Fig 2 and 3). Therefore, the kinetics of phos-TBK1 and phos-OPTN generation are consistent with the recruitment of OPTN-TBK1 to the contact site.
The data presented in Fig 2 clearly indicate that the autophagy components are involved in phos-TBK1 generation during Parkin-mediated mitophagy. Therefore, the claim that ubiquitination machinery is sufficient for TBK1 activation is incorrect. Although we agree that the autophagy gene deletions cannot completely inhibit TBK1 autophosphorylation, mitophagy-dependent generation of phos-TBK1 is largely impaired by ATG9A KO (Fig 2j and k). Thus, there is no doubt that isolation membrane formation is important for TBK1 activation following Parkin-mediated mitophagy.
Fig 1e - The reviewer is correct that phos-TBK1 is reduced in the NDP52 knockout. We have rewritten the main text. It is also true that NDP52 has a smaller effect on TBK1 autophosphorylation as compared to OPTN.
Fig 1g - Immunoblots using total cell lysates prepared from six different cell lines (WT, Penta KO alone, Penta KO stably expressing low or high OPTN or NDP52) under four different conditions (DMSO, valinomycin 1 hr, valinomycin 3 hrs, valinomycin + bafilomycin 3 hrs) showed that OPTN is a rate-limiting factor for TBK1 phosphorylation. Please see Fig 1g and h in the revised manuscript
Fig 2c - The recruitment of OPTN WT and associated mutants to the contact site was re-examined by immunostaining with WIPI2 labeling. We found that OPTN WT was both efficiently recruited to and formed the contact site. In contrast, the OPTN 4LA/F178A mutant was unable to interact with FIP200/LC3/ATG9A and was uniformly (i.e. homogenously) distributed on damaged mitochondria with the rate of autophagosome site formation reduced. Please see Fig 2e, f, g in the revised manuscript.
Fig 2e and f - KO of the autophagy core components FIP200 and ATG9A increased phos-TBK1 under basal, non-mitophagy-associated conditions (see Fig 3). The levels of autophagy adaptors other than OPTN also increased in FIP200 KO and ATG9A KO cells. Furthermore, as shown in Fig 3a and supplementary Fig 7, both phos-TBK1 and the autophagy adaptors accumulated in Ub-positive condensates. Based on previous reports (Mejlvang 2018 J Cell Biol), TAX1BP1, p62, and NBR1 have short half-lives and are quicky degraded by macro/micro autophagy. The accumulation of phos-TBK1 in the absence of autophagy occurs because autophagy-dependent degradation of TAX1BP1 (and other adaptors) is inhibited. This allows for the formation of Ub-positive condensates, which brings TBK1 into sufficient proximity for activation. This has been noted in the revised manuscript.
Fig 3b and 3c - We wonder if the "loading controls are missing for Fig 3b and 3c" statement might be a misinterpretation by the reviewer as TOMM20 was used as the loading control in the original Fig 3b. It was recovered in the supernatant fractions of WT, FIP200 KO, and ATG9A KO cells, indicating that the accumulation of autophagy adaptors in the pellet fractions depends on autophagy gene deletion. Similarly, actin and TOMM20 were used as loading controls in the original manuscript Fig 3c.
Fig 2d (perhaps meant to be Fig 3d) – A previous study reported that phos-tag PAGE blot of OPTN in TBK1 KO cells alone revealed no differences between WT and TBK1 KO cells (Heo et al 2015 Mol Cell). We cited this reference in the revised manuscript.
Fig 2b (perhaps meant to be Fig 4b) - Immunoblots of phos-TBK1 have been incorporated into the results of Fig 4b in the revised manuscript.
Fig 4 - We show in Fig 2 that induction of Parkin-mediated mitophagy promotes OPTN accumulation at contact sites formed by isolation membranes and ubiquitinated mitochondria, and that autophagy core subunits are required for efficient generation of phos-TBK1. Fig 3 shows that phos-TBK1 accumulates in Ub-positive condensates with TAX1BP1, rather than OPTN, and that it is responsible for phos-TBK1 accumulation. Together, these results suggest a model in which TBK1 is activated when OPTN and TBK1 are positioned near each other. We hypothesized that if we could force OPTNs into close proximity the autophagy machinery requirement for TBK1 activation might be bypassed. To assess this model, we designed the fluoppi assay shown in Fig 4. This assay was critical in that it provided an important clue for the molecular mechanism that OPTN and the autophagy machinery use to cooperatively induce TBK1 trans-autophosphorylation. Because the original manuscript may not have sufficiently conveyed our reasoning for the fluoppi analysis, we have rewritten this section. The main point of the fluoppi assay is that engineered OPTN multimerization was able to bypass the autophagy requirement for TBK1 activation.
Fig 5 - For easier interpretation, the L693Q and V700Q data, which are not related to ALS pathology, have been removed.
Fig 5d – Statistical significance has been determined for the mitophagy results and the main text has been rewritten for better clarity.
Fig 6c, d, and I – The experiments were independently replicated more than three times with statistical support and error bars incorporated into the associated graphs.
*Reviewer #2 (Significance (Required)): *
*this manuscript represents a lot of work but the experiments often lack controls and are over-interpretated. The manuscript is for a broad audience. *
For the revised manuscript, additional experiments were carefully performed with appropriate controls and the manuscript was rewritten to address concerns regarding over-interpretation. We hope that we have adequately addressed the reviewer’s comments.
*Reviewer #3 (Evidence, reproducibility and clarity (Required)): *
*The authors investigated the mechanisms by which TBK1 is phosphorylated and thus activated in PINK1/Parkin-mediated mitophagy. They show data indicating that OPTN, by interacting both with ubiquitin-coated mitochondria and with the autophagy machinery, provides a platform where OPTN-bound TBK1 can be hetero-autophosphorylated by adjacent TBK1. *
*According to the prevailing model (prior to this manuscript), TBK1 activation via autophosphorylation leads to TBK1-mediated phosphorylation of OPTN S177 and subsequent pOPTN-mediated recruitment of autophagic isolation membranes to the mitochondria. However, based on the model provided in this manuscript, OPTN needs to interact first with both autophagic membranes and ubiquitin before TBK1 can become activated. *
*This is an important topic. Overall, the experimental data are of high scientific quality. For the most part, the manuscript is clearly written. The figures have been made with great care. The novel insights are relevant. However, a number of issues need to be addressed or clarified. *
*Major comments: *
Furthermore, we show that Parkin expression levels affect the amount of phos-TBK1 generated during mitophagy. Please see supplementary Fig 4 in the revised manuscript. When PARKIN was integrated into HeLa cells under a CMV promoter via an AAVS1 (Adeno-associated virus integration site 1)-locus, the resultant cell line (referred to as high-Parkin) had higher Parkin levels than HeLa cells in which PARKIN was introduced by retrovirus infection (referred to as low-Parkin). In high-Parkin HeLa cells, phos-TBK1 levels reached a maximum after 30 min with valinomycin, while in low-Parkin HeLa cells, phos-TBK1 levels were comparable after 30 min and 1 hr. High-Parkin HeLa was used for Fig 1a, b, c, and d as well as supplementary Fig 1, 2, 3 and 4. For all other Figs, PARKIN genes were introduced by retrovirus infection. This is one of the reasons why val was used for 30 min in Fig1, but 1-3 hrs for the other Figs. Because 3 hrs valinomycin treatment may be unsuitable for evaluating OPTN recruitment to mitochondria/isolation membrane contact sites, we deleted the original Fig 2c and replaced it with the val 1 hr data (Please see Fig 2e in the revised manuscript).
Figure 2 demonstrates that both ubiquitin on mitochondria and formation of the isolation membranes are needed to provide a platform for OPTN to assemble in close proximity to each other and subsequently induce TBK1 autophosphorylation. To determine if OPTN proximity is sufficient for TBK1 autophosphorylation (i.e., if engineered OPTN multimerization can bypass the autophagy machinery requirement for TBK1 autophosphorylation), we used the fluoppi assay. The results clearly showed that engineered OPTN multimerization induced TBK1 autophosphorylation without the need for the autophagy machinery. Although this is not a mitophagy experiment, the fluoppi assay provided crucial insights into the molecular mechanism underlying OPTN-mediated TBK1 autophosphorylation. The main text was rewritten to provide more clarity regarding the purpose of the fluoppi experiments.
The amount of phos-TBK1 during Parkin-mediated mitophagy was reduced in cells with the OPTN 4LA/F178A mutant, which cannot interact with the autophagy machinery (e.g. FIP200, ATG9A, and LC3) but can be targeted to mitochondria (see Fig 2c, d). ATG9AKO cells also had reduced amounts of phos-TBK1 relative to WT cells (See Fig 2j, k). Therefore, rather than OPTN-ubiquitin freely diffusing laterally on the outer membrane, we suggest that the contact site OPTN forms with ubiquitin and the autophagy machinery provides a more suitable platform for TBK1 autophosphorylation because it maintains TBK1 in a proximal position for a longer period of time.
The OPTN UBAN domain binds a ubiquitin-chain composed of two ubiquitin molecules (Oikawa et al. 2016 Nat Comm), and during Parkin-mediated mitophagy only shorter length poly-ubiquitin chains are generated on the mitochondrial surface (Swatek et al. 2019 Nature). Based on those findings, it is unlikely that multiple OPTN bind to one ubiquitin chain. Of course, we cannot rule out the possibility that TBK1 autophosphorylation does not occur on mitochondria in the absence of autophagy components. While full activation of TBK1 requires OPTN to associate with the isolation membrane, initial TBK autophosphorylation at the onset of mitophagy may occur based only on the OPTN-ubiquitin interaction. These explanations have been added to the Discussion in the revised manuscript.
Furthermore, based on comments from Reviewer 2, we performed time-lapse microscopy to observe OPTN dynamics during Parkin-mediated mitophagy (please see Fig 2l). HeLa cells stably expressing GFP-OPTN and pSu9-mCherry (a mitochondrial marker) were treated with valinomycin. GFP-OPTN was initially a peri- mitochondrial dot-like structure that elongated over time to a cup-shaped structure and which eventually ended up forming a spherical structure. The time-laps imaging showed that, at least in WT cells, OPTN is directly recruited to the contact sites and elongates along with the isolation membranes. We thus concluded that TBK1 is activated (autophosphorylated) at the contact site rather than on the outer membrane where OPTN-TBK can move freely.
We regret the confusion. Reviewer 2 also noted that the main text for Fig 5 was difficult to interpret. One of the reasons that complicated interpretation of the data is the number of TBK1 mutants used. The L693Q and V700Q mutations used by Li et al. (2016 Nat Commun) were expected to inhibit mitophagy since those authors reported that the mutations prevented interactions with OPTN. However, our in-cell assay showed that the two mutants only moderately affected Parkin-mediated mitophagy. Furthermore, both L693Q and V700Q were engineered based on the X-ray structure and are not ALS pathogenic mutations. To simplify the data and to make data interpretation easier, we decided to delete the L693Q and V700A data. We also determined statistical significance and rewrote this section.
We apologize for the lack of statistical analyses. We repeated experiments (if the experiments had not been independently performed more than three times) with statistical significance and error bars incorporated into the relevant figures.
*Other comments: *
Fig. 1a: how do they know that the upper OPTN band is ubiquitinated OPTN? Reviewer 2 raised the same question. To demonstrate that the upper OPTN band is ubiquitinated, cell lysates after mitophagy induction were incubated in vitro* with a recombinant USP2 core domain, and the samples immunoblotted. As shown in supplementary Fig 1 in the revised manuscript, the upper OPTN band disappeared in a USP2-dependent manner. The upper NDP52 and TOMM20 bands similarly disappeared. Therefore, the upper OPTN, NDP52 and TOMM20 bands observed after mitophagy induction are ubiquitinated.
Fig. 1a,b: the bafilomycin stabilization of pTBK1, OPTN and pOPTN indicates that these proteins are substantially degraded by autophagy within 30-60 minutes. This seems extremely fast for mitophagy completion. Please discuss.*
According to Kulak et al. (2014 Nat Methods), autophagy adaptor abundance (OPTN: 2.32E+4 and NDP52: 3.34E+4 in HeLa cell line) is low compared to that of mitochondria (TOMM20: 1.45E+6 in HeLa cell line). This is one of the reasons why autophagic degradation of adaptors is easier to see. Degradation of phos-TBK1 was likewise easy to detect, whereas total TBK1 was not. This discrepancy is likely based on differences in the abundance of phos-TBK1 and total TBK1. In addition, because autophagy adaptors are localized outside of the mitochondrial membrane they may be easier targets for lysosomal degradation than matrix proteins, which are localized inside the outer and inner membranes.
Both mass spectrometry and mutational analyses indicated that OPTN S473 is phosphorylated during Parkin-mediated mitophagy and that OPTN phosphorylated at S473 strongly binds ubiquitin chains (Richter et al. 2016 PNAS and Heo et al. 2015 Mol Cell). However, because a phos-S473 OPTN antibody is, to the best of our knowledge, currently not commercially available, we did not focus on S473 phosphorylation.
We regret the over-interpretation. As the reviewer indicated, the amount of phos-TBK generated in response to mitophagy was reduced in NDP52 KO cells relative to that in WT cells. This has been corrected. We would like to emphasize that, unlike OPTNdeletion, NDP52 deletion has relatively minor effects on TBK1 phosphorylation.
We apologize for this error. We intended to state “TBK1 phosphorylation was not apparent in the Penta KO cells without OPTN expression even after 3 hrs with valinomycin, indicating that autophagy adaptors are essential for TBK1 activation”. This sentence has been corrected in the revised manuscript.
The reviewer is correct, we have corrected the error.
The newly generated pTBK1 levels following Parkin-mediated mitophagy were calculated as pTBK1 [val & baf 3 hrs] minus pTBK1 [DMSO]. Since pTBK1 [val & baf 3 hrs] in WT cells is set to 100%, the newly generated pTBK1 in WT cells was 100% - 5% = 95%. The calculated values for pTBK1 [DMSO] and pTBK1 [val & baf 3 hrs] in ATG9A KO cells were ~55% and ~85%, respectively. Consequently, newly generated pTBK1 in the ATG9A KO cells is ~85% - ~55% = 30%. For clarity, we modified the figure to make the meaning of the numbers more apparent.
According to Kishi-Itakura et al. (2014 J Cell Sci), ferritin accumulates in the p62 condensates in FIP200 KO and ATG9A KO cells. However, it is unknown if the ferritin in the condensates is ubiquitinated. In the original manuscript, we confirmed by immunostaining that the p62-NBR1 condensates contain ferritin (Fig 3a in the original manuscript and supplementary Fig 7b in the revised manuscript).
We thank the reviewer for the suggestion. As mentioned, ferritin is a known substrate that accumulates in p62 condensates, we thus sought to confirm its presence. We have included this explanation in the revised manuscript.
We thank the reviewer for the suggestion. We obtained a rabbit anti-NBR1 antibody that allowed us to co-immunostain with the mouse anti-ubiquitin antibody (please see supplementary Fig 7b in the revised manuscript).
To determine if the two bands are genuine OPTN, total cell lysates prepared from HeLa cells treated with control siRNA or OPTN siRNA were subjected to phos-tag PAGE followed by immunoblotting with an anti-OPTN antibody. As shown below (Figure 2 for reviewers), the two bands (indicated as blue arrowheads) were absent in the OPTN knock down cells, indicating that both are derived from OPTN. Since phosphorylated species migrate slower in phos-tag PAGE, the upper band might be a phosphorylated form. The specific Ser/Thr phosphorylated in OPTN, however, remains to be determined. Heo et al. (2015 Mol Cell) also reported the two OPTN bands on phos-tag PAGE and that both were unchanged in TBK1 KO cells, suggesting that at least the upper band is not affected by TBK1.
We immunoblotted phos-TBK1. Please see Fig 4b in the revised manuscript.
*Reviewer #3 (Significance (Required)): *
*The novel insights are relevant. *
*According to the prevailing model (prior to this manuscript), TBK1 activation via autophosphorylation leads to TBK1-mediated phosphorylation of OPTN S177 and subsequent pOPTN-mediated recruitment of autophagic isolation membranes to the mitochondria. However, based on the model provided in this manuscript, OPTN needs to interact first with both autophagic membranes and ubiquitin before TBK1 can become activated. *
Based on our time-lapse microscopy observations (Fig 2l), OPTN recruited to the vicinity of mitochondria was visible as a small dot-like structures that likely correspond to contact sites between mitochondria and the isolation membrane since OPTN colocalizes with WIPI1 (please see supplementary Fig 2). These results support our proposed model that OPTN interacts with both isolation membranes and ubiquitin at the onset of mitophagy. Without TBK1 activation, OPTN can interact with ATG9A vesicles, a seed for isolation membrane formation (Yamano et al 2020 JCB), and TBK1 can interact with the PI3K complex (Nguyen et al 2023 Mol Cell). Therefore, OPTN-TBK1 can be recruited to the contact site from the very beginning of mitophagy induction prior to TBK1 being fully activated. Furthermore, the proposed model also includes an OPTN-TBK1 positive feedback loop; however, the earliest reactions in the positive feedback loop are too difficult to observe. For example, it’s widely known that PINK1 and Parkin form a positive feedback loop to generate ubiquitin-chains on damaged mitochondria, but the initial reaction has yet to be observed. It remains unclear if PINK1 is the first to phosphorylate mitochondrial ubiquitin (if this is the case, it remains unknown how ubiquitin comes to mitochondria) or if cytosolic Parkin first adds ubiquitin to the outer membrane albeit with very weak activity. Similarly, in our proposed model, we cannot determine the earliest OPTN-TBK1 reaction. As described in the Discussion in the revised manuscript, it remains possible that in the absence of autophagy machinery OPTN distributed freely on the outer membrane can induce trans-autophosphorylation, albeit weakly, as the earliest reaction.
We would like to thank Reviewer 3 for the critical comments and suggestions. We have performed several of the suggested experiments, added new data, and rewritten the text. We hope that these changes have sufficiently addressed the reviewer’s concerns.
China hat einen lange angekündigten Plan zur Methan-Reduktion veröffentlicht. Der Zeitpunkt am Ende von Gesprächen des chinesischen Klima-Beauftragten Xie Zhenhua mit seinem amerikanischen Pendant John Kerry gilt als Signal für mehr Kooperation zwischen China und den USA im Vorfeld der COP28. https://www.theguardian.com/world/2023/nov/08/china-methane-plan-climate-agreement-us
Mehr zu Methan-Emissionen und -Reduktion: https://hypothes.is/search?q=tag%3A%27process%3A+methane+reduction%27
Author Response
The following is the authors’ response to the original reviews.
Cook, Watt, and colleagues previously reported that a mouse model of Spinocerebellar ataxia type 6 (SCA6) displayed defects in BDNF and TrkB levels at an early disease stage. Moreover, they have shown that one month of exercise elevated cerebellar BDNF expression and improved ataxia and cerebellar Purkinje cell firing rate deficits. In the current work, they attempt to define the mechanism underlying the pathophysiological changes occurring in SCA6. For this, they carried out RNA sequencing of cerebellar vermis tissue in 12-month-old SCA6 mice, a time when the disease is already at an advanced stage, and identified widespread dysregulation of many genes involved in the endo-lysosomal system. Focusing on BDNF/TrkB expression, localization, and signaling they found that, in 7-8 month-old SCA6 mice early endosomes are enlarged and accumulate BDNF and TrkB in Purkinje cells. Curiously, TrkB appears to be reduced in the recycling endosomes compartment, despite the fact that recycling endosomes are morphologically normal in SCA6. In addition, the authors describe a reduction in the Late endosomes in SCA6 Purkinje cells associated with reduced BDNF levels and a probable deficit in late endosome maturation.
We would like to thank the reviewers for their careful reading of the paper, their feedback has helped us to add information and experiments to the paper that enhance the clarity of the findings.
Strengths:
The article is well written, and the findings are relevant for the neuropathology of different neurodegenerative diseases where dysfunction of early endosomes is observed. The authors have provided a detailed analysis of the endo-lysosomal system in SCA6 mice. They have shown that TrkB recycling to the cell membrane in recycling endosomes is reduced, and the late endosome transport of BDNF for degradation is impaired. The findings will be crucial in understanding underlying pathology. Lastly, the deficits in early endosomes are rescued by chronic administration of 7,8-DHF.
We thank the reviewers for their positive feedback on this work.
Weaknesses:
The specificity of BDNF and TrkB immunostaining requires additional controls, as it has been very difficult to detect immunostaining of BDNF. In addition, in many of the figures, the background or outside of Purkinje cell boundaries also exhibits a positive signal.
We agree with the reviewers that the performance of the BDNF and TrkB antibodies is an important concern. We have ourselves had difficulties with the performance of many antibodies and the images in this paper are the result of many years of optimization. We have therefore added further detail about the antibody optimization to the methods section of this paper, and have carried out new staining experiments with additional controls. We have added 2 new figure panels in supplementary figures 3 and 4 to demonstrate these tests.
In the case of anti-BDNF antibodies, we have tested several antibodies and staining protocols and found that in our hands, the only antibody that reliably stained BDNF with a good signal to noise ratio was the one used in this paper (abcam ab108319). Even for this antibody, the staining was greatly enhanced by the use of a heat induced epitope retrieval (HIER) step, which allowed the visualization of BDNF within intracellular structures such as endosomes. When we quantified the intensity of this staining in our previous paper, the results were in agreement with those from a BDNF ELISA used to measure levels of BDNF in the cerebellar vermis of WT and SCA6 mice (Cook et al., 2022), which corroborates these results. As the staining was carried out in tissue sections and not dissociated cells, we also see positive signal from the BDNF staining outside of the Purkinje cells, since BDNF acts on cell-surface receptors and is thus released into the extracellular space around cells (Kuczewski et al., 2008) and is detectable in the extracellular matrix (Lam et al., 2019) and presynaptic terminals around neurons (Camuso et al., 2022; Choo et al., 2017). This is in contrast to studies that image BDNF mRNA with in-situ hybridization, which labels BDNF mRNA predominantly found in cells, and cannot tell us about sub-cellular or extracellular localization of BDNF protein. Together, these factors explain why we observe staining that is not cell- limited, but extends into the space around the cells of interest.
We have added an additional supplemental figure to demonstrate the importance of using HIER when staining slices with anti-BDNF (Supplementary figure 3). We tested HIER protocols that involved heating the slices to 95°C in a variety of buffers. The buffers tested were sodium citrate buffer (10 mM sodium citrate, 0.05% Tween 20, pH 6), Tris buffer (10mM TBS, 0.05% Tween 20, pH 10), EDTA buffer (1mM EDTA, 0.05% Tween 20, pH 8) and neutral PBS. The PBS produced the best result, enhancing the staining of both anti-BDNF and anti-EEA1 antibodies (Supplementary figure 3). Therefore all slices stained using those antibodies were heated to 95°C in PBS using a heat block or thermocycler for 10 minutes, then allowed to cool before staining proceeded.
The antibody we use (abcam ab108319) has been used in hundreds of other publications, including Javed et al., 2021 who ectopically expressed BDNF and noted colocalization between the antibody staining and the GFP tag of the BDNF construct, and Lejkowska et al., 2019 who overexpressed BDNF and saw a dramatic increase in antibody staining as well. The colocalization between ectopically expressed BDNF and the antibody in these studies demonstrates the specificity of the antibody.
However, to further validate antibody specificity we used liver tissue as a negative control. In liver tissue from rodents and humans, the majority of the liver contains negligible levels of BDNF (Koppel et al., 2009; Vivacqua et al., 2014), see also the Human Protein Atlas. The exception is some cholangiocytes: epithelial cells that express BDNF at high levels (Vivacqua et al., 2014). We obtained liver tissue from a WT mouse that was undergoing surgery for an unrelated project and fixed and processed the tissue as we did for brain tissue (outlined in methods section). As we would expect, most of the cells in the liver showed BDNF immunoreactivity that was comparable to background levels (Supplementary figure 3). Interestingly, we were also able to detect sparse highly BDNF-positive cells in the liver, presumed cholangiocytes (Supp. Fig. 3). This pattern of liver BDNF expression is as predicted in the literature, and thus acts as a control for our antibody. We therefore believe that in our hands this antibody is able to stain BDNF with an appropriate degree of specificity.
We also carried out staining experiments using a second anti-TrkB antibody that we had previously used to detect TrkB via Western bloing. We carried out immunohistochemistry as previously described using tissue sections from a WT mouse. The staining with the two different antibodies was carried out at the same time and all other reagents were kept constant. We found that both antibodies labelled TrkB in a similar pattern of localization, including in the early endosomes of the Purkinje cells (Supplementary figure 4). The second antibody however did have a lower signal to noise ratio and so we believe that the original anti-TrkB antibody used in this manuscript (EMD Millipore ab9872) is optimal for staining cerebellar tissue sections in our hands.
One important concern about the conclusions is that the RNAseq experiment was conducted in 12-month- old SCA6 mice suggesting that the defects in the endo-lysosomal system may be caused by other pathophysiological events and, likewise, the impairment in BDNF signaling may also be indirect, as also noted by the authors. Indeed, Purkinje cells in SCA6 mice have an impaired ability to degrade other endocytosed cargo beyond BDNF and TrkB, most likely because of trafficking deficits that result in a disruption in the transport of cargo to the lysosomes and lysosomal dysfunction.
We agree with the reviewers that the defects in the endo-lysosomal system may be caused by other events occurring in the course of disease progression. As mentioned by the reviewers, we have noted this possibility in the text. Detailed investigation into the sequence of events and the root causes of signaling disruption in SCA6 merits future study and we aim to address this in future work. We have expanded this explanation in the text.
Moreover, the beneficial effects of 7,8-DHF treatment on motor coordination may be caused by 7,8-DHF properties other than the putative agonist role on TrkB. Indeed, many reservations have been raised about using 7,8-DHF as an agonist of TrkB activity. Several studies have now debunked (Todd et al. PlosONE 2014, PMID: 24503862; Boltaev et al. Sci Signal 2017, PMID: 28831019) or at the very least questioned (Lowe D, Science 2017: see Discussion: https://www.science.org/content/blog-post/those-compounds-aren-t- what-you-think-they-are Wang et al. Cell 2022 PMID: 34963057). Another interpretation is that 7,8-DHF possesses antioxidant activity and neuroprotection against cytotoxicity in HT-22 and PC12 cells, both of which do not express TrkB (Chen et al. Neurosci Lett 201, PMID: 21651962; Han et al. Neurochem Int. 2014, PMID: 24220540). Thus, while this flavonoid may have a beneficial effect on the pathophysiology of SCA6, it is most unlikely that mechanistically this occurs through a TrkB agonistic effect considering the potent anti-oxidant and anti-inflammatory roles of flavonoids in neurodegenerative diseases (Jones et al. Trends Pharmacol Sci 2012, PMID: 22980637).
We thank the reviewers for raising this important point. We have noted in our previous paper (Cook et al., 2022) that 7,8-DHF may not be acting as a TrkB agonist in SCA6 mice, and are in agreement that other explanations are possible. We have now added information to the text of this paper to highlight this possibility. We did show in our previous paper that 7,8-DHF administration activates Akt signaling in the cerebellum of SCA6 mice, a signaling event that is known to take place downstream of TrkB activation. Additionally, 7,8-DHF treatment led to the increase of TrkB levels in the cerebellum of SCA6 mice (Cook et al., 2022), implicating TrkB in the mechanism of action, even if mechanistically, this is not via direct TrkB activation alone. However, even if the mechanism is currently incompletely explained, we believe that 7,8- DHF remains a valuable treatment strategy for SCA6. We have tried to rewrite the Discussion to highlight what we think is the most important takeaway: that 7,8-DHF can rescue endosomal and other deficits in SCA6, even if we do not currently know the full mechanism of action. We have therefore amended the text to add more detail about other potential explanations for the mechanism of action of 7,8-DHF.
References
Camuso S, La Rosa P, Fiorenza MT, Canterini S. 2022. Pleiotropic effects of BDNF on the cerebellum and hippocampus: Implications for neurodevelopmental disorders. Neurobiol Dis. doi:10.1016/j.nbd.2021.105606
Choo M, Miyazaki T, Yamazaki M, Kawamura M, Nakazawa T, Zhang J, Tanimura A, Uesaka N, Watanabe M, Sakimura K, Kano M. 2017. Retrograde BDNF to TrkB signaling promotes synapse elimination in the developing cerebellum. Nat Commun 8:195. doi:10.1038/s41467-017-00260-w
Cook AA, Jayabal S, Sheng J, Fields E, Leung TCS, Quilez S, McNicholas E, Lau L, Huang S, Watt AJ. 2022. Activation of TrkB-Akt signaling rescues deficits in a mouse model of SCA6. Sci Adv 8:3260. doi:10.1126/sciadv.abh3260
Javed S, Lee YJ, Xu J, Huang WH. 2021. Temporal dissection of Rai1 function reveals brain-derived neurotrophic factor as a potential therapeutic target for Smith-Magenis syndrome. Hum Mol Genet 31:275–288. doi:10.1093/HMG/DDAB245
Koppel I, Aid-Pavlidis T, Jaanson K, Sepp M, Pruunsild P, Palm K, Timmusk T. 2009. Tissue-specific and neural activity-regulated expression of human BDNF gene in BAC transgenic mice. BMC Neurosci 10:68. doi:10.1186/1471-2202-10-68
Kuczewski N, Porcher C, Ferrand N, Fiorentino H, Pellegrino C, Kolarow R, Lessmann V, Medina I, Gaiarsa JL. 2008. Backpropagating action potentials trigger dendritic release of BDNF during spontaneous network activity. J Neurosci 28:7013–7023. doi:10.1523/JNEUROSCI.1673-08.2008
Lam D, Enright HA, Cadena J, Peters SKG, Sales AP, Osburn JJ, Soscia DA, Kulp KS, Wheeler EK, Fischer NO. 2019. Tissue-specific extracellular matrix accelerates the formation of neural networks and communities in a neuron-glia co-culture on a multi-electrode array. Sci Rep 9. doi:10.1038/s41598- 019-40128-1
Lejkowska R, Kawa MP, Pius-Sadowska E, Rogińska D, Łuczkowska K, Machaliński B, Machalińska A. 2019. Preclinical Evaluation of Long-Term Neuroprotective Effects of BDNF-Engineered Mesenchymal Stromal Cells as Intravitreal Therapy for Chronic Retinal Degeneration in Rd6 Mutant Mice. Int J Mol Sci 2019, Vol 20, Page 777 20:777. doi:10.3390/IJMS20030777
Vivacqua G, Renzi A, Carpino G, Franchitto A, Gaudio E. 2014. Expression of brain derivated neurotrophic factor and of its receptors: TrKB and p75NT in normal and bile duct ligated rat liver. Ital J Anat Embryol 119:111–129. doi:10.13128/IJAE-15138
Author Response
The following is the authors’ response to the original reviews.
We thank the reviewers and editor for their thoughful and careful evaluation of our manuscript. We appreciate your time and effort and have incorporated many of these suggestions to improve our revised manuscript.
Reviewer #1 (Public Review):
Summary: Cullinan et al. explore the hypothesis that the cytoplasmic N- and C-termini of ASIC1a, not resolved in x-ray or cryo-EM structures, form a dynamic complex that breaks apart at low pH, exposing a C-terminal binding site for RIPK1, a regulator of necrotic cell death. They expressed channels tagged at their N- and C-termini with the fluorescent, non-canonical amino acid ANAP in CHO cells using amber stop-codon suppression. Interaction between the termini was assessed by FRET between ANAP and colored transition metal ions bound either to a cysteine reactive chelator attached to the channel (TETAC) or metal-chelating lipids (C18-NTA). A key advantage to using metal ions is that they are very poor FRET acceptors, i.e. they must be very close to the donor for FRET to occur. This is ideal for measuring small distances/changes in distance on the scales expected from the initial hypothesis. In order to apply chelated metal ions, CHO cells were mechanically unroofed, providing access to the inner leaflet of the plasma membrane. At high pH, the N- and C- termini are close enough for FRET to be measured, but apparently too far apart to be explained by a direct binding interaction. At low pH, there was an apparent increase in FRET between the termini. FRET between ANAP on the N-and Ctermini and metal ions bound to the plasma membrane suggests that both termini move away from the plasma membrane at low pH. The authors propose an alternative hypothesis whereby close association with the plasma membrane precludes RIPK1 binding to the C-terminus of ASIC1a.
Strengths: The findings presented here are certainly valuable for the ion channel/signaling field and the technical approach only increases the significance of the work. The choice of techniques is appropriate for this study and the results are clear and high quality. Sufficient evidence is presented against the starting hypothesis.
Weaknesses: I have a few questions about certain controls and assumptions that I would like to see discussed more explicitly in the manuscript.
My biggest concern is with the C-terminal citrine tag. Might this prevent the hypothesized interaction between the N- and C-termini? What about the serine to cysteine mutations? The authors might consider a control experiment in channels lacking the C-terminal FP tag.
While it is certainly possible that the C-terminal citrine tag is preventing the hypothesized interaction between the intracellular termini, there are a few things that mitigate (but not eliminate) this concern. First, previous work looking at the interaction between the intracellular termini used FPs on both the N- and C-termini and concluded that in fact there is an interaction (PMID:31980622). Our channels have only a single FP, and we use a higher resolution FRET approach. Second, we aVach our citrine tag with a 11-residue linker, allowing for enhanced flexibility of the region and hopefully allowing for more space for an interaction that was posited to be between the very proximal part of the C-terminus (near the membrane and away from the tag) and the untagged N-terminus. Third, we previously showed that Stomatin, a much larger protein than the NTD, could bind the distal C-terminus of rASIC3 with a large fluorescent protein connected by the same linker on the C-terminus. In the case of Stomatin, the interaction involved the residues at the distal portion of the C-terminus close to the bulky FP. Interestingly, while we did not publish this, without this flexible linker, Stomatin could not regulate the channel and likely did not bind.
Despite this, we agree that this is possible and have added a statement in our limitations section explicitly saying this.
Figure 2 supplement 1 shows apparent read-through of the N-terminal stop codons. Given that most of the paper uses N-terminal ANAP tags, this figure should be moved out of the supplement. Do Nterminally truncated subunits form functional channels? Do the authors expect N-terminally truncated subunits to co-assemble in trimers with full-length subunits? The authors should include a more explicit discussion regarding the effect of truncated channels on their FRET signal in the case of such co-assembly.
The positions that show readthrough (E6, L18, H515) were not used in the study. We eliminated them largely on the basis of these westerns. We elected to put the bulk of the blots in the supplement simply because of how many there were. We believe this is the best compromise. It allows us to show representative blots for all our positions without making an illegible figure with 7 blots.
The N-terminally truncated subunits would create very short peptides that are not able to create functional channels. A premature stop at say E8 would create a 7-mer. Our longest N-terminal truncation would only create a protein of 32 amino acids. These don’t contain the transmembrane segments and thus cannot make functional channels.
As the epitope used for the western blots in Figure 2 and supplements is part of the C-terminal tag, these blots do not provide an estimate of the fraction of C-terminally truncated channels (those that failed to incorporate ANAP at the stop codon). What effect would C-terminally truncated channels have on the FRET signal if incorporated into trimers with full-length subunits?
Alternatively, C-terminally truncated subunits would be able to form functional channels because they contain the full N-terminus, the transmembrane domains, the extracellular domain and a portion of the C-terminus. We don’t think this is a major contaminant to our experiments. The only two C-terminal ANAP positions we use are 464 and 505. In each of these cases, they are only used for memFRET. The ones that do not contain ANAP are essentially “invisible” to the experiment. Since we are measuring their proximity to the membrane, having some missing should not maVer. However, there is some chance that truncations in some subunits could allosterically affect the position of the CT in other subunits. We have added a discussion of this in the manuscript.
Some general discussion of these results in the context of trimeric channels would be helpful. Is the putative interaction of the termini within or between subunits? Are the distances between subunits large enough to preclude FRET between donors on one subunit and acceptor ions bound on multiple subunits?
Thank you for this comment. We did not directly test whether the distances are within or between subunits. We considered using a concatemer to do this, however, the concatemeric channels do not express particularly well. Then, UAA incorporation hurts the expression as well. It was unlikely we would be able to get sufficient expression for tmFRET.
However, the Maclean group has previously tested this using FRET between concatenated subunits and determined that FRET is stronger within than between subunits. We have updated the manuscript to reflect a more thorough discussion of our results in the context of their trimeric assembly.
The authors conclude that the relatively small amount of FRET between the cytoplasmic termini suggests that the interaction previously modeled in Rosetta is unlikely. Is it possible that the proposed structure is correct, but labile? For example, could it be that the FRET signal is the time average of a state in which the termini directly interact (as in the Rosetta model) and one in which they do not?
The proposed RoseVa model does not include the reentrant loop of the channel, so it is probable that this model would change if it were redone to include this new feature of the channel.
However, we do discuss the limitation of FRET as a method that measures a time average that is weighted towards closest approach in our discussion section. The termini are most certainly dynamic and it is possible that spend some time in close proximity. Given that FRET is biased towards closest approach, we actually think this strengthens our argument that the termini don’t spend a great deal of time in complex. In addition, our MST data suggests that the termini do not bind. We have added some commentary on this to the discussion section for clarity.
Reviewer #2 (Public Review):
Summary:
The authors use previously characterised FRET methods to measure distances between intracellular segments of ASIC and with the membrane. The distances are measured across different conditions and at multiple positions in a very complete study. The picture that emerges is that the N- and C-termini do not associate.
Strengths:
Good controls, good range of measurements, advanced, well-chosen and carefully performed FRET measurements. The paper is a technical triumph. Particularly, given the weak fluorescence of ANAP, the extent of measurements and the combination with TETAC is noteworthy.
The distance measurements are largely coherent and favour the interpretation that the N and C terminus are not close together as previously claimed.
Weaknesses:
One difficulty is that we do not have a positive control for what binding of something to either N- or Cterminus would look like (either in FRET or otherwise).
We acknowledge that this is a challenge for the approach. Having a positive control for binding would be great but we are not sure such a thing exists. You could certainly imagine a complex between two domains where each label (ANAP and TETAC) are pointed away from one other (giving comparatively modest quenching) or one where they are very close (giving comparatively large quenching), both of which could still be bound. This is essentially a less significant version of the problem with using FPs to measure proximity…they are not very good proxies for the position of the termini. These small labels are certainly beVer proxies but still not perfect. Our conclusion here is based more on the totality of the data. We tried many combinations and saw no sign of distances closer than ~ 20A at resting pH. We think the simplest explanation is that they are not close to one another but we tried to lay out the limitations in the discussion.
One limitation that is not mentioned is the unroofing. The concept of interaction with intracellular domains is being examined. But the authors use unroofing to measure the positions, fully disrupting the cytoplasm. Thus it is not excluded that the unroofing disrupts that interaction. This should be mentioned as a possible (if unlikely) limitation.
Thank you for your comment. We discuss unroofing as a potential limitation because it exposes both sides of the plasma membrane to changes in pH. We have updated this section to include acknowledgement of the possibility that unroofing disrupts the interaction via washout of other critical proteins.
Reviewer #3 (Public Review):
Summary: The manuscript by Cullinan et al., uses ANAP-tmFRET to test the hypothesis that the NTD and CTD form a complex at rest and to probe these domains for acid-induced conformational changes. They find convincing evidence that the NTD and CTD do not have a propensity to form a complex. They also report these domains are parallel to the membrane and that the NTD moves towards, and the CTD away, from the membrane upon acidification.
Strengths:
The major strength of the paper is the use of tmFRET, which excels at measuring short distances and is insensitive to orientation effects. The donor-acceptor pairs here are also great choices as they are minimally disruptive to the structure being studied.
Furthermore, they conduct these measurements over several positions with the N and C tails, both between the tails and to the membrane. Finally, to support their main point, MST is conducted to measure the association of recombinant N and C peptides, finding no evidence of association or complex formation.
Weaknesses:
While tmFRET is a strength, using ANAP as a donor requires the cells to be unroofed to eliminate background signal. This causes two problems. First, it removes any possible low affinity interacting proteins such as actinin (PMID 19028690). Second, the pH changes now occur to both 'extracellular' and 'intracellular' lipid planes. Thus, it is unclear if any conformational changes in the N and CTDs arise from desensitization of the receptor or protonation of specific amino acids in the N or CTDs or even protonation of certain phospholipid groups such as in phosphatidylserine. The authors do comment that prolonged extracellular acidification leads to intracellular acidification as well. But the concerns over disruption by unroofing/washing and relevance of the changes remain.
We acknowledge that unroofing is a limitation of our approach and noted it in the discussion. However, we have updated the section to include the possibility that the act of unroofing and washing could also disrupt the potential interaction between the intracellular domains as well as between these domains and other intracellular proteins. This was the best approach we could use to address our questions and it required that we unroof the cells. However, we look forward to future studies or new techniques that do not require the unroofing of the cells.
The distances calculated depend on the R0 between donor and acceptor. In turn, this depends on the donor's emission spectrum and quantum yield. The spectrum and yield of ANAP is very sensitive to local environment. It is a useful fluorophore for patch fluorometry for precisely this reason, and gating-induced conformational changes in the CTD have been reported just from changes in ANAP emission alone (PMID 29425514). Therefore, using a single R0 value for all positions (and both pHs at a single position) is inappropriate. The authors should either include this caveat and give some estimate of how big an impact changes spectrum and yield might have, or actually measure the emission spectra at all positions tested.
This is a reasonable concern and one we considered. Measuring the quantum yield would be quite difficult. However, we have measured spectra at a number of positions and see a relatively minimal shik in the peak. Most positions peak between 481 and 484nm. If you calculate the difference in R0 using theoretical spectra with a blue shik of 20nm, the difference in R0 is only ~1.5A. A shik of 20nm is on the higher side of anything we have seen in the literature (PMID 30038260) and since even with that large a shik, the difference is minimal we do not think measuring spectra for each position would impact the overall conclusions presented. As you noted, though, the quantum yield also changes. Assuming a change in yield from 0.22 to 0.47, the largest we found reported in the literature (PMID:29923827) , the R0 would increase by 2A. This same paper showed that the blue shiked position was the one with the higher extinction coefficient so these changes would be working in opposition to one another making the difference in R0 even smaller. It is important to note, that while tmFRET is a much more powerful measure of distance than standard FRET, these distances, as you point out, are quite challenging to measure precisely. Our conclusions are based less on the absolute distances and more on the observation that no positions show large quenching and that if there is any change upon acidification, it is in the wrong direction.
Overall, the writing and presentation of figures could be much improved with specific points mentioned in the recommendations for authors section.
See below.
The authors argue that the CTD is largely parallel to the plasma membrane, yet appear to base this conclusion on ANAP to membrane FRET of positions S464 and M505. Two positions is insufficient evidence to support such a claim. Some intermediate positions are needed.
We do not see in the paper where we suggest that the CTD is parallel. However, your point that we could try and determine if this was the case is correct. However, we aVempted to create several other CTD TAG mutants but struggled with readthrough and poor expression of these mutants so we opted to just include S464 and M505. Our point from these data is only that the distal CTD (505) must spend significant time near the membrane to explain our FRET data.
Upon acidification, NTD position Q14 moves towards the plasma membrane (Figure 8B). Q14 also gets closer to C515 or doesn't change relative to 505 (Figures 7C and B) upon acidification. Yet position 505 moves away from the membrane (Figure 8D). How can the NTD move closer to the membrane, and to the CTD but yet the CTD move further from the membrane? Some comment or clarification is needed.
This is a reasonable question and one that is hard to definitively answer. Our goal here was to test the hypothesis that the termini are bound at rest. Mapping the precise positions of the termini is difficult for reasons we will enumerate in the question that asks why we didn’t make a model. There are potentially multiple explanations but the easiest one would be that the CTD could move away from the membrane but closer to Q14, for instance, if the distal termini, say, rotated towards the NTD. This would move 505 closer and have no impact on whether or not the NTD and CTD moved away or toward the membrane.
Reviewer #1 (Recommendations For The Authors):
Minor concerns
The authors show the spectrum of ANAP attached to beads and use this spectrum to calculate R0 for their FRET measurements. Peak ANAP fluorescence is dependent on local environment and many reports show ANAP in protein blue-shiked relative to the values reported here. How would this affect the distance measurements reported?
This is an important point. See above for the answer.
Could the lack of interaction between the N- and C-terminal peptides in Figure 7 arise from the cysteine to serine mutations or lack of structure in the synthetic peptides. How were peptide concentrations measured/verified for the experiment?
It is possible that cysteine to serine mutations could prevent the interaction. It is also possible that these peptides are not capable of adopting their native fold without the presence of the plasma membrane or due to being synthetically created. However, the termini are thought to be largely unstructured. We received these peptides in lyophilized form at >95% purity and resuspended to our desired stock concentration (3 mM C-terminus, 1 mM N-terminus). Even if our concentration was off, we see no signs of interaction up to quite a high concentration.
How was photobleaching measured for correcting the data?
We executed several mock experiments at various TAG positions using either pH 8 and pH 6, where we performed the experiments as usual but with a mock solution exchange when we would normally add the metal. We normalized the L-ANAP fluorescence to the first image and averaged together these values for pH 8 and pH 6. We then corrected using Equation 2 in the manuscript..
We have updated the methods to include how we adjusted for bleaching.
The authors may wish to make it more explicit that their Zn2+ controls also preclude the possibility that a changing FRET signal between ANAP and citrine may affect their data.
Thank you for this comment. We agree, it would strengthen the manuscript to include this statement. We have now included this.
It might be useful to the reader if the authors could include (as a supplement) plots of their data (like in Figure 6), in which FRET efficiency has been converted to distance.
We considered this idea as well but felt like showing the actual data in the figures and the distances in a table would be best.
Figure 5D is mentioned in the text before any other figures. This is unconventional. Could this panel be moved to Figure 1 or the mention moved to later?
Changed
western blot is not capitalized.
Changed.
Figure 1, the ANAP structure shown is the methyl ester, which is presumably cleaved before ANAP is conjugated to the tRNA. The authors may wish to replace this with the free acid structure.
This is a fair point. We originally used the methyl ester structure to indicate the version of ANAP we chose to use. However, you are correct that the methyl ester is cleaved before conjugation to the tRNA. We replaced the methyl ester with the free acid structure to clarify this.
Figures 1 and 4 should have scale bars for the images.
Scale bars have been added to figures 1, 4, and 5.
In Figure 3, the letters in the structures (particularly TETAC) are way too small. Please increase the font size.
Changed
In Figure 3 and Figure 3 supplement 1, the axes are labeled "Absorbance (M-1cm-1)." Absorbance is dimensionless. The authors are likely reporting the extinction coefficient.
Thank you for catching this. We adjusted the axes to extinction coefficient.
In Figures 5 B and C, it might be clearer if the headers read "Initial, +Cu2+/TETAC, DTT" rather than "Initial, FRET, Recovery."
Changed
The panel labels for Figure 8 seem to be out of order.
Changed
The L for L-ANAP should be rendered, by convention, in small caps.
This is a good example of learning something new from the review process. This is the first I have ever heard of small caps. We can find no other papers that use small caps for L-ANAP so I am not 100% sure what convention this is referring to and don’t want to change the wrong thing in the paper. We are happy to change if the editorial staff at eLife agree but have lek this for now.
Reviewer #2 (Recommendations For The Authors):
With so many distances measured, why was not even a basic structural model attempted?
We certainly considered it, but a number of things lead us to conclude that it might imply more certainty about the structure of these termini than we hope to give. 1) Given that the FRET is a time average of positions, these distance constraints would not do much constraining. 2) Given that the termini are likely unstructured and flexible this makes the problem in 1 worse. 3) There is no structural information to use as a starting point for a model. 4) The flexibility of the linkers for each FRET pair also introduces uncertainty. This can, in theory, be modeled as they do in EPR but all of this together made us decide not to do this. What we hope readers take home, is the overall picture of the data is not consistent with the original RIPK1 hypothesis.
Maybe it would be good to draw a band on the graphs in Figure 6 for the FRET signal expected for interaction (and thus, disfavoured by these data). This would at least give context.
We agree this could be helpful, but it is not so easy to do. What distance would we choose? We could put a line at ~5Å (the model predicted distance). As we noted above, a number of distances could be compatible with an interaction. However, we think it’s unlikely that if a complex was formed that none of our measurements would show a distance closer than 20Å at rest and that an unbinding event would then lead to a decrease in distance. This, to us, is the take home message.
Minor points:
"Aker unroofing the cells, only fluorescence associated with the "footprint", or dorsal surface, of the cell membrane is lek behind."
The authors use dorsal and ventral in this section to describe parts of an adherent cell. But in the first instance, they remove the dorsal part of the cell, and then in this phrase, the dorsal part is lek behind....I am a bit confused.
Thank you for pointing out this mistake, we have fixed this. It is indeed the ventral surface lek behind.
"bind at rest an" - and?
Changed
"One previous study used a different approach to try and map the topography of the intracellular termini of ASIC1a comparable to our memFRET experiments." I think a citation is due.
Citation added
"great deal of precedent" even if this result is from my own lab, I would prefer that the authors note that it's one study from one lab! I think best just to delete "great deal of".
“Great deal of” deleted
I think the column "Significance" in the tables is unnecessary when the P value is given.
Thank you for this suggestion. We agree and have made the change.
Figure 7a Q14TAG has a clearly bimodal distribution at pH 8. What could be the meaning of this result? The authors do not mention it that I could find. Perhaps there is no meaning. The authors should state what they think is (or is not) going on.
This is a good question and we don’t have a good answer. It appears to be experimental variability. The data from the “low fret” in this experimental condition all came from the same days. So something was different that day. We considered that they might be outliers to exclude but thought showing all of our data was the beVer path. We reperformed the ANOVA here separating out the “outlier” day and nothing of substance changed. Both populations were still different with P value less than 0.001.
Typo: Lumencore
Changed
Maybe just a matter of taste but the panel created with Biorender in Figure 8 is not attractive and depicts the channel differently to in Figure 5D, which is again different from Figure 1A. Surely one advantage of using computer-generated artwork could be to have consistency.
We agree and have used the same cartoon for all of our images with the one exception being the schematics that are just meant to show the positions that are present in each bar graph.
Figure 4A was squashed to fit (text aspect ratio is wrong).
Fixed
Reviewer #3 (Recommendations For The Authors):
Citrine is used to report incorporation. Yet citrine has a strong tendency to dimerize (PMID 27240257). Did the authors use mCitrine or just Citrine? This is quite important in interpreting their data.
Thank you for pointing out this important distinction. We use mCitirine which we have added to the methods.
The manuscript has numerous instances of imprecise language. For example, page 10, last para, first line, "previous studies have looked at..." or page 7, final paragraph "tell a similar story". Related, the figures could be much better. For example, in Figure 1, where the authors depict the anap chemical in red, as opposed to the blue one might expect of a blue emiqng fluorophore. In figure 6, ANAP is also in red with the quenching group in green. This is opposite to how one typically thinks of FRET with the warmer color being the acceptor not the donor. Moreover, the pH 6 condition is also colored the same shade of red as the ANAP. Labels of Cys positions would again be useful here. In Figure 3, the heteroatoms of TETAC and C18-NTA are very small and difficult to see. It would also be good to label these structures, and the spectra below, so the reader can tell at a glance without looking at the caption, what the structures and spectra arise from. Also, how are the absorption spectra normalized? This is not discussed in the methods. The lack of attention to presentation mars an otherwise nice study.
Thank you for these points. We have made modifications to the manuscript to address these comments.
Abstract, second last line "Aker prolonged acidification, ...", 'prolonged' could be interpreted as 'it takes a while for the domain to move' or 'the movement only happens aker a while'. This not what the authors intend to convey. Consider modifying to just 'Aker acidification,'
We updated the main text to indicate that prolonged acidification is intended to describe acidification that occurs over the minutes timescale.
Pdf page 6, bottom para on Anap incorporation not altering channel function: What is meant by 'steady state pH dependence of activation'? This implies the authors applied a pH stimulus, then waited until equilibrium was achieved ie. until desensitization was complete and measured the current at that point. It seems more likely they simply applied different pH stimuli and measured the peak response and that the use of 'steady state' here is a typo.
We removed the phrase steady state.
Same section, controls of electrophysiology allude to 485, 505 and 515 ANAP-containing channels. In fact, the authors have no way of determining what fraction (if any) of the pH evoked currents arise from channels containing Anap in those positions versus from simply having a translation stop but still functioning. This should be mentioned.
This is correct. We cannot be sure the CTD TAG positions are not a mixture of ANAP-containing channels and truncations. See above for why we do not think this a big concern for the FRET experiments. Functionally, though, you are correct that we cannot tell. We now mention this in the paper.
Methods, the abbreviation for SBT should be defined somewhere.
Added.
Methods, unroofing section, middle paragraph, the authors use nM not nm to list wavelengths of light.
Changed.
Figure 3C-D: There's an unexpected blip in the Anap emission spectra at ~500 nm. Are the grating efficiency of the spectrograph and quantum efficiency of the camera accounted for in these spectra?
This is a good question. The data are not corrected for either camera efficiency or grating efficiency. We don’t have easy access to the actual data (although we can see a pdf version of each). There is a liVle blip in the grating efficiency graph that could partly explain the blip in our spectra.
Figure 5C, were recovery experiments routinely done? If so, would be good to show more than n = 1 in the plot to get an idea of reproducibility.
Recovery experiments were done in every experiment but are not shown for simplicity. We have included all FRET and recovery data for position Q14TAG-C469 at pH 6 in figure 5C to show reproducibility of our FRET and recovery data.
Table 1, considering adding a Δ distance column (pH 8 versus 6) so the magnitude of changes are more easily seen.
This is a reasonable suggestion but we decided not to include a Δ distance column. The data are whole numbers and people can easily determine the Δ distance. We felt that including that column would bring too much focus on what we think are preVy small changes. Our hope is that readers take away that the data are not consistent with complex formation between the determine and focus less on absolute distances.
Figure 7A, Q14tag pH 8 condition has a quite a bit of spread and, likely, two populations. These data, as well as G11, are unlikely to be parametric and hence ANOVA is inappropriate. A normality test, and likely Kruskal-Wallis test is called for.
Aker testing for normality, the data for Q14TAG C485 pH8 are non-normally distributed. However, a Kruskal Wallis is a non-parametric test for a one-way ANOVA and not applicable here. We separated the data out into population 1 and 2 and repeated the two-way ANOVA statistical test. When Q14TAG pH 8 is split into 2 populations, the statistics hardly change. When the data is not separated, Q14TAG pH 8 relative to pH 6 has a p-value <0.0001. When the 2 populations are separated, both populations relative to Q14TAG pH 6 still have a p-value of <0.0001.
Reviewer #1 (Public Review):
Cullinan et al. explore the hypothesis that the cytoplasmic N- and C-termini of ASIC1a, not resolved in x-ray or cryo-EM structures, form a dynamic complex that breaks apart at low pH, exposing a C-terminal binding site for RIPK1, a regulator of necrotic cell death. They expressed channels tagged at their N- and C-termini with the fluorescent, non-canonical amino acid ANAP in CHO cells using amber stop-codon suppression. Interaction between the termini was assessed by FRET between ANAP and colored transition metal ions bound either to a cysteine reactive chelator attached to the channel (TETAC) or metal-chelating lipids (C18-NTA). A key advantage to using metal ions is that they are very poor FRET acceptors, i.e. they must be very close to the donor for FRET to occur. This is ideal for measuring small distances/changes in distance on the scales expected from the initial hypothesis. In order to apply chelated metal ions, CHO cells were mechanically unroofed, providing access to the inner leaflet of the plasma membrane. At high pH, the N- and C- termini are close enough for FRET to be measured, but apparently too far apart to be explained by a direct binding interaction. At low pH, there was an apparent increase in FRET between the termini. FRET between ANAP on the N-and C-termini and metal ions bound to the plasma membrane suggests that both termini move away from the plasma membrane at low pH. The authors propose an alternative hypothesis whereby close association with the plasma membrane precludes RIPK1 biding to the C-terminus of ASIC1a.
The findings presented here are certainly valuable for the ion channel/signaling field and the technical approach only increases the significance of the work. The choice of techniques is appropriate for this study and the results are clear and high quality. Sufficient evidence is presented against the starting hypothesis. I have a few questions about certain controls and assumptions that I would like to see discussed more explicitly in the manuscript.
--As discussed by the authors, the C-terminal citrine could potentially disrupt the hypothesized interaction between the N- and C-termini.
--There is apparent read-through of some of the stop codons in the absence of ANAP, which could complicate interpretation of the experiments. The largest amount of read-through is for the E6TAG, L18TAG, and H515TAG constructs, which were not used for further experiments. However, some degree of read-through is evident from western blots for V10TAG, Q14TAG, L41TAG, and A44TAG as well.
Since the epitope used for western blots is on the C-terminus of the protein, the blots do not show the fraction of truncated protein. As discussed by the authors, N-terminally truncated constructs would be too small to assemble into channels. In constructs with the TAG codon towards the C-terminus, there is the potential for co-assembly of full-length and truncated subunits into trimers. Truncated subunits would not contribute directly to the fluorescence signal, but could potentially have allosteric effects on the position of the C-termini of full-length ANAP-tagged constructs in the context of a mixed channel.
When does annotating books become a distraction? .t3_17pitv9._2FCtq-QzlfuN-SwVMUZMM3 { --postTitle-VisitedLinkColor: #8c8c8c; --postTitleLink-VisitedLinkColor: #8c8c8c; --postBodyLink-VisitedLinkColor: #989898; }
reply to u/Low-Appointment-2906 at https://www.reddit.com/r/books/comments/17pitv9/when_does_annotating_books_become_a_distraction/
Through the middle ages, bookmakers would not only leave significant margins for readers to annotate, but they also illuminated books and included drolleries which readers in the know would use in conjunction with the arts of memory (from rhetoric) to memorize portions of texts more easily. I strongly suspect this isn't what booktokkers are doing; their practice is likely more like the sorts of decorative #ProductivityPorn one sees in the Bullet journal and journaling spaces. It's performative content creation.
Those interested in refining their practices of "reading with a pen in hand", continuing the "great conversation" or having "conversations with their texts" might profitably start with Mortimer J. Adler's essay: “How to Mark a Book” (Saturday Review of Literature, July 6, 1941). In his 1975 KCET series How to Read a Book, which was based on their book of the same name, Adler mentioned to Charles Van Doren that he would buy new copies of books so he could re-annotate them without being distracted by his older annotations.
Some have solved the problem of distracting annotations by interleaving their books so they've got lots of blank space to write their notes. It's a rarer practice now, but some publishers still print Bibles with blank pages every other page for this practice. Others put their annotations and notes into commonplace books or on index cards for their card index/zettelkasten.
As some have mentioned, friends and lovers through time have shared books with annotations as a way of sharing their thoughts. George Custer and his wife Elizabeth did this with Tennyson.
If you're interested in annotating digitally online, perhaps check out Hypothes.is where I've seen teachers and students using social annotation to read and make sense of books [example]. I've also seen groups of people use this tool for hosting online book groups/clubs.
If you're in it for fun, you might appreciate:
And those wishing to delve more deeply into the history and power of annotation might look at: Kalir, Remi H., and Antero Garcia. Annotation. The MIT Press Essential Knowledge Series. MIT Press, 2019. https://mitpressonpubpub.mitpress.mit.edu/annotation.
Good luck annotating! 📝
Author Response
The following is the authors’ response to the previous reviews
Reviewer #1 (Public Review):
Comments on the original submission:
Trypanosoma brucei undergoes antigenic variation to evade the mammalian host's immune response. To achieve this, T. brucei regularly expresses different VSGs as its major surface antigen. VSG expression sites are exclusively subtelomeric, and VSG transcription by RNA polymerase I is strictly monoallelic. It has been shown that T. brucei RAP1, a telomeric protein, and the phosphoinositol pathway are essential for VSG monoallelic expression. In previous studies, Cestari et al. (ref. 24) has shown that PIP5pase interacts with RAP1 and that RAP1 binds PI(3,4,5)P3. RNAseq and ChIPseq analyses have been performed previously in PIP5pase conditional knockout cells, too (ref. 24). In the current study, Touray et al. did similar analyses except that catalytic dead PIP5pase mutant was used and the DNA and PI(3,4,5)P3 binding activities of RAP1 fragments were examined. Specifically, the authors examined the transcriptome profile and did RAP1 ChIPseq in PIP5pase catalytic dead mutant. The authors also expressed several C-terminal His6-tagged RAP1 recombinant proteins (full-length, aa1300, aa301-560, and aa 561-855). These fragments' DNA binding activities were examined by EMSA analysis and their phosphoinositides binding activities were examined by affinity pulldown of biotin-conjugated phosphoinositides. As a result, the authors confirmed that VSG silencing (both BES-linked and MES-linked VSGs) depends on PIP5pase catalytic activity, but the overall knowledge improvement is incremental. The most convincing data come from the phosphoinositide binding assay as it clearly shows that N-terminus of RAP1 binds PI(3,4,5)P3 but not PI(4,5)P2, although this is only assayed in vitro, while the in vivo binding of full-length RAP1 to PI(3,4,5)P3 has been previously published by Cestari et al (ref. 24) already. Considering that many phosphoinositides exert their regulatory role by modulate the subcellular localization of their bound proteins, it is reasonable to hypothesize that binding to PI(3,4,5)P3 can remove RAP1 from the chromatin. However, no convincing data have been shown to support the author's hypothesis that this regulation is through an "allosteric switch".
Comments on revised manuscript:
In this revised manuscript, Touray et al. have responded to reviewers' comments with some revisions satisfactorily. However, the authors still haven't addressed some key scientific rigor issues, which are listed below:
1) It is critical to clearly state whether the observations are made for the endogenous WT protein or the tagged protein. It is good that the authors currently clearly indicate the results observed in vivo are for the RAP1-HA protein. However, this is not as clearly stated for in vitro EMSA analyses. In addition, in discussion, the authors simply assumed that the c-terminally tagged RAP1 behaves the same as WT RAP1 and all conclusions were made about WT RAP1.
There are two choices here. The authors can validate that RAP1-HA still retains RAP1's essential function as a sole allele in T. brucei cells (as was recommended previously). Indeed, HA-tagged RAP1 has been studied before, but it is the N-terminally HA-tagged RAP1 that has been shown to retain its essential functions. Adding the HA tag to the C-terminus of RAP1 may well cause certain defects to RAP1. For example, N-terminally HA-tagged TERT does not complement the telomere shortening phenotype in TERT null T. brucei cells, while C-terminally GFP-tagged TERT does, indicating that HA-TERT is not fully functional while TERT-GFP likely has its essential functions (Dreesen, RU thesis). Although RAP1-HA behaves similar to WT RAP1 in many ways, it is still not fully validated that this protein retains essential functions of RAP1. By the way, it has been published that cells lacking one allele of RAP1 behave as WT cells for cell growth and VSG silencing (Yang et al. 2009, Cell; Afrin et al. 2020, mSphere). In addition, although RAP1 may interact with TRF weakly, the interaction is direct, as shown in yeast 2-hybrid analysis in (Yang et al. 2009, Cell).
Alternatively, if the authors do not wish to validate the functionality of RAP1-HA, they need to add one paragraph at the beginning of the discussion to clearly state that RAP1-HA may not behave exactly as WT RAP1. This is important for readers to better interpret the results. In addition, the authors need to tune down the current conclusions dramatically, as all described observations are made on RAP1-HA but not the WT RAP1.
The results with RAP1-HA are consistent with previous knowledge of RAP1 interactions with telomeric proteins and DNA. Hence, the C-terminal HA-tagged RAP1 seems, by all measures, functional. Nevertheless, to make it clear for the reader, we added a note in the discussion, lines 244-246: “Although we showed that C-terminal HA-tagged RAP1 protein has telomeric localization (Cestari et al. 2015, PNAS) and interactions with other telomeric proteins (Cestari et al. 2019 Mol Cell Biol); we cannot rule out potential differences between HA-tagged and non tagged RAP1.”
For a similar reason, the current EMSA results truly reflect how C-terminally His6-tagged RAP1 and RAP1 fragments behave. If the authors choose not to remove the His6 tag, it is essential that they use "RAP1-His6" to refer to these recombinant proteins. It is also essential for the authors to clearly state in the discussion that the tagged RAP1 fragments bind DNA, but the current data do not reveal whether WT RAP1 binds DNA. In addition, the authors incorrectly stated that "disruption of the MybL domain sequence did not eliminate RAP1-telomere binding in vivo" (lines 165-166). In ref 29, deletion of Myb domain did not abolish RAP1-telomere association. However, point mutations in MybL domain that abolish RAP1's DNA binding activities clearly disrupted RAP1's association with the telomere chromatin. Therefore, the current observation is not completely consistent with that published in ref 29.
We stated in line 149-150 “…we expressed and purified from E. coli recombinant 6xHistagged T. brucei RAP1 (rRAP1)”. To clarify to the authors, we replaced rRAP1 with rRAP1-His throughout the manuscript and figures. As for the statement that “disruption of the MybL domain sequence did not eliminate RAP1-telomere binding in vivo" (lines 165-166).”. We removed the statement from the manuscript.
2) There is no evidence, in vitro or in vivo, that binding PI(3,4,5)P3 to RAP1 causes conformational change in RAP1. The BRCT domain of RAP1 is known for its ability to homodimerize (Afrin et al. 2020, mSphere). It is therefore possible that binding of PI(3,4,5)P3 to RAP1 simply disrupts its homodimerization function. The authors clearly have extrapolated their conclusions based on available data. It is therefore important to revise the discussion and make appropriate statements.
We did not state that PI(3,4,5)P3 causes RAP1 conformational changes. We discussed the possibility. We stated in lines 199-201: “PI(3,4,5)P3 inhibition of RAP1-DNA binding might be due to its association with RAP1 N-terminus causing conformational changes that affect Myb and MybL domains association with DNA.” This is a reasonable discussion, given the data presented in the manuscript.
Reviewer #2 (Public Review):
In this manuscript, Touray et al investigate the mechanisms by which PIP5Pase and RAP1 control VSG expression in T. brucei and demonstrate an important role for this enzyme in a signalling pathway that likely plays a role in antigenic variation in T. brucei. While these data do not definitively show a role for this pathway in antigenic variation, the data are critical for establishing this pathway as a potential way the parasite could control antigenic variation and thus represent a fundamental discovery.
The methods used in the study are generally well-controlled. The authors provide evidence that RAP1 binds to PI(3,4,5)P3 through its N-terminus and that this binding regulates RAP1 binding to VSG expression sites, which in turn regulates VSG silencing. Overall their results support the conclusions made in the manuscript. Readers should take into consideration that the epitope tags on RAP1 could alter its function, however.
There are a few small caveats that are worth noting. First, the analysis of VSG derepression and switching in Figure 1 relies on a genome which does not contain minichromosomal (MC) VSG sequences. This means that MC VSGs could theoretically be mis-assigned as coming from another genomic location in the absence of an MC reference. As the origin of the VSGs in these clones isn't a major point in the paper, I do not think this is a major concern, but I would not over-interpret the particular details of switching outcomes in these experiments.
We agree with the reviewer and thus made no speculations on minichromosomes. The data analysis must rely on the available genome, and the reference genome used is well-assembled with PacBio sequences and Hi-C data (Muller et al. 2018, Nature).
Another aspect of this work that is perhaps important, but not discussed much by the authors, is the fact that signalling is extremely poorly understood in T. brucei. In Figure 1B, the RNA-seq data show many genes upregulated after expression of the Mut PIP5Pase (not just VSGs). The authors rightly avoid claiming that this pathway is exclusive to VSGs, but I wonder if these data could provide insight into the other biological processes that might be controlled by this signaling pathway in T. brucei.
We published that the inositol phosphate pathway also plays a role in T. brucei development (Cestari et al. 2018, Mol Biol Cell; reviewed by Cestari I 2020, PLOS Pathogens)
Overall, this is an excellent study which represents an important step forward in understanding how antigenic variation is controlled in T. brucei. The possibility that this process could be controlled via a signalling pathway has been speculated for a long time, and this study provides the first mechanistic evidence for that possibility.
Reviewer #1 (Recommendations For The Authors):
Please see the public review for recommendations.1. It is critical to clearly state whether the observations are made for the endogenous WT protein or the tagged protein. It is good that the authors currently clearly indicate the results observed in vivo are for the RAP1-HA protein. However, this is not as clearly stated for in vitro EMSA analyses. In addition, in discussion, the authors simply assumed that the c-terminally tagged RAP1 behaves the same as WT RAP1 and all conclusions were made about WT RAP1.
There are two choices here. The authors can validate that RAP1-HA still retains RAP1's essential function as a sole allele in T. brucei cells (as was recommended previously). Indeed, HA-tagged RAP1 has been studied before, but it is the N-terminally HA-tagged RAP1 that has been shown to retain its essential functions. Adding the HA tag to the C-terminus of RAP1 may well cause certain defects to RAP1. For example, N-terminally HA-tagged TERT does not complement the telomere shortening phenotype in TERT null T. brucei cells, while C-terminally GFP-tagged TERT does, indicating that HA-TERT is not fully functional while TERT-GFP likely has its essential functions (Dreesen, RU thesis). Although RAP1-HA behaves similar to WT RAP1 in many ways, it is still not fully validated that this protein retains essential functions of RAP1. By the way, it has been published that cells lacking one allele of RAP1 behaves as WT cells for cell growth and VSG silencing (Yang et al. 2009, Cell; Afrin et al. 2020, mSphere). In addition, although RAP1 may interact with TRF weakly, the interaction is direct, as shown in yeast 2-hybrid analysis in (Yang et al. 2009, Cell).
Alternatively, if the authors do not wish to validate the functionality of RAP1-HA, they need to add one paragraph at the beginning of the discussion to clearly state that RAP1-HA may not behave exactly as WT RAP1. This is important for readers to better interpret the results. In addition, the authors need to tune down the current conclusions dramatically, as all described observations are made on RAP1-HA but not the WT RAP1.
The results with RAP1-HA are consistent with previous knowledge of RAP1 interactions with telomeric proteins and DNA. Hence, the C-terminal HA-tagged RAP1 seems, by all measures, functional. Nevertheless, to make it clear for the reader, we added a note in the discussion, lines 244-246: “Although we showed that C-terminal HA-tagged RAP1 protein has telomeric localization (Cestari et al. 2015, PNAS) and interactions with other telomeric proteins (Cestari et al. 2019 Mol Cell Biol); we cannot rule out potential differences between HA-tagged and non tagged RAP1.”
For a similar reason, the current EMSA results truly reflect how C-terminally His6-tagged RAP1 and RAP1 fragments behave. If the authors choose not to remove the His6 tag, it is essential that they use "RAP1-His6" to refer to these recombinant proteins. It is also essential for the authors to clearly state in the discussion that the tagged RAP1 fragments bind DNA, but the current data do not reveal whether WT RAP1 binds DNA. In addition, the authors incorrectly stated that "disruption of the MybL domain sequence did not eliminate RAP1-telomere binding in vivo" (lines 165-166). In ref 29, deletion of Myb domain did not abolish RAP1-telomere association. However, point mutations in MybL domain that abolish RAP1's DNA binding activities clearly disrupted RAP1's association with the telomere chromatin. Therefore, the current observation is not completely consistent with that published in ref 29.
We stated in lines 149-150 “…we expressed and purified from E. coli recombinant 6xHistagged T. brucei RAP1 (rRAP1)”. To clarify to the authors, we replaced rRAP1 with rRAP1-His throughout the manuscript text. As for the statement that “disruption of the MybL domain sequence did not eliminate RAP1telomere binding in vivo" (lines 165-166).”. We removed the statement from the manuscript.
2) There is no evidence, in vitro or in vivo, that binding PI(3,4,5)P3 to RAP1 causes conformational change in RAP1. The BRCT domain of RAP1 is known for its ability to homodimerize (Afrin et al. 2020, mSphere). It is therefore possible that binding of PI(3,4,5)P3 to RAP1 simply disrupts its homodimerization function. The authors clearly have extrapolated their conclusions based on available data. It is therefore important to revise the discussion and make appropriate statements.
We did not state that PI(3,4,5)P3 causes RAP1 conformational changes. We discussed the possibility. We stated in lines 199-201: “PI(3,4,5)P3 inhibition of RAP1-DNA binding might be due to its association with RAP1 N-terminus causing conformational changes that affect Myb and MybL domains association with DNA.” This is a reasonable discussion, given the data presented in the manuscript.
en idioma japonés de las palabras «Wa» (armonía o círculo) y «Komu» (computadora).
tag
Reviewer #2 (Public Review):
Casp11 is a cytosolic sensor for LPS in mice (orthologue of Casp4/5 in human). It is an important innate sensor of intracellular infection. Casp11 activity results in cleavage and activation of the pore-forming protein Gasdemin D (GSDMD) leading to lytic death (pyroptosis), of an infected cell. How exactly Casp11 signals upon LPS detection is beginning to be understood, but the picture is incomplete. Previous reports suggested that upon LPS detection, Casp11 dimerizes and undergoes auto-processing to form a pyroptosis-competent enzyme. The prediction from these studies was that the formation of a fully functional Casp11 signalling complex involves two steps: inducible dimerization and auto-processing.
In this study, authors used fluorescently tagged Casp11 reporter fusions, to report that detection of cytosolic LPS induces Casp11 assembly into a large perinuclear speck to form a signalling complex, where GSDMD can be processed. Such signalling complex resembles signalling specks formed upon the activation of other canonical inflammasomes.
Strengths:
Results are clean, experiments well controlled, and support the conclusions. Overall conclusions fit nicely in the general principle of innate signalling, whereby activation of many innate sensors results in their inducible assembly into higher-order oligomeric signalling complexes, called supra-molecular organizing centers (SMOCs).
A surprising finding from this work was that catalytically inactive Casp11 (C254A mutant) did not form signalling specks, despite being able to bind LPS and dimerise. This model is proposed where LPS binding to the CARD domain of Casp11 and Casp11 dimerization is necessary but not sufficient to mediate Casp11 speck formation within cells. The Casp11 catalytic activity is needed to facilitate the assembly of the higher-order, pyroptosis-competent Casp11 signalling platform. The model is further supported by experimental evidence that auto-processing of Casp11, by an exogenous protease TEV, (i.e. in the absence of LPS), is sufficient to mediate speck assembly in cells expressing wild type, but not catalytically inactive Casp11 mutant.
Possible technical improvements:
In general, the authors achieved their aims, and the results support the conclusions.
For technical robustness, it would be nice to consider a few controls:<br /> (a) Visualise Casp11 specks using constructs with smaller tags, and test whether tag placement on N or C terminus matters for speck formation; or<br /> (b) Biochemically crosslink and isolate endogenous, untagged Casp11 specks upon LPS transfection of primed macrophages (e.g. after priming through IFNs or TLRs). This would mimic the natural upregulation and activation of endogenous Casp11.<br /> (c) Test what happens after actual intracellular pathogen detection when the pathogen itself serves as a signalling platform? Are specks stills formed (or even needed)?
The broad impact of the work, implication, and questions for future work:
Results of this study would suggest that the enzymatic activity of Casp11 in macrophages may be highly restricted to the speck location, similar to what was described for Casp1. This may explain the very restricted substrate repertoire of Casp11 in cells, likely controlled by the substrate recruitment to the speck. This also opens avenues for follow-up work to answer several emerging questions:
1. After LPS binding and dimerization, why Casp11 must undergo intra-molecular processing to induce the formation of a pyroptosis-competent speck? Is there any substrate for LPS-bound, uncleaved Casp11 (beyond Casp11 itself), before Casp11 forms a full speck for GSDMD processing? The only currently known targets of Casp11 activity are itself, and GSDMD. Also, after intradomain linker cleavage of Casp11, what additional substrate must the cleaved Casp11 process to allow full speck formation?
2. Can activity probes be designed to detect the location of the active Casp11, and if so, would the activity of Casp11 be restricted to the speck? Is there a second cleavage event that would eventually dissociate Casp11 from the speck, to terminate its signalling? If not, how is speck activity terminated? If specks are released by lysis, are they capable of seeding new speck formation in neighbouring phagocytes, in prion-like behaviour previously described for canonical ASC speck?
3. What is the role of macrophage priming in speck formation, and what roles, if any GBPs play in speck formation?
4. Does this model apply to human orthologues, Casp4/5?
Reviewer #2 (Public Review):
Summary:
Chew et al describe interaction of the flavivirus protein NS1 with HDL using primarily cryoEM and mass spec. The NS1 was secreted from dengue virus infected Vero cells, and the HDL were derived from the 3% FBS in the culture media. NS1 is a virulence factor/toxin and is a biomarker for dengue infection in patients. The mechanisms of its various activities in the host are incompletely understood. NS1 has been seen in dimer, tetramer and hexamer forms. It is well established to interact with membrane surfaces, presumably through a hydrophobic surface of the dimer form, and the recombinant protein has been shown to bind HDL. In this study, cryoEM and crosslinking-mass spec are used to examine NS1 secreted from virus-infected cells, with the conclusion that the sNS1 is predominantly/exclusively HDL-associated through specific contacts with the ApoA1 protein.
Strengths:
The experimental results are consistent with previously published data.
Weaknesses:
CryoEM:
Some of the neg-stain 2D class averages for sNS1 in Fig S1 clearly show 1 or 2 NS1 dimers on the surface of a spherical object, presumably HDL, and indicate the possibility of high-quality cryoEM results. However, the cryoEM results are disappointing. The cryo 2D class averages and refined EM map in Fig S4 are of poor quality, indicating sub-optimal grid preparation or some other sample problem. Some of the FSC curves (2 in Fig S7 and 1 in Fig S6) have extremely peculiar shapes, suggesting something amiss in the map refinement. The sharp drop in the "corrected" FSC curves in Figs S5c and S6c (upper) indicate severe problems. The stated resolutions (3.42 & 3.82 Å) for the sNS1ts-Fab56.2 are wildly incompatible with the images of the refined maps in Figs 3 & S7. At those resolutions, clear secondary structural elements should be visible throughout the map. From the 2D averages and 3D maps shown in the figures this does not seem to be the case. Local resolution maps should be shown for each structure.
The samples were clearly challenging for cryoEM, leading to poor quality maps that were difficult to interpret. None of the figures are convincing that NS1, Ab56.2 or Fab56.2 are correctly fit into EM maps. There is no indication of ApoA1 helices. Details of the fit of models to density for key regions of the higher-resolution EM maps should be shown and the models should be deposited in the PDB. An example of modeling difficulty is clear in the sNS1ts dimer with bound Fab56.2 (figs 3c & S7e). For this complex, the orientation of the Fab56.2 relative to the sNS1ts dimer in this submission (Fig 3c) is substantially different than in the bioRxiv preprint (Fig 3c). Regions of empty density in Fig 3c also illustrate the challenge of building a model into this map.
Mass spec:
Crosslinking-mass spec was used to detect contacts between NS1 and ApoA1, providing strong validation of the sNS1-HDL association. As the crosslinks were detected in a bulk sample, they show that NS1 is near ApoA1 in many/most HDL particles, but they do not indicate a specific protein-protein complex. Thus, the data do not support the model of an NS1-ApoA1 complex in Fig 4d. Further, a specific NS1-ApoA1 interaction should have evidence in the EM maps (helical density for ApoA1), but none is shown or mentioned. If such exists, it could perhaps be visualized after focused refinement of the map for sNS1ts-HDL with Fab56.2 (Fig S7d). The finding that sNS1-ApoA1 crosslinks involved residues on the hydrophobic surface of the NS1 dimer confirms previous data that this NS1 surface engages with membranes and lipids.
Sample quality:
The paper lacks any validation that the purified sNS1 retains established functions, for example the ability to enhance virus infectivity or to promote endothelial dysfunction. Peculiarities include the gel filtration profiles (Fig 2a), which indicate identical elution volumes (apparent MWs) for sNS1wt-HDL bound to Ab562 (~150 kDa) and to the ~3X smaller Fab56.2 (~50 kDa). There should also be some indication of sNS1wt-HDL pairs crosslinked by the full-length Ab, as can be seen in the raw cryoEM micrograph (Fig S5b).
Obtaining high quality structures is often more demanding of sample integrity than are activity assays. Given the low quality of the cryoEM maps, it's possible that the acidification step in immunoaffinity purification damaged the HDL complex. No validation of HDL integrity, for example with acid-treated HDL, is reported. Acid treatment is perhaps discounted by a statement (line 464) that another group also used immunoaffinity purification in a recent study (ref 20) reporting sNS1 bound to HDL. However the statement is incorrect; the cited study used affinity purification via a strep-tag on recombinant sNS1.
Discussion:
The Discussion reflects a view that the NS1 secreted from virus-infected cells is a 1:1 sNS1dimer:HDL complex with the specific NS1-ApoA1 contacts detected by crosslinking mass spec. This is inconsistent with both the neg-stain 2D class average with 2 sNS1 dimers on an HDL (Fig S1c) and with the recent study of Flamand & co-workers showing 1-3 NS1 dimers per HDL (ref 20). It is also ignores the propensity of NS1 to associate with membranes and lipids. It is far more likely that NS1 association with HDL is driven by these hydrophobic interactions than by specific protein-protein contacts. A lengthy Discussion section (lines 461-522) includes several chemically dubious or inconsistent statements, all based on the assumption that specific ApoA1 contacts are essential to NS1 association with HDL and that sNS1 oligomers higher than the dimer necessarily involve ApoA1 interaction, conclusions that are not established by the data in this paper.
Analog zettelkasten for natural sciences .t3_17kui2u._2FCtq-QzlfuN-SwVMUZMM3 { --postTitle-VisitedLinkColor: #9b9b9b; --postTitleLink-VisitedLinkColor: #9b9b9b; --postBodyLink-VisitedLinkColor: #989898; }
Reply to u/Wooden-School-4091 at https://www.reddit.com/r/Zettelkasten/comments/17kui2u/analog_zettelkasten_for_natural_sciences/
Given that Carl Linnaeus "invented" the standardized 3x5 inch index card and used it heavily in his scientific work (read Isabelle Charmantier and Staffan Müller-Wille's works for more on his practice), and a variety of others including me, use it for mathematics, physics, chemistry, biology, etc., Zettelkasten can certainly be used for STEM, STEAM, and any of the natural sciences.
See also, notes and links at: https://hypothes.is/users/chrisaldrich?q=tag%3A%22zettelkasten+for+studying%22
If I were using it for classes/university/general studying via lectures, I'd base my practice primarily on Cornell Notes in combination with creating questions/cards for spaced repetition and/or a variation on Leitner's System.
Some of the best material on spaced repetition these days can be found via:
and other material on their sites.
Beyond this, I'd focus my direct zettelkasten practice less on the learning portion and more on the developing or generating ideas portion of the work. Some of my practice with respect to mathematics can be found here: https://www.reddit.com/r/Zettelkasten/comments/17bqztm/applying_zettelkasten_for_math_heavy_subjects/
For those interested, it may bear mentioning that Bjornstad, an engineer at Remnote, has a TiddlyWiki-based zettelkasten at https://zettelkasten.sorenbjornstad.com/#PublicHomepage:PublicHomepage which he demonstrates with a walk through at https://www.youtube.com/watch?v=GjpjE5pMZMI
Joint Public Review:
In this manuscript, Kipfer et al describe a method for a fast and accurate SARS-CoV2 rescue and mutagenesis. This work is based on a published method termed ISA (infectious subgenomic amplicons), in which partially overlapping DNA fragments covering the entire viral genome and additional 5' and 3' sequences are transfected into mammalian cell lines. These DNA fragments recombine in the cells, express the full length viral genomic RNA and launch replication and rescue of infectious virus.
CLEVER, the method described here significantly improves on the ISA method to generate infectious SARS-CoV2, making it widely useful to the virology community.
Specifically, the strengths of this method are:<br /> 1) The successful use of various cell lines and transfection methods.<br /> 2) Generation of a four-fragment system, which significantly improves the method efficiency due to lower number of required recombination events.<br /> 3) Flexibility in choice of overlapping sequences, making this system more versatile.<br /> 4) The authors demonstrated how this system can be used to introduce point mutations as well as insertion of a tag and deletion of a viral gene.<br /> 5) Fast-tracking generation of infectious virus directly from RNA of clinical isolates by RT-PCR, without the need for cloning the fragments or using synthetic sequences.<br /> 6) The authors further expanded this method to work on additional plus-strand RNA viruses beyond SARS-CoV-2 (CHIKV, DENV)
The manuscript clearly presents the findings, and the proof-of-concept experiments are well designed.
Overall, this is a very useful method for SARS-CoV2 research. Importantly, it can be applicable to many other viruses, speeding up the response to newly emerging viruses than threaten the public health.
Reviewer #1 (Public Review):
Summary:<br /> In this manuscript, the authors have applied an asymmetric split mNeonGreen2 (mNG2) system to human iPSCs. Integrating a constitutively expressed long fragment of mNG2 at the AAVS1 locus, allows other proteins to be tagged through the use of available ssODN donors. This removes the need to generate long AAV donors for tagging, thus greatly facilitating high-throughput tagging efforts. The authors then demonstrate the feasibility of the method by successfully tagging 9 markers expressed in iPSC at various, and one expressed upon endoderm differentiation. Several additional differentiation markers were also successfully tagged but not subsequently tested for expression/visibility. As one might expect for high-throughput tagging, a few proteins, while successfully tagged at the genomic level, failed to be visible. Finally, to demonstrate the utility of the tagged cells, the authors isolated clones with genes relevant to cytokinesis tagged, and together with an AI to enhance signal-to-noise ratios, monitored their localization over cell division.
Strengths:<br /> Characterization of the mNG2 tagged parental iPSC line was well and carefully done including validation of a single integration, the presence of markers for continued pluripotency, selected off-target analysis, and G-banding-based structural rearrangement detection.
The ability to tag proteins with simple ssODNs in iPSC capable of multi-lineage differentiation will undoubtedly be useful for localization tracking and reporter line generation.
Validation of clone genotypes was carefully performed and highlights the continued need for caution with regard to editing outcomes.
Weaknesses:<br /> IF and flow cytometry figures lack quantification and information on replication. How consistent is the brightness and localization of the markers? How representative are the specific images? Stability is mentioned in the text but data on the stability of expression/brightness is not shown.
The localization of markers, while consistent with expectations, is not validated by a second technique such as antibody staining, and in many cases not even with Hoechst to show nuclear vs cytoplasmic.
For the multi-germ layer differentiation validation, NCAM is also expressed by ectoderm, so isn't a good solo marker for mesoderm as it was used. Indeed, the kit used for the differentiation suggests Brachyury combined with either NCAM or CXCR4, not NCAM alone.
Only a single female parental line has been generated and characterized. It would have been useful to have several lines and both male and female to allow sex differences to be explored.
The AI-based signal-to-noise enhancement needs more details and testing. Such models can introduce strong assumptions and thus artefacts into the resolved data. Was the model trained on all markers or were multiple models trained on a single marker each? For example, if trained to enhance a single marker (or co-localized group of markers), it could introduce artefacts where it forces signal localization to those areas even for others. What happens if you feed in images with scrambled pixel locations, does it still say the structures are where the training data says they should be? What about markers with different localization from the training set? If you feed those in, does it force them to the location expected by the training data or does it retain their differential true localization and simply enhance the signal?
Reviewer #3 (Public Review):
The authors report on the engineering of an induced Pluripotent Stem Cell (iPSC) line that harbours a single copy of a split mNeonGreen, mNG2(1-10). This cell line is subsequently used to take endogenous protein with a smaller part of mNeonGreen, mNG2(11), enabling the complementation of mNG into a fluorescent protein that is then used to visualize the protein. The parental cell is validated and used to construct several iPSC lines with endogenously tagged proteins. These are used to visualize and quantify endogenous protein localisation during mitosis.
I see the advantage of tagging endogenous loci with small fragments, but the complementation strategy has disadvantages that deserve some attention. One potential issue is the level of the mNG2(1-10). Is it clear that the current level is saturating? Based on the data in Figure S3, the expression levels and fluorescence intensity levels show a similar dose-dependency which is reassuring, but not definitive proof that all the mNG2(11)-tagged protein is detected.
Do the authors see a difference in fluorescence intensity for homo- and heterozygous cell lines that have the same protein tagged with mNG2(11)? One would expect two-fold differences, or not?
Related to this, would it be favourable to have a homozygous line for expressing mNG2(1-10)?
The complementation seems to work well for the proteins that are tested. Would this also work for secreted (or other organelle-resident) proteins, for which the mNG2(11) tag is localised in a membrane-enclosed compartment?
The authors present a technological advance and it would be great if others could benefit from this as well by having access to the cell lines.
eLife assessment
This study describes a method to track MHC class II binding peptides on dendritic cell (DC) surfaces using a tetracystein tag and a thiol-reactive dye, which can then be investigated in vitro and in vivo. This is a valuable study for the impact on immunology and potentially other areas where the detection of cell-associated peptides is required. The methods are convincing based on the use of MHC class I/II deficient mice that have significantly reduced signal, but the non-zero background is detected, and it is not clear that this is lower than if the peptides were directly labelled with fluorophores.
Reviewer #1 (Public Review):
Summary:<br /> The authors develop a method to fluorescently tag peptides loaded onto dendritic cells using a two-step method with a tetracystein motif modified peptide and labelling step done on the surface of live DC using a dye with high affinity for the added motif. The results are convincing in demonstrating in vitro and in vivo T cell activation and efficient label transfer to specific T cells in vivo. The label transfer technique will be useful to identify T cells that have recognised a DC presenting a specific peptide antigen to allow the isolation of the T cell and cloning of its TCR subunits, for example. It may also be useful as a general assay for in vitro or in vivo T-DC communication that can allow the detection of genetic or chemical modulators.
Strengths:<br /> The study includes both in vitro and in vivo analysis including flow cytometry and two-photon laser scanning microscopy. The results are convincing and the level of T cell labelling with the fluorescent pMHC is surprisingly robust and suggests that the approach is potentially revealing something about fundamental mechanisms beyond the state of the art.
Weaknesses:<br /> The method is demonstrated only at high pMHC density and it is not clear if it can operate at at lower peptide doses where T cells normally operate. However, this doesn't limit the utility of the method for applications where the peptide of interest is known. It's not clear to me how it could be used to de-orphan known TCR and this should be explained if they want to claim this as an application. Previous methods based on biotin-streptavidin and phycoerythrin had single pMHC sensitivity, but there were limitations to the PE-based probe so the use of organic dyes could offer advantages.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
The study performed by Niphadkar et al. seeks to uncover the role of the phosphatase Ppg1 in regulating gluconeogenesis during post-diauxic shift in S. cerevisiae. Thea authors show that loss or inactivation of Ppg1p affects production of gluconeogenic products incl. trehaloase and glycogen. The authors show that assembly of the Far complex required the activity of Ppg1 and is required to maintain gluconeogenic outputs after glucose depletion.
The manuscript is clearly written and methods well considered, no omics-methods have been included. Especially phosphoproteomics would be relevant to include. Specifically, the tracing experiments are an interesting and appropriate approach to confirm effects on gluconeogeneisis etc. Yet, working with regulation of posttranslational modifications (phosporylations) it is surprising that the authors only to a limited extent examine phosphorylation events, and not all examine or discuss specific phosphorylation events of e.g. Far11.
The study is interesting and provides new insights into regulation of glucose metabolism in yeast, however, there are serious concerns that need to be addressed before it can be reconsidered for publication.
Major points:
The authors use electrophoretic mobility assays w/wo CIP to address the phosphorylation state of Far11. They show in figure 3E that the mobility of Far11 depends on Ppg1 activity and can be affected by CIP. Why is the mobility of Far11 not affected in e.g. figure 3D?
There are several sites in Far11 previously reported to be phosphorylated, see e.g. Bodenmiller et al 2010 (Science Signal.) Are there sites that are specifically regulated (dephosphorylated) by Ppg1? or by other phosphatases? kinases?
Here, it would be appropriate to apply phosphoproteomics to examine Far11 phosphorylation in Ppg1 knock out cells or in cells with inactivated Ppg1.
The authors show that the levels of Ppg1 remain constant during growth in YPD medium, while the levels of Far11 increased after 24hrs of growth in YPD medium, and thus argue that the amount of Far complex itself increases in post-diauxic phase. The authors need to show that the level of complex indeed increases.
The authors also apply fluorescence microscopy to address the localization of the Far11 complex etc. The quality of the shown images should be improved, also merged images should be shown. Only one single image containing one cell is shown, images should ideally show additional cells in the same image, alternatively, additional images should be shown.
Minor points:
Does the FLAG tag affect activity of Ppg1?
The study is interesting and provides new insights into regulation of glucose metabolism in yeast, however, there are serious concerns that need to be addressed before it can be reconsidered for publication.
The manuscript is of broad interests for an audience primarily interested glucose metabolism and signalling in yeast.
cops n’ robbers
The name's that kids gave to their games back then were insane. Why was this such a fun game to play though? in Mexico tag was called* la roña * which was a name for chicken pox
39:28 - Note graph and tag discussion
Obsidian for Academic Publishing - A Walkthrough with Jason Yuh (6)
Jason menciona que su forma de investigación está hecha a partir de capítulos y no de ideas.
Anthony comenta sobre "serendipity": encontrar conexiones que no encontraría por sí mismo. No las encuentra buscando "serendipity connections", sino cuando está buscando otra cosa siguiendo links. Y de repente aparece, cuando está haciendo links con otras notas.
Jason usa tags como keywords. Pero eventualmente, cuando el concepto crece, hace una nota: ej. "Innovation".
Tiene secciones de:
Importante hacer esto, con mis notas de cada concepto. O quizás con una nota más extensa para trabajar en varios conceptos que se relacionan.
Jason Yuh is a Ph.D. student at the University of Toronto working on his dissertation entitled: "The Dialectics of Traditions and Innovations in the Didache through Canon and Ritual." In this session Jason shares how he uses Obsidian to organize and process notes for developing his thesis. 00:00 - Intro, Obsidian discovery, and Jason's thesis 16:08 - Jason screen sharing 17:55 - Folder structure 20:30 - Tracking scholar references 23:40 - Daily notes 29:05 - Taking notes on multi-author volumes 32:00 - Writing abstracts (and using transclusion) 38:16 - Reading vs. Writing discussion 39:28 - Note graph and tag discussion 47:32 - Deciding what to work on next 49:27 - Synthesizing notes for publication 1:01:16 - Wrap-up
Obsidian for Academic Publishing - A Walkthrough with Jason Yuh (0)
Anthony's Desk
1:02:33
HTML had blown open document publishing on the internet
... which may have really happened, per se, but it didn't wholly incorporate (subsume/cannibalize) conventional desktop publishing, which is still in 2023 dominated by office suites (a la MS Word) or (perversely) browser-based facsimiles like Google Docs. Because the Web as it came to be used turned out to be as a sui generis medium, not exactly what TBL was aiming for, which was giving everything (everything—including every existing thing) its own URL.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
This manuscript presents the cis-regulatory analysis of the enhancers controlling prox1a gene in zebrafish. Authors used both evolutionary conservation and existing single-cell ATAC data to highlight the major role of two elements. I feel that the transgenesis work is quite solid and the main conclusions interesting. However, I feel the authors need to provide some extra validations for some of the analysis.
I also feel that the evolutionary aspect could be discussed a bit more, what are the differences between the diffeerent vertebrate lineage, etc...
(p7) active enhancer in a tissue: while ATAC gives a good indicated of accessibility it is not an indicate of activity as for instance H3K27Ac would be.
I think this is an interesting piece of work, which elaborates on previous studies on prox1a involvment in the lymphatic system but it doesn not bring essentially new perspective on the question.
RRID:SCR_022636
DOI: 10.1101/2023.10.16.562573
Resource: SCR_022636
Curator: @scibot
SciCrunch record: RRID:SCR_022636
Note: This rebuttal was posted by the corresponding author to Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
[The “revision plan” should delineate the revisions that authors intend to carry out in response to the points raised by the referees. It also provides the authors with the opportunity to explain their view of the paper and of the referee reports.
In this paper we describe the new finding that the epicardial deposits the extracellular matrix component laminin onto the apical ventricular surface during cardiac development. We identify a novel role for the apicobasal polarity protein Llgl1in timely emergence of the epicardium and deposition of this apical laminin, alongside a requirement for Llgl1 in maintaining integrity of the ventricular wall at the onset of trabeculation.
We thank the reviewers for their very positive appraisal of our manuscript, and for their helpful suggestions for useful revisions. In particular we would like to highlight the broad interest they feel this manuscript holds, not only contributing conceptual advances to our understanding of multiple aspects of cardiac development, but also to cell and developmental biologists working in epithelial polarity and extracellular matrix function. We also note their positive appraisal of the rigor of the study and quality of the manuscript.
Reviewer 1
1a) It is mentioned that llgl1 CRISPR/Cas9 mutants are viable as adults on pg. 3 of the Results section. Have the authors examined heart morphology in these mutants in juvenile or adult fish?
We have some historical data on adult llgl1 mutant survival that we plan to include in the study.
Reviewer 2
2a) The authors note an interesting observation with apical and basal laminin deposition dynamics surrounding cardiomyocytes, and that Llg1 has a role in apical Laminin deposition (however, highly variable at 80 hpf as Figure 3M shows). They carry out a very nice study in which they overexpress Llgl1 tagged with mCherry in the myocardium and show that there is no rescue of the extruding cardiomyocyte defect or Laminin deposition. However, there is still a possibility that the tagged Llgl1 in the transgene Tg(myl7:Llg1-mCherry)sh679 might not be functional due to improper protein folding or interference by the mCherry tag. The authors should supplement their approach with a transplantation experiment to generate mosaic llgl1 mutant animals and assess whether llgl1 mutant cardiomyocytes extrude at a higher rate than the control. This would provide definitive evidence that Llg1l acts in a cell non-autonomous manner.
We agree with the reviewer, and propose to perform transplant experiments, transplanting cells from llgl1 mutants into wild type siblings, and quantify cell extrusion to determine whether llgl1 mutant cells are extruded more frequently than wild type.
2b) The data in this manuscript appears to point that Llgl1 regulates Laminin deposition mainly in epicardial cells to regulate their dissemination/migration across the ventricular myocardial surface. It would be important to test this cell-autonomous function with the transplant experiment (above point) and examine whether llgl1 mutant epicardial cells fail to migrate and deposit Laminin. It might be possible to perform a rescue experiment through overexpression of Llgl1 in epicardial cells (if possible, there is a tcf21:Gal4 line available).
Similar to above, we propose to perform transplant experiments, transplanting cells from llgl1 mutants or wild type siblings into wild type siblings or llgl1 mutants, respectively, and in this instance quantify contribution of transplanted cells to epicardial coverage.
2c) In the Discussion, the authors propose that Llgl1 acts in two ways: Laminin deposition in epicardial cells that suppress cell extrusion and polarity regulation in cardiomyocytes to promote trabeculation. It would be important to test the second hypothesis on trabeculation and polarity regulation by using the myocardial-specific overexpression/rescue of Llgl1 in llgl1 mutants, and then quantifying the trabeculating cardiomyocytes and analyze Crb2a localization. This experiment can distinguish whether this trabeculation phenotype is rescued independently of the apical Laminin deposition that has been included in Figure S5.
To help address the second part of our hypothesis laid out in the discussion, we propose to quantify trabecular organisation and Crb2a localisation in llgl1 mutants either carrying the myl7:llgl1-mCherry construct, or mCherry-negative controls.
2d) The potential mis-localization of Crb2a in the llgl1 mutants is interesting, but this effect appears to be quite mild, and as the authors note, resolve by 80 hpf. Considering the role of Lgl in Drosophila in shifting Crb complex localization during early epithelial morphogenesis, it would be worth performing the analysis at earlier timepoints (between 55 and 72 hpf) to determine whether Llgl1 is indeed important for the progressive apical relocalization of Crb2a.
We will expand our description of this in the mutants by performing analysis of Crb2a at earlier timepoints in the llgl1 mutant (55hpf and 60hpf).
2e) OPTIONAL: It might be worth testing other antibodies that could mark the apical (particularly aPKC which is known to phosphorylate and regulate the Crb complex) and basolateral domains (Par1, Dlg) of the cardiomyocytes to definitively conclude that the epithelial integrity of the cells is affected. Although there are no reports of working antibodies marking the basal domain in zebrafish, there is at least a Tg(myl7:MARCK3A-RFP) line published (Jimenez-Amilburu et al. (2016)) - which the authors can inject to examine the localization in mosaic hearts.
We plan to assess localisation of aPKC (see section 4 for response to other suggested polarity protein analyses).
2f) Have the authors quantified the numbers of total cardiomyocytes in llgl1 mutants to correlate how many cells are lost as a consequence of extrusion? What is the physiological impact of this extrusion (ejection fraction, total cardiac volumes, sarcomere organization)?
We have some of this data already which we will include in the manuscript (cell number, myocardial volume). We agree that the analysis of cardiac function could be more extensive, and we will perform more detailed analysis of cardiac function, including e.g. ejection fraction. Sarcomere organisation has been previously described in llgl1 mutants by Flinn et al, 2020, so we do not plan to replicate this data.
2g) The lamb1a and lamc1 mutant phenotypes were nicely analyzed. However, there is basement membrane deposition on both the apical and basal sides of the cardiomyocytes. Therefore, it is unclear whether the cardiomyocyte extrusion is completely caused by loss of apical basement membrane, or whether the loss of basal basement membrane could compromise the myocardial tissue integrity. The authors should clarify this conclusion in the text.
We will address this further in the text, but will also include 55hpf Laminin staining data for llgl1 mutants to reinforce our message.
2h) The authors note that Llgl1-mCherry in the Tg(myl7:Llg1-mCherry)sh679 line localizes to the basolateral domain of the cardiomyocytes, which is valuable confirmation that Llgl1 protein is spatially restricted. However, only 1 timepoint (55 hpf) is noted. It would be important to perform Llgl1 localization across different developmental timepoints (at least until 80 hpf) to examine the dynamics of this protein during trabeculation and apical extrusion, and potentially correlate it with Crb2a localization for a better understanding of the apicobasal machinery in cardiomyocytes.
We already have some of this data and will include extra timepoints in a revised version of the manuscript
2i) The phenotypes of llgl1 mutants described here differ compared to the previous study by Flinn et al. (2020). In particular, whereas the mutants generated in this study have only mild pericardial edema and are adult viable, approximately one third of llgl1mw3 (Flinn et al. (2020)) died at 6 dpf. Is this caused by the different natures of the mutations in the llgl1 gene? Is there a possibility that the llgl1sh598 is a hypomorphic allele since the targeted deletion is in a more downstream sequence (in exon 2) compared to the llgl1mw3 (deletion in exon 1) allele?
We thank the reviewer for noticing these subtle differences between the two llgl1 mutants. Indeed, while we occasionally see llgl1sh598 mutants with the severe phenotype described by Flinn et al, this is a small minority which we did not quantify. Our mutation is indeed slightly further downstream than that described by Flinn et al, however we believe that this will have a neglible effect on Llgl1 function. Our llgl1sh589 mutation results in truncation shortly into the WD40 domain, and importantly completely lacks the Lgl-like domain, which is responsible for the specific function of Llgl1 likely through its ability to interact with SNAREs to regulate cargo delivery to membranes (Gangar et al, Current Biology 2005).
Interestingly, Flinn et al report no increased phenotypic severity in their maternal-zygotic llgl1 mutants when compared to zygotic mutants. Conversely, we often observed very severe phenotypes in MZ llgl1sh589 mutants, including failure of embryos during blastula stages, apparently through poor blastula integrity. We did not include this information in the manuscript due to space constraints. However, we argue that together these differences between the two alleles may not be due to hypomorphism of our llgl1sh589 allele, but rather differences in genetic background that may amplify specific phenotypes. We plan to include a short sentence summarising the above in combination with planned experiments described below to address the reviewer’s next comment.
2j) Suggested experiment: qPCR of regions downstream of the deletion to make sure that the transcript is absent/reduced in the llgl1sh598 mutants. Alternatively, immunostaining or Western blot would be an even better option to ensure there is no Llgl1 protein production - there is an anti-Llgl1 antibody available that works for Western blots in zebrafish (Clark et al. (2012)).
We plan to analyse llgl1 expression in llgl1 mutants using qPCR.
Reviewer 3
3a) Major - the authors describe that llgl1 mutants exhibit transient cardiac edema at 3 dpf, which is resolved by 5 dpf, and claim that the mutants are viable. This statement needs to be better supported - What is the proportion of mutants that survive to adulthood? The embryonic phenotypes are pretty variable - are the mutants that survive the ones with a less severe phenotype? Is there a gross defect in the adult heart of these animals?
In line with comments from Reviewers 1 and 2 above, we will include a description of the data we have from adult animals (historical data, not generation of new animals).
3b) Major - Many of the phenotypes described here -most importantly, the defects on epicardial development- could result from hemodynamic defects in llgl1 mutants. The authors claim that function is unaffected in these animals, but this has only been addressed by measuring heartbeat. The observation that the cardiac function in these animals is normal would conflict with a previous description (PMID: 32843528) that demonstrates that llgl1 mutant animals show significant hemodynamic defects, which would cause epicardial defects. Thus, this aspect of the work needs to be better addressed.
In line with our comments to point 2f) from Reviewer 2, we will perform a more in-depth functional analysis on llgl1 mutant larvae.
3c) The phenotypes related to forming multiple layers in the heart (Fig. 1) could be more convincing. In some figures, the authors use a reporter that labels the myocardial cell membrane, but in Figure 1 this is not used. Showing a myocardial membrane marker (for example, the antibody Alcama, Zn-8) would significantly strengthen this observation.
We will describe trabecular phenotypes in more detail using the suggested antibody to highlight membranes.
3d) The analysis of Crumbs redistribution (Fig. 2) is quite interesting. Still, given that the authors have a transgenic model to rescue llgl1 expression in cardiomyocytes, they could move from correlative evidence to experimental demonstration of the role of llgl1 in Crumbs localization.
Similar to our response to comment 2c) from Reviewer 2, we plan to address this
Reviewer 1:
Although information is provided in the introduction and discussion on the role of the Llgl1 homolog in Drosophila and speculation on LLGL1 contributing to heart defects in SMS patients in the discussion, have Llgl1 homologs been examined in other vertebrate animal models during heart development or regeneration?
With the exception of the Flinn et al paper, we find no published studies assessing the role of Llgl1 in heart development or regeneration in other vertebrates, and have updated the introduction to highlight this fact:
‘Zebrafish have two Lgl homologues, llgl1 and llgl2, and llgl1 has previously been shown to be required for early stages of heart morphogenesis (Flinn et al. 2020). However, although Llgl1 expression has also been reported in the developing mouse heart and both adult mouse and human hearts (Uhlén et al. 2015; Klezovitch et al. 2004), whether llgl1 plays a role in ventricular wall development has not been examined.’
In Fig. 4J-M', there is no Cav1 signals after wt1a MO but still laminin signals. Where these laminins come from?
The residual laminin staining observed in wt1a morphants is located at the basal surface of cardiomyocytes (while the apical laminin signal is lost, in line with the epicardial deposition of laminin at the apical ventricle surface). This basal laminin is likely deposited earlier during heart tube development by either the myocardium, endocardium or both, and thus unaffected by later formation of the epicardium. We reason this since a) it is present at the basal cardiomyocyte surface at 55hpf (see Fig 2); b) we have previously identified both myocardial and endocardial expression of laminin subunits at 26hpf and 55hpf (Derrick et al, Development, 2021); c) sc-RNA-seq analysis of hearts at 48hpf demonstrates that laminin subunits, e.g. lamc1 are expressed in myocardial and endocardial cells (Nahia et al, bioRxiv, 2023), also in line with our previous ISH analysis. We have included a sentence to reflect this in the results section:
Conversely, *wt1a* morphants retain deposition of laminin at the basal CM surface, likely from earlier expression and deposition of laminin by either myocardial or endocardial cells (Derrick et al. 2021; Nahia et al. 2023), which is unaffected by later epicardial development.
On page 3 of the manuscript, Fig. 1A should be included with Fig. 1B in the first sentence of paragraph 2 of the Results subsection "Llgl1 regulates ventricular wall integrity and trabeculation".
Amended
It would be beneficial to readers to briefly describe what cell type the transgenic reporters label when mentioned in the Results section to help readers unfamiliar with zebrafish.
We have updated the text to read:
We further analysed heart morphology using live lightsheet microscopy of *Tg(myl7:LifeActGFP);Tg(fli1a:AC-TagRFP)* double transgenic wild-type and *llgl1* mutant embryos, allowing visualisation of myocardium (green) and endocardium (magenta) respectively. Comparative analysis of overall heart morphology between 55hpf and 120hpf when looping morphogenesis is complete, revealing that *llgl1* mutants continue to exhibit defects in heart morphogenesis (Fig S1S-X).
Reviewer 3
(Optional) There is laminin in the luminal side of the heart before there is any epicardial invasion. What is the source of this laminin? The techniques the authors have used (i.e., chromogenic ISH) are fine, but a more detailed analysis using fluorescent ISH (i.e., RNAScope) would be much more definitive.
This is related to our response to Reviewer 1 (above) – where we have included the following text included in manuscript: Conversely, *wt1a* morphants retain deposition of laminin at the basal CM surface, likely from earlier expression and deposition of laminin by either myocardial or endocardial cells (Derrick et al. 2021; Nahia et al. 2023), which is unaffected by later epicardial development. We hope this clarifies our proposed origins for the earlier laminin deposition.
Reviewer 1:
As pan-epicardial transgenes like tcf21 reporters have been widely used, the authors should use such reporters to verify the expression of laminin gene expression in epicardial cells, and the efficacy and efficiency of depleting epicardial cells after wt1 MO injection.
Several studies have demonstrated that the epicardium is not a heterogeneous population – for example, tcf21 is not expressed in all epicardial cells and thus not a pan-epicardial reporter (Plavicki et al, BMC Dev Biol, 2014, Weinberger et al, Dev Cell, 2020) – the suggested analysis would not necessarily be conclusive, and more detailed study would require acquisition of three new transgenic lines. Furthermore, we believe the evidence we present in the paper supports our claim: 1) We show expression of two laminin subunits in a thin mesothelial layer directly adjacent to the myocardium, specifically in the location of the epicardium; 2) sc-RNA seq analyses have also identified laminin expression in epicardial cells at 72hpf (where lamc1a is identified as a marker of the epicardium); 3) We demonstrate 100% efficacy of our wt1a knockdown as assayed by Cav1 expression, an established epicardial marker (Grivas et al, 2020, Marques et al, 2022) which in sc-RNA seq data is expressed at high levels broadly in the epicardial cell population (Nahia et al, 2023), representing a good assay for presence of epicardium. However, we propose to perform ISH analysis of laminin subunit expression in wt1a MO to investigate whether the mesothelial laminin-expressing layer we observe adjacent to the myocardium is absent upon loss of wt1a.
Reviewer 2:
The data in this manuscript appears to point that Llgl1 regulates Laminin deposition mainly in epicardial cells to regulate their dissemination/migration across the ventricular myocardial surface. It would be important to test this cell-autonomous function with the transplant experiment (above point) and examine whether llgl1 mutant epicardial cells fail to migrate and deposit Laminin. It might be possible to perform a rescue experiment through overexpression of Llgl1 in epicardial cells (if possible, there is a tcf21:Gal4 line available).
We do not propose to perform this experiment using a tcf21:Gal4 line, as this would likely require at least 6 months to either import and quarantine, or generate the necessary stable lines. Furthermore, as mentioned above, tcf21 is not a pan-epicardial marker, and the extent and timing of the Gal4:UAS system may make this challenging to determine whether llgl1 has been expressed early or broadly enough. We will instead attempt transplantation experiments.
OPTIONAL: It might be worth testing other antibodies that could mark the apical (particularly aPKC which is known to phosphorylate and regulate the Crb complex) and basolateral domains (Par1, Dlg) of the cardiomyocytes to definitively conclude that the epithelial integrity of the cells is affected. Although there are no reports of working antibodies marking the basal domain in zebrafish, there is at least a Tg(myl7:MARCK3A-RFP) line published (Jimenez-Amilburu et al. (2016)) - which the authors can inject to examine the localization in mosaic hearts.
We will assess localisation of aPKC, but we do not plan to analyse the other components. Analysis of basolateral domains (Par1, Dlg, Mark3a-RGP), will not necessarily assess epithelial integrity, as suggested, but rather apicobasal polarity – which we already assess using Crb2a, and additionally plan to assess aPKC to accompany the Crb2a analysis. Since the reviewer suggests this as an optional experiment we prioritise their other suggested experiments that we think more directly address the main messages of the manuscript.
OPTIONAL: Gentile et al. (2021) found that reducing heartbeat led to decreased cardiomyocyte extrusion in snai1b mutants. The authors could look into the contribution of mechanical pressure through contraction in the apical cardiomyocyte extrusion, and test whether reducing contraction (tnnt2 morpholino, chemical treatments) partly rescues the llgl1 mutant phenotypes.
The relationship between cardiac function and myocardial wall integrity appears to be complex. The paper referred to by the reviewer indeed finds that reduction in heartbeat leads to decreased CM extrusion upon loss of the EMT-factor Snai1b. Previous studies have also found that endothelial flow-responsive genes klf2a/b are required to maintain myocardial ventricular wall integrity at later stages in a contractility-dependent manner (Rasouli et al, 2018). However, contractility is also required early for pro-epicardial emergence, but plays a lesser role in expansion of the epicardial layer on the myocardial surface (Peralta, 2013). Unpicking the relationship between the forces induced by mechanical contraction of the ventricular wall, contractility-based induction of e.g klf2 expression, and the impact of contractile forces on proepicardial development or epicardial expansion will be complex. We therefore think the proposed experiment will be difficult to interpret whatever the outcome, and argue that dissecting this relationship is beyond the scope of revisions for this paper.
Reviewer 3
How llgl1 relates to epicardial biology is left entirely unexplored in this work. Do proepicardial cells show any defect in cell polarization related to llgl1 absence?
We agree with the reviewer that we do not delve into the mechanisms underlying regulation of epicardial development by llgl1, and that this is an interesting question. Our scope for this manuscript was to understand the mechanisms by which llgl1 regulates integrity of the ventricular wall, and feel that uncovering the molecular mechanisms by which llgl1 regulates timely epicardial emergence is a larger question that would require substantial investigation (for example, if and when llgl1 PE cells do exhibit apicobasal defects, how this impacts timing of cluster release etc). We think these are important questions that would be better answered in detail in a separate manuscript.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary: The manuscript by Pollitt et al. explores the functions of llgl1, which encodes a critical component of the basolateral domain complex, during cardiac development in zebrafish. The authors observed that llgl1 mutants exhibited compromised myocardial tissue integrity with significantly higher numbers of apically extruding cardiomyocytes. Llgl1 appears to primarily function during epicardial cell spreading on the myocardial tissue, as myocardial-specific overexpression of llgl1 did not rescue llgl1 mutant phenotypes. llgl1 mutants exhibited impaired epicardial coverage and subsequently Laminin deposition on the apical side of the cardiomyocytes. Functional linkage between Laminin/the basement membrane was identified, as extruding cardiomyocytes were also observed in mutants of two core laminin genes, lamb1a and lamc1. The epicardial defects were transmitted to myocardial tissue defects, marked by mis-localization of the apical polarity protein Crumbs2a during early heart development. Overall, the authors provide a nice study that strengthens the role of apicobasal factors in myocardial tissue morphogenesis and that sheds light on the role of epicardial-derived basement membrane in maintaining myocardial tissue integrity.
Major comments
Minor comments
As someone with expertise in cardiac development and cellular behaviours, I find this study provides strong and convincing quantitative data on the role of Llgl1 in suppressing cardiomyocyte extrusion and promoting epicardial dissemination on the ventricular surface. The genetic experiments, including mutant analysis and myocardial-specific rescue, were carefully performed in a region-specific manner, which provides much insight into the non-uniformity of myocardial tissue integrity. The generation of Tg(myl7:llgl1-mCherry) line is also a valuable tool for researchers in the field interested in understanding apicobasal polarity and cardiomyocyte development and regeneration.
A limitation of the study is the unclear link between epithelial polarity and basement membrane deposition, and how they synchronize to regulate cardiomyocyte integrity. The llgl1 mutant phenotype in increasing cardiomyocyte apical extrusion and Crb2 localization is interesting; however, the authors note that this appears to be a phenotype induced by epicardial defects. Epicardial cells are not known to exhibit apicobasal polarity and are fibroblastic by nature. Thus, the cellular mechanisms by which Llg1 regulates epicardial cell morphology or behaviours, and how it functions to regulate polarity in cardiomyocytes are not clearly defined in this work. In addition, clarification of the cell autonomous functions of Llgl1 in epicardial cells and/or cardiomyocytes would strengthen the findings.
Overall, the findings of this study would be of interest to cell and developmental biologists in the fields of epithelial polarity, cardiac morphogenesis, and extracellular matrix function. It provides nice conceptual advance in further elucidating the mechanisms that underlie myocardial tissue integrity and epicardial-myocardial interactions.
lustig. schon bald (diesen winter? nächsten winter?) wird die mehrheit verhungern und erfrieren,<br /> und die sogenannte "opposition" beschäftigt sich mit kleinscheiss.
wir brauchen viel mehr extremismus und viel mehr radikale lösungen (selbstversorgung...),<br /> und nicht diese schwule weichgespülte "wir sind was besseres" gelaber.
die AFD hat jeden tag angst vor parteiverbot,<br /> also werden so "radikale typen" wie ich von anfang an rausselektiert.
dabei sollten so "radikale typen" wie ich eine blutige revolution (einen militärputsch) führen,<br /> und die ganzen versager und verräter vom alten system an die wand stellen.
aber auf revolution habt ihr gar keine lust, ihr wollt auch nur reformen, und noch mehr gesetze,<br /> aber es gibt keine politische lösung, weil das system ist radikal falsch, also nicht reformierbar.
deswegen, für mich sind graue wölfe 1000 mal interessanter als die AFD.<br /> ich bin sowieso kein christ oder jude, aber auch kein moslem.<br /> einfach nur pragmatisch, ein echter "wissenschaftler", einer der wissen schafft.
"Their evil designs run against nature." -- Kevin Alfred Strom
(ich freu mich schon auf den prozess wegen "gefühlsverletzung", ich lach mich tot...)
Der Sturm Otis, der die Küste bei Acapulco verwüstet hat, hat sich in nur einem Tag von einem tropischen Sturm zu einem Hurrikan der Kategorie 5 entwickelt. Es ist noch unklar worauf dieser enorme Gewinn an Intensität zurückgeht. https://www.nytimes.com/2023/10/25/world/americas/hurricane-otis-mexico-intensity-surprise.html
That’s the honest-to-goodness HTML I have in the Markdown for this post. That’s it! There’s no special setup; I don’t have to remember to put specific elements on the page before calling a function or load a bunch of extra resources.1 Of course, I do need to keep the JS files around and link to them with a <script> tag.
There's nothing special about Web Components; the author could have just as easily put the script block itself there.
Die Internationale Energieagentur IEA hält eine Begrenzung der globalen Erhitzung aufgrund des schnellen Wachstums bei den erneuerbaren Energien für sehr schwierig, aber noch möglich. In ihrem Jahresbericht kommt sie zu dem Ergebnis, dass der Höhepunkt der Nachfrage nach Kohle, Gas und Öl bis 2030 erreicht werden wird. Die Energiepolitik der wichtigen Staaten ist aber bei der Umstellung auf Erneuerbare bei weitem nicht so ehrgeizig, als es nötig ist. https://www.liberation.fr/environnement/grace-aux-energies-bas-carbone-limiter-le-rechauffement-climatique-reste-possible-affirme-lagence-internationale-de-lenergie-20231024_YF7ZJA7WBFACRFIVCBRONJPKAA/
World Energy Outlook 2023: https://origin.iea.org/reports/world-energy-outlook-2023
Mehr zum World Energy Outlook 2023: https://hypothes.is/search?q=tag%3A%22report%3A%20World%20Energy%20Outlook%202023%22
Der Standardbericht zu dem neuen State of the climate-Report konzentriert sich auf die Forderung nach sofortigen Veränderungen in der Wirtschaft. Er enthält auch eine Übersicht zu den Temperaturanomalien in Österreich in diesem Jahr. https://www.derstandard.at/story/3000000192443/hoechsttemperaturen-wie-2023-gab-es-womoeglich-erstmals-seit-100000-jahren
Mehr zum State of the climate 2023-Report: https://hypothes.is/search?q=tag%3A%22report%3A%202023%20state%20of%20the%20climate%22;
Reviewer #2 (Public Review):
Summary:<br /> This study seeks to understand the connection between protein sequence and function in disordered regions enriched in polar amino acids (specifically Q, N, S and T). While the authors suggest that specific motifs facilitate protein-enhancing activities, their findings are correlative, and the evidence is incomplete. Similarly, the authors propose that the re-assignment of stop codons to glutamine-encoding codons underlies the greater user of glutamine in a subset of ciliates, but again, the conclusions here are, at best, correlative. The authors perform extensive bioinformatic analysis, with detailed (albeit somewhat ad hoc) discussion on a number of proteins. Overall, the results presented here are interesting, but are unable to exclude competing hypotheses.
Strengths:<br /> Following up on previous work, the authors wish to uncover a mechanism associated with poly-Q and SCD motifs explaining proposed protein expression-enhancing activities. They note that these motifs often occur IDRs and hypothesize that structural plasticity could be capitalized upon as a mechanism of diversification in evolution. To investigate this further, they employ bioinformatics to investigate the sequence features of proteomes of 27 eukaryotes. They deepen their sequence space exploration uncovering sub-phylum-specific features associated with species in which a stop-codon substitution has occurred. The authors propose this stop-codon substitution underlies an expansion of ploy-Q repeats and increased glutamine distribution.
Weaknesses:<br /> The preprint provides extensive, detailed, and entirely unnecessary background information throughout, hampering reading and making it difficult to understand the ideas being proposed.<br /> The introduction provides a large amount of detailed background that appears entirely irrelevant for the paper. Many places detailed discussions on specific proteins that are likely of interest to the authors occur, yet without context, this does not enhance the paper for the reader.
The paper uses many unnecessary, new, or redefined acronyms which makes reading difficult. As examples: (1) Prion forming domains (PFDs). Do the authors mean prion-like domains (PLDs), an established term with an empirical definition from the PLAAC algorithm? If yes, they should say this. If not, they must define what a prion-forming domain is formally. (2) SCD is already an acronym in the IDP field (meaning sequence charge decoration) - the authors should avoid this as their chosen acronym for Serine(S) / threonine (T)-glutamine (Q) cluster domains. Moreover, do we really need another acronym here (we do not). (3) Protein expression-enhancing (PEE) - just say expression-enhancing, there is no need for an acronym here.
The results suggest autonomous protein expression-enhancing activities of regions of multiple proteins containing Q-rich and SCD motifs. Their definition of expression-enhancing activities is vague and the evidence they provide to support the claim is weak. While their previous work may support their claim with more evidence, it should be explained in more detail. The assay they choose is a fusion reporter measuring beta-galactosidase activity and tracking expression levels. Given the presented data they have shown that they can drive the expression of their reporters and that beta gal remains active, in addition to the increase in expression of fusion reporter during the stress response. They have not detailed what their control and mock treatment is, which makes complete understanding of their experimental approach difficult. Furthermore, their nuclear localization signal on the tag could be influencing the degradation kinetics or sequestering the reporter, leading to its accumulation and the appearance of enhanced expression. Their evidence refuting ubiquitin-mediated degradation does not have a convincing control.
Based on the experimental results, the authors then go on to perform bioinformatic analysis of SCD proteins and polyX proteins. Unfortunately, there is no clear hypothesis for what is being tested; there is a vague sense of investigating polyX/SCD regions, but I did not find the connection between the first and section compelling (especially given polar-rich regions have been shown to engage in many different functions). As such, this bioinformatic analysis largely presents as many lists of percentages without any meaningful interpretation. The bioinformatics analysis lacks any kind of rigorous statistical tests, making it difficult to evaluate the conclusions drawn.
The methods section is severely lacking. Specifically, many of the methods require the reader to read many other papers. While referencing prior work is of course, important, the authors should ensure the methods in this paper provide the details needed to allow a reader to evaluate the work being presented. As it stands, this is not the case.
Overall, my major concern with this work is that the authors make two central claims in this paper (as per the Discussion).
The authors claim that Q-rich motifs enhance protein expression. The implication here is that Q-rich motif IDRs are special, but this is not tested. As such, they cannot exclude the competing hypothesis ("N-terminal disordered regions enhance expression"). The authors also do not explore the possibility that this effect is in part/entirely driven by mRNA-level effects (see Verma Na Comms 2019). As such, while these observations are interesting, they feel preliminary and, in my opinion, cannot be used to draw hard conclusions on how N-terminal IDR sequence features influence protein expression. This does not mean the authors are necessarily wrong, but from the data presented here, I do not believe strong conclusions can be drawn.
That re-assignment of stop codons to Q increases proteome-wide Q usage. I was unable to understand what result led the authors to this conclusion. My reading of the results is that a subset of ciliates has re-assigned UAA and UAG from the stop codon to Q. Those ciliates have more polyQ-containing proteins. However, they also have more polyN-containing proteins and proteins enriched in S/T-Q clusters. Surely if this were a stop-codon-dependent effect, we'd ONLY see an enhancement in Q-richness, not a corresponding enhancement in all polar-rich IDR frequencies? It seems the better working hypothesis is that free-floating climate proteomes are enriched in polar amino acids compared to sessile ciliates. Regardless, the absence of any kind of statistical analysis makes it hard to draw strong conclusions here.
Almost all thirty informants immediately focused on outdoor activities—tag, hide-n-seek, jumping rope, picnics, hiking, swimming, bike riding, random adventures with friends, and so on. Regardless of whether our informants grew up in a rural or urban setting, they typically recalled their girlhood as a time when media and popular culture were peripheral or absent from their lives
This is a concept hard for many to imagine nowadays given how reliant and incorporated media is in various popular culture
About 150,000transformed plants (T1 plants) expressing aT-DNA-located kanamycin-resistance gene(NPTID
Signature tagged mutagenesis, tag is kanamycin resistance; integrate into genome and then look for unusual phenotypes (kanamycin resistance), when found, this phenotype indicates that the gene relating to the usual phenotype was inactivated
Author Response
Reviewer #1 (Public Review):
This is a very exciting manuscript from Meng Wang's lab on lysosomal proteomics. They used several different protein tags to identify the lysosomal proteome. The exciting findings include A) specific lysosomal proteins exist in a tissue-specific manner B) lipl-4 overexpression and daf-2 extend life span using different mechanisms C) identification of novel lysosomal proteins D) demonstration of the function of several lysosomal proteins in regulation lysosome abundance and function.
We thank the reviewer for finding our manuscript exciting.
Reviewer #2 (Public Review):
In this manuscript, Yu and colleagues profile the lysosome content in C. elegans. They implement lysosome immunoprecipitation (Lyso-IP) for C. elegans and they convincingly show that this method successfully isolates lysosomes from whole worms. The authors find that the lysosomes of worms overexpressing the lysosomal lipase lipl4 are enriched for AMPK subunits and nucleoporins and that these proteins are required for the longevity of lipl-4 overexpressing worms. The authors also show that this is specific to this longevity pathway given that another long-lived worm strain (daf2) does not exhibit enrichment for nucleoporins nor does it require them for longevity. The authors go on to express the Lyso-IP tag in different tissues of C. elegans (muscle, hypodermis, intestine, neurons) and identify the tissue-specific lysosome proteomes. Finally, the authors use this method to identify lysosome proteins in mature lysosomes and they find new proteins that regulate lysosomal acidification.
The authors present a powerful tool to unbiasedly identify lysosome-associated proteins in C. elegans, and they provide an in-depth assessment of how this method can be used to understand longevity pathways and identify novel proteins. Understanding lysosomal differences in specific tissues or in response to different longevity conditions are exciting as it provides new insight into how organelles could control specific homeostasis responses. This tool and proteomics datasets also represent a great resource for the C. elegans community and should pry open new studies on the regulation and role of the lysosome at the organismal level.
We truly appreciate that the reviewer’s positive comment on our work.
Addressing the following suggestions would help strengthen this already strong manuscript. First, it would be helpful to validate selected candidates from the tissuespecific Lyso-IP to verify that the protocol is still specific with lower sample amounts. Second, it would be helpful to provide more details on the methods, notably for sample preparation and analysis, so that it can serve as a guideline for the community. Third, the manuscript contains a lot of data and conditions, which is great, but they may also feel disconnected in some cases and it could be helpful to focus the study on the main key findings.
We thank the reviewer’s comments. As suggested by the reviewer, we have also generated a CRISPR knock-in line for one hypodermis-specific candidate Y58A7A.1 that encodes a copper transporter and validated its hypodermis-specific lysosomal localization (new Supplementary Figure 2E).
As suggested by the reviewer, we have extended the method section on Lyso-IP to include more details. We believe that the new version should be sufficient for any lab to follow this protocol and conduct their own analyses. We will also take advantage of the eLife “Request a Protocol” feature to share the detailed version of the Lyso-IP method with researchers who are interested.
We have thoroughly reorganized the manuscript to increase the textual clarity and improve the connection between different analyses and results.
Reviewer #3 (Public Review):
The manuscript by Ji et al dissects the important role of lysosomes in cellular metabolism and signaling and their regulation by various associated proteins. The authors utilized deep proteomic profiling in C.Elegans to identify lysosome-associated proteins involved in regulating longevity and discovered the recruitment of AMPK and nucleoporin proteins in response to increased lysosomal lipolysis. Additionally, the authors found lysosomal heterogeneity across different tissues and specific enrichment of the Ragulator complex on Cystinosin-positive lysosomes.
Strengths of this work include the utilization of deep proteomic profiling to identify novel lysosome-associated proteins involved in longevity regulation, as well as the discovery of lysosomal heterogeneity and specific protein enrichments across different worm tissues. These findings point to a complex interplay between lysosomal protein dynamics, signal transduction, organelle crosstalk, and organism longevity.
One weakness of this work may be the limited scope of the study, as it focuses primarily on the identification and characterization of lysosome-associated proteins involved in longevity regulation, with limited mechanistic follow-up and some unsubstantiated claims.
We thank the reviewer for her/his helpful comments and suggestions. The primary goal of this manuscript is to provide new methods and resource to the community. We did have several biological findings from the current study, and mechanistic follow-up with these findings will be interesting future topics but may beyond the scope of the current manuscript. In addition, we have provided new experimental results to further support several claims that the reviewer has commented on.
Reviewer #2 (Public Review):
In this manuscript, Yu and colleagues profile the lysosome content in C. elegans. They implement lysosome immunoprecipitation (Lyso-IP) for C. elegans and they convincingly show that this method successfully isolates lysosomes from whole worms. The authors find that the lysosomes of worms overexpressing the lysosomal lipase lipl-4 are enriched for AMPK subunits and nucleoporins and that these proteins are required for the longevity of lipl-4 overexpressing worms. The authors also show that this is specific to this longevity pathway given that another long-lived worm strain (daf-2) does not exhibit enrichment for nucleoporins nor does it require them for longevity. The authors go on to express the Lyso-IP tag in different tissues of C. elegans (muscle, hypodermis, intestine, neurons) and identify the tissue-specific lysosome proteomes. Finally, the authors use this method to identify lysosome proteins in mature lysosomes and they find new proteins that regulate lysosomal acidification.
The authors present a powerful tool to unbiasedly identify lysosome-associated proteins in C. elegans, and they provide an in-depth assessment of how this method can be used to understand longevity pathways and identify novel proteins. Understanding lysosomal differences in specific tissues or in response to different longevity conditions are exciting as it provides new insight into how organelles could control specific homeostasis responses. This tool and proteomics datasets also represent a great resource for the C. elegans community and should pry open new studies on the regulation and role of the lysosome at the organismal level.
Addressing the following suggestions would help strengthen this already strong manuscript. First, it would be helpful to validate selected candidates from the tissue-specific Lyso-IP to verify that the protocol is still specific with lower sample amounts. Second, it would be helpful to provide more details on the methods, notably for sample preparation and analysis, so that it can serve as a guideline for the community. Third, the manuscript contains a lot of data and conditions, which is great, but they may also feel disconnected in some cases and it could be helpful to focus the study on the main key findings.
Any recommendations on Analog way of doing it? Not the Antinet shit
reply to u/IamOkei at https://www.reddit.com/r/Zettelkasten/comments/17beucn/comment/k5s6aek/?utm_source=reddit&utm_medium=web2x&context=3
u/IamOkei, I know you've got a significant enough practice that not much of what I might suggest may be helpful beyond your own extension of what you've got and how it is or isn't working for you. Perhaps chatting with a zettelkasten therapist may be helpful? Does anyone have "Zettelkasten Whisperer" on a business card yet?! More seriously, I occasionally dump some of my problems and issues into a notebook, unpublished on my blog, or even into a section of my own zettelkasten, which I never index or reconsult, as a helpful practice. Others like Henry David Thoreau have done something like this and there's a common related practice of writing "Morning Pages" that you can explore. My own version is somewhat similar to the idea of rubber duck debugging but focuses on my own work. You might try doing something like this in one of Bob Doto's cohorts or by way of private consulting sessions. Another free version of this could be found by participating in Will's regular weekly posts/threads "Share with us what is happening in your ZK this week" at https://forum.zettelkasten.de/. It's always a welcoming and constructive space. There are also some public and private (I won't out them) Discords where some of the practiced hands chat and commiserate with each other. Even the Obsidian PKM/Zettelkasten Discord channels aren't very Obsidian/digital-focused that you couldn't participate as an analog practitioner. I've even found that participating in book clubs related to some of my interests can be quite helpful in talking out ideas before writing them down. There are certainly options for working out and extending your own practice.
Beyond this, and without knowing more of your specific issues, I can only offer some broad thoughts which expand on some of the earlier discussion above.
I recommend stripping away Scheper's religious fervor, some of which he seems to have thrown over lately along with the idea of a permanent note or "main card" (something I think is a grave mistake), and trying something closer to Luhmann's idea of ZKII.
An alternate method, especially if you like a nice notebook or a particular fountain pen, might be to take all of your basic literature/fleeting notes along with the bibliographic data in a notebook and then just use your analog index cards/slips to make your permanent notes and your index.
Ultimately it's all a lot of the same process, though it may come down to what you want to call it and your broad philosophy. If you're anti-antinet, definitely quit using the verbiage for the framing there and lean toward the words used by Ahrens, Dan Allosso, Gerald Weinberg, Mark Bernstein, Umberto Eco, Beatrice Webb, Jacques Barzun & Henry Graff, or any of the dozens of others or even make up your own. Goodness knows we need a lot more names and categories for types of notes—just like we all need another one page blog post about how the Zettelkasten method works by someone who's been at it for a week. Maybe someone will bring all these authors to terms one day?
Generally once you know what sorts of ideas you're most interested in, you take fewer big notes on administrivia and focus more of your note taking towards your own personal goals and desires. (Taking notes to learn a subject are certainly game, but often they serve little purpose after-the-fact.) You can also focus less on note taking within your entertainment reading (usually a waste) and focusing more heavily on richer material (books and journal articles) that is "above you" in Adler's framing. You might make hundreds of highlights and annotations in a particular book, but only get two or three serious ideas and notes out of it ultimately. Focus on this and leave the rest. If you're aware of the Pareto principle or the 80/20 rule, then spend the majority of your time on the grander permanent notes (10-20%), and a lot less time worrying about the all the rest (the 80-90%).
In the example above relating to Marx, you can breeze through some low level introductory material for context, but nothing is going to beat reading Marx himself a few times. The notes you make on his text will have tremendously more value than the ones you took on the low level context. A corollary to this is that you're highly unlikely to earn a Ph.D. or discover massive insight by reading and taking note posts on Twitter, Medium, or Substack (except possibly unless your work is on the cultural anthropology of those platforms).
A lot of the zettelkasten spaces focus heavily on the note taking part of the process and not enough on the quality of what you're reading and how you're reading it. This portion is possibly more valuable than the note taking piece, but the two should be hand-in-glove and work toward something.
I suspect that most people who have 1000 notes know which five or ten are the most important to where they're going and how they're growing. Focus on those and your "conversations with texts" relating to those. The rest is either low level context for where you're headed or either pure noise/digital exhaust.
If you think of ideas as incunables, which notes will be worth of putting on your tombstone? In other words: What are your "tombstone notes"? (See what I did there? I came up with another name for a type of note, a sin for which I'm certainly going to spend a lot of time in zettelkasten purgatory.)
This is a test. I'm trying out syncing Hypothes.is with Obsidian
Eines der Rituale, das Mohammed von den Juden übernimmt, ist das dreimalige Beten am Tag, das er durch zwei weitere ergänzt hat. Von der anfänglichen Gebetsrichtung nach Jerusalem war bereits weiter oben die Rede. Für die Muslime führte Mohammed den Freitag-Gottesdienst als zentrales rituelles Geschehen ein — damit wollte er sich von den Juden unterscheiden, für die der Samstag, der Schabbath, der heilige Tag der Woche ist. Und auch das „Fasten am Aschura-Tag“, dem zehnten Tag im hebräischen Monat Tischri, erweiterte Mohammed zum Ramadan-Fasten. Ebenso hat Mohammed die Beschneidung für alle männlichen Kinder übernommen, allerdings nicht am achten Tag nach der Geburt wie bei den Juden, sondern generell erst sehr viel später. Juden nehmen vor jeder rituellen Handlung und auch vor dem Thora-Studium das rituelle Händewaschen vor, Muslime waschen bekanntlich Gesicht, Hände und Füße, bevor sie die Moschee zum Gebet betreten. Auch dabei dürfte es sich vermutlich um mehr als bloße Ähnlichkeiten handeln.
Der Teufel ist ein Imitator, aber ein schlechter.
eLife assessment
This study examines the expression of HDAC3 within DC compartment. Taking advantage of tamoxifen inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to occur during development around the CLP stage. Tamoxifen treated mice lack almost all pDC besides lymphoid progenitors. RNA seq studies identify multiple DC specific target genes within the remaining pDC - using Cut and Tag technology they validate some of the identified targets of HDAC3. Taken together, this study shows the requirement of HDAC3 on pDC but not cDC, congruent with the recent findings of a lymphoid origin of pDC.
Reviewer #1 (Public Review):
The work by Yijun Zhang and Zhimin He at al. analyzes the role of HDAC3 within DC subsets. Using an inducible ERT2-cre mouse model they observe the dependency of pDCs but not cDCs on HDAC3. The requirement of this histone modifier appears to be early during development around the CLP stage. Tamoxifen treated mice lack almost all pDCs besides lymphoid progenitors. Through bulk RNA seq experiment the authors identify multiple DC specific target gens within the remaining pDCs and further using Cut and Tag technology they validate some of the identified targets of HDAC3.<br /> Collectively the study is well executed and shows the requirement of HDAC3 on pDCs but not cDCs, in line with the recent findings of a lymphoid origin of pDC.
While the authors provide extensive data on the requirement of HDAC3 within progenitors, the high expression of HDAC3 in mature pDCs may underly a functional requirement. Have you tested INF production in CD11c cre pDCs? Are there transcriptional differences between pDCs from HDAC CD11c cre and WT mice?
A more detailed characterization of the progenitor compartment that is compromised following depletion would be important, as also suggested in the specific points.
Reviewer #2 (Public Review):
In this article Zhang et al. report that the Histone Deacetylase-3 (HDAC3) is highly expressed in mouse pDC and that pDC development is severely affected both in vivo and in vitro when using mice harbouring conditional deletion of HDAC3. However, pDC numbers are not affected in Hdac3fl/fl Itgax-Cre mice, indicating that HDCA3 is dispensable in CD11c+ late stages of pDC differentiation. Indeed, the authors provide wide experimental evidence for a role of HDAC3 in early precursors of pDC development, by combining adoptive transfer, gene expression profiling and in vitro differentiation experiments. Mechanistically, the authors have demonstrated that HDAC3 activity represses the expression of several transcription factors promoting cDC1 development, thus allowing the expression of genes involved in pDC development. In conclusion, these findings reveals HDAC3 as a key epigenetic regulator of the expression of the transcription factors required for pDC vs cDC1 developmental fate.
These results are novel and very promising. However, supplementary information and eventual further investigations are required to improve the clarity and the robustness of this article.
Major points<br /> 1) The gating strategy adopted to identify pDC in the BM and in the spleen should be entirely described and shown, at least as a Supplementary Figure. For the BM the authors indicate in the M & M section that they negatively selected cells for CD8a and B220, but both markers are actually expressed by differentiated pDC. However, in the Figures 1 and 2 pDC has been shown to be gated on CD19- CD11b- CD11c+. What is the precise protocol followed for pDC gating in the different organs and experiments?
2) pDC identified in the BM as SiglecH+ B220+ can actually contain DC precursors, that can express these markers, too. This could explain why the impact of HDAC3 deletion appears stronger in the spleen than in the BM (Figures 1A and 2A). Along the same line, I think that it would important to show the phenotype of pDC in control vs HDAC3-deleted mice for the different pDC markers used (SiglecH, B220, Bst2) and I would suggest to include also Ly6D, taking also in account the results obtained in Figures 4 and 7. Finally, as HDCA3 deletion induces downregulation of CD8a in cDC1 and pDC express CD8a, it would important to analyse the expression of this marker on control vs HDAC3-deleted pDC.
3) How do the authors explain that in the absence of HDAC3 cDC2 development increased in vivo in chimeric mice, but reduced in vitro (Figures 2B and 2E)? More generally, as reported also by authors (line 207), the reconstitution with HDAC3-deleted cells is poorly efficient. Although cDC seem not to be impacted, are other lymphoid or myeloid cells affected? This should be expected as HDAC3 regulates T and B development, as well as macrophage function. This should be important to know, although this does not call into question the results shown, as obtained in a competitive context.
4) What are the precise gating strategies used to identify the different hematopoietic precursors in the Figure 4 ? In particular, is there any lineage exclusion performed? Moreover, what is the SiglecH+ CD11c- population appearing in the spleen of mice reconstituted with HDAC3-deleted CDP? Data shown in Figure 4F should be expressed as log2 and not10. Finally, how do the authors explain that Hdac3fl/fl express Il7r, while they are supposed to be sorted CD127- cells?
5) What is known about the expression of HDAC3 in the different hematopoietic precursors analysed in this study? This information is available only for a few of them in Supplementary Figure 1. If not yet studied, they should be addressed.
6) It would be highly informative to extend CUT and Tag studies to Irf8 and Tcf4, if this is technically feasible.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary: Sharma, et al. report the characterization of the polar tube (PT) from the microsporidian species, Vairimorpha necatrix, using a combination of optical microscopy, cryo-ET, and proteomics. The polar tube is a fascinating invasion apparatus which mediates the translocation of the parasite into the inside of a host cell to initiate infection. Similar to results obtained previously in other species, the authors show that PT firing in Vairimorpha necatrix is extremely fast, occurring on the order of 1 sec, and that the extruded PT is over 100 microns long in this species. Using cryo-ET to image the PT at a high resolution, they find that it exists in two major states: both an empty state and a state filled with cargo, and that the thickness of the tube wall changes when cargo is present. Strikingly, the authors observed that one of the cargo components, the ribosomes, are organized ordered array that may have helical symmetry. Finally, the authors took advantage of a naturally occurring "His tag" on PTP3 to affinity purify PTP3-containing protein complexes and analyze the composition using proteomics.
Major comments
ln 139-140: The absolute handedness of something can be very tricky to determine by cryo-ET (but certainly is possible). Variable hardware configurations between microscopes and differing conventions between software packages (e.g., for what direction is a positive tilt angle) can lead to inversion of the apparent handedness in the final tomogram. How certain are the authors that the absolute handedness is indeed right handed, as this seems to vary between the various display items in the manuscript? For example, in Fig 1c, my impression is that ribosome helices are left handed, as they are also in the supplemental movie. If this isn't known with certainty, perhaps it would be sufficient to describe the apparent helical symmetry but state that the handedness is ambiguous.
Minor comments
ln 39-40: Perhaps also cite the E. cuniculi genome paper?
ln 97-98: It is interesting that the PT shortens in V. necatrix as well, and while I can pick this out in some of the individual traces in Sup Fig. 1b, it seems to get washed out in the trend line and isn't super obvious. If it isn't to laborious, it could be nice to add a panel showing the quantification of this (e.g., plotting the final length of each PT as a percentage of the maximum length achieved).
ln 98-100: Strictly speaking, I don't think the referenced figure shows the sporoplasm being transformed into an extended conformation, only that it is spherical upon exit. Simply reword this to make clear that the deformations are inferred to occur but not directly observed.
Because PT firing is so fast, the probability of trapping a PT in the process of transporting cargo would be pretty low. So then why does the PT still contain cellular cargo like ribosomes inside in the tomograms? Should these not have emerged in the sporoplasm which would enter the host cell? Are these "defective" spores that have failed to complete sporoplasm transport? Perhaps this is worth discussing.
ln 118: The authors note an apparent correlation between the phase of germination and the thickness of the tube wall but don't specify what this correlation is. Is it thicker in the early phase and thinner in later phase, or vice versa? One could imagine "empty" tubes existing before or after sporoplasm transport, for example, so I'm not sure I follow how the phase is being inferred from the tomograms.
ln 119-120: What is the evidence that the outer layer is made of PTPs, or that it is even protein (for example, as opposed to cell wall-like carbohydrate polymers)? I think this seems like a very reasonable hypothesis, but I would suggest explaining the logic and ensuring the degree of uncertainty is conveyed clearly. In light of this, I would also suggest changing figure labels, etc, that refer to the PTP layer (e.g., Fig. 3, PTPc and PTPe labels).
ln 121, 123: "sheathed by a thin layer" and "enveloped by a thick outer layer": is this an additional layer being described? Or is this referring to the putative PTP layer, and that its thickness is variable?
ln 125-126: While I understand how some features, like ribosomes, proteasomes, and generic membrane compartments could be identified, it is unclear to me how one would recognize the nucleus when inside the PT, nor are any examples shown. If the data is clear, perhaps the authors could show it in a figure? Otherwise, I suggest removing the claim regarding the nucleus.
The arrangement of the ribosomes in a subset of tubes is really fascinating! While the number of observations is relatively small (n=5), it seems like it should be possible to comment preliminarily on whether there is much variability in their helical arrangement. Do the helical parameters vary much between observations? Does the til, pitch, etc vary much, are the 5 occurrences very similar? Is there any sign that they are associated with a membrane? Also, since the ribosomes form a lattice-like arrangement, it seems like it would be possible to trace ribosome helices in both the left and right handed directions. How did the authors decide between the two possibilities? This doesn't seem to be discussed.
Fig. 2e: Are the two different colors/orientations meant to represent the two protamers of the ribosome dimer? When refined subvolumes are mapped back onto the original tomogram do the authors observe a similar crystalline arrangement of particles as in their segmentation? Are the orientations of the ribosomes correlated, and do the provide any evidence for the dimeric arrangement mentioned? The PlaceObjects plugin for Chimera can be very helpful for visualizing this: https://www.biochem.mpg.de/7939908/Place-Object
Supp figure 4(b-d): Perhaps these models could be colored by pLDDT scores (with a key indicating the color scheme), so the reader can assess the quality of the predictions?
How were the measurements of the membrane thickness and putative PTP layer carried out? On the tomogram projections? STAs? How were the boundaries of the layers established (e.g., map threshholding if STA?)? This information appears to be missing from the methods.
Some tubes that are labeled as 'PTempty' actually contain cargo and look dense (example supp. Fig 2c, left and middle panels). Is it fair to classify these as empty tubes?
Fig. 3d: I am not entirely clear on what is being shown here. Are independent reconstructions of PTcargo and PTempty superposed (aligned on membrane)? The description in the figure legend doesn't clearly say what is being displayed. I think it might be more clear to show these side-by-side instead of superposed (i.e., 4 panels instead of 2).
Sup Fig 1: Define S and SP in legend or just spell out on figure? Missing x-axis label on panel b.
Fig. 4b and Sup Fig 2a: The depictions of the PT in the spore here are left-handed. In a few species, the coil of the PT was found to form a right-handed helix (Jaroenlak, et al.), and it seems plausible that this may be a general feature that would be conserved across microsporidia. I appreciate that it might not be actually known to be right-handed in V. necatrix, but if there is no strong data either way, perhaps it would make sense for these depictions of the PT to be right-handed.
I think all three of us are more or less in consensus about this manuscript, and I largely agree with the other reviewers comments. I think after addressing reviewer suggestions, this will be a pretty nice story.
Overall, this manuscript from Sharma, et al. presents interesting new findings about the structure and cargo transport function of the microsporidian PT. Microsporidia infect a wide range of hosts, including humans, and how the PT mediates parasite entry into cells is poorly understood. The approaches used in this study are appropriate for tackling the questions at hand, and appear to be generally well executed and interpreted. The observation that ribosomes assemble into an array within the PT is very unexpected and quite fascinating, and may be of broader interest to researchers working on ribosome structure and function, in addition to researchers studying microsporidia. The approach to investigating proteins interacting with PTP3 was quite elegant, and yielded a list of potential interactors that appears to be of very high quality and is highly plausible based on the literature field. We think this work is a substantial advance in the field and provides important new insights into the organization of the PT. - Please define your field of expertise with a few keywords to help the authors contextualize your point of view:
Structural biology, microsporidia - Indicate if there are any parts of the paper that you do not have sufficient expertise to evaluate.
We are not experts in proteomics/mass spectrometry
Should I use zettelkasten? .t3_172ujnk._2FCtq-QzlfuN-SwVMUZMM3 { --postTitle-VisitedLinkColor: #9b9b9b; --postTitleLink-VisitedLinkColor: #9b9b9b; --postBodyLink-VisitedLinkColor: #989898; } questionI am a student in college in the UK studying A levels (Advanced levels), this includes mathematics, biology, chemistry and physics. I dont really take notes for mathematics so I wont be using any type of note taking system for that but for the sciences IDK what to do.
reply to u/Wooden-School-4091 at https://www.reddit.com/r/Zettelkasten/comments/172ujnk/should_i_use_zettelkasten/
This comes up fairly frequently. See https://hypothes.is/users/chrisaldrich?q=tag%3A%27zettelkasten+for+studying%27 and related links for other variations and advice on this theme.
ja. fehler:
In Bayern hat sich eine konservative Mehrheit gebildet, die 67,4 Prozent der abgegebenen Stimmen erhalten hat: CSU (die Union wird hier mitgerechnet unter zumindest potentiell noch konservativ, die Red.), Freie Wähler und AfD.
die einzige "konservative" partei ist die AFD, alle anderen sind betrüger ("conservatives in name only").
aber die AFD war noch nie teil einer regierung, sonst würde man sehen: auch die AFD kann nichts ändern, weil alle entscheidungen kommen von oben, und politiker verkaufen diese entscheidungen nach unten, wie in einer dauerwerbesendung. deswegen, auch die AFD ist nur "controlled opposition".
wenn politiker wirklich mal rebellieren gegen "oben", dann wird die zentralbank einfach die geldzahlungen einstellen, und am nächsten tag ist die regierung pleite. schon heute sind alle regierungen pleite, und leben von kredit von der zentralbank.
Gebt mir die Kontrolle über die Währung einer Nation, und es ist mir gleichgültig, wer die Gesetze macht.
-- Amschel Mayer Rothschild
deswegen: politik ist zeitverschwendung. aktivismus heisst selbsthilfe, selbstorganisation, kleinstaaten.
Reviewer #1 (Public Review):
Summary:<br /> Cincotta et al set out to investigate the presence of glucocorticoid receptors in the male and female embryonic germline. They further investigate the impact of tissue-specific genetically induced receptor absence and/or systemic receptor activation on fertility and RNA regulation. They are motivated by several lines of research that report inter and transgenerational effects of stress and or glucocorticoid receptor activation and suggest that their findings provide an explanatory mechanism to mechanistically back parental stress hormone exposure-induced phenotypes in the offspring.
Strengths:<br /> - A chronological immunofluorescent assessment of GR in fetal and early life oocyte and sperm development.<br /> - RNA seq data that reveal novel cell type specific isoforms validated by q-RT PCR E15.5 in the oocyte.<br /> - 2 alternative approaches to knock out GR to study transcriptional outcomes. Oocytes: systemic GR KO (E17.5) with low input 3-tag seq and germline-specific GR KO (E15.5) on fetal oocyte expression via 10X single cell seq and 3-cap sequencing on sorted KO versus WT oocytes - both indicating little impact on polyadenylated RNAs<br /> - 2 alternative approaches to assess the effect of GR activation in vivo (systemic) and ex vivo (ovary culture): here the RNA seq did show again some changes in germ cells and many in the soma.<br /> - They exclude oocyte-specific GR signaling inhibition via beta isoforms.<br /> - Perinatal male germline shows differential splicing regulation in response to systemic Dex administration, results were backed up with q-PCR analysis of splicing factors.
Weaknesses:<br /> - The presence of a protein cannot be entirely excluded based on IF data (staining of spermatids is referred to but not shown).<br /> - The authors do not consider post-transcriptional level a) modifications also trigged by GR activation b) non-coding RNAs (not assessed by seq).<br /> - Sequencing techniques used are not total RNA but either are focused on all polyA transcripts (10x) or only assess the 3' prime end and hence are not ideal to study splicing, The number of replicates in the low input seq is very low and hence this might be underpowered. Since Dex treatment showed some (modest) changes in oocyte RNA - effects of GR depletion might only become apparent upon Dex treatment as an interaction.<br /> - Effects in oocytes following systemic Dex might be indirect due to GR activation in the soma.<br /> - Even though ex vivo culture of ovaries shows GR translocation to the nucleus it is not sure whether the in vivo systemic administration does the same.
The conclusion that fetal oocytes are "intrinsically buffered to GR signalling" is very strong, given that "only" poly A sequencing and few replicates of 3-prime sequencing have been analyzed and information is lacking on whether GR is activated in germ cells in the systemically dex-injected animals.
This work is a good reference point for researchers interested in glucocorticoid hormone signaling fertility and RNA splicing. It might spark further studies on germline-specific GR functions and the impact of GR activation on alternative splicing.
While the study provides a characterization of GR and some aspects of GR perturbation, and the negative findings in this study do help to rule out a range of specific roles of GR in the germline, there is still a range of other potential unexplored options. The introduction of the study eludes to implications for intergenerational effects via epigenetic modifications in the germline, however, it does not mention that the indirect effects of reproductive tissue GR signaling on the germline have indeed already been described in the context of intergenerational effects of stress. Also, the study does not assess epigenetic modifications.
The conclusion that the persistence of a phenotype for up to three generations suggests that stress can induce lasting epigenetic changes in the germline is misleading. For the reader who is unfamiliar with the field, it is important to define much more precisely what is referred to as "a phenotype". Furthermore, this statement evokes the impression that the very same epigenetic changes in the germline have been observed across multiple generations.
The evidence of the presence of GR in the germline is also somewhat limited - since other studies using sequencing have detected GR in the mature oocyte and sperm.
The discussion ends again on the implications of sex-specific differences of GR signaling in the context of stress-induced epigenetic inheritance. It states that the observed differences might relate to the fact that there is more evidence for paternal lineage findings, without considering that maternal lineage studies in epigenetic inheritance are generally less prevalent due to some practical factors - such as more laborious study design making use of cross-fostering or embryo transfer. Since the authors comment on RNA-mediated inheritance it seems inevitable to again consider indirect effects.
Comparing the two points, you can see that E is Pareto inefficient because both the rich and poor are better off at R than at E. The income distribution at R is also the one at which the poor are as rich as they can possibly be in this economy, as indicated by the feasible frontier. This is the point that Rawls favoured (and why we called it point R).
The impact of free international financial markets on East Asia, as viewed through the lens of this section's framework, is complex. On one hand, these markets have stimulated economic growth in the region by facilitating investment and access to global capital, which has the potential to positively affect individuals' financial wealth and physical assets. However, the benefits have not been equally distributed, contributing to growing income disparities. According to SSRN, "growth-promoting economic freedoms hamper future progress by raising inequalities." The disparities are rooted in differences in education, gender, and social class, influenced by institutions and technology.
Therefore, while free international financial markets have brought economic opportunities to East Asia, they have also exacerbated income inequality, highlighting the need to address these disparities by considering the interplay between institutions, technology, and individual endowments to achieve a more balanced outcome in the region.
Works Cited: Ilkay Yılmaz , Mehmet Murat Balkan , Mehmet Nasih Tağ Income Inequality and Economic Freedom: The 2000s
Total cost of ownership (TCO) addresses the total cost of software development from inception to sun setting. In 2011, the CRASH report stated the total cost of ownership for software code was $18/Line of Code (LOC). Of this, it is generally accepted that the majority of this cost is related to the maintenance of the software after its initial creation, with estimates ranging from 60-90%.
$18 /LOC
Reviewer #1 (Public Review):
The manuscript by Royall et al. builds on previous work in the mouse that indicates that neural progenitor cells (NPCs) undergo asymmetric inheritance of centrosomes and provides evidence that a similar process occurs in human NPCs, which was previously unknown.
The authors use hESC-derived forebrain organoids and develop a novel recombination tag-induced genetic tool to birthdate and track the segregation of centrosomes in NPCs over multiple divisions. The thoughtful experiments yield data that are concise and well-controlled, and the data support the asymmetric segregation of centrosomes in NPCs. These data indicate that at least apical NPCs in humans undergo asymmetric centrosome inheritance. The authors attempt to disrupt the process and present some data that there may be differences in cell fate, but this conclusion would be better supported by a better assessment of the fate of these different NPCs (e.g. NPCs versus new neurons) and would support the conclusion that younger centriole is inherited by new neurons.
Reviewer #2 (Public Review):
Royall et al. examine the asymmetric inheritance of centrosomes during human brain development. In agreement with previous studies in mice, their data suggest that the older centrosome is inherited by the self-renewing daughter cell, whereas the younger centrosome is inherited by the differentiating daughter cell. The key importance of this study is to show that this phenomenon takes place during human brain development, which the authors achieved by utilizing forebrain organoids as a model system and applying the recombination-induced tag exchange (RITE) technology to birthdate and track the centrosomes.
Overall, the study is well executed and brings new insights of general interest for cell and developmental biology with particular relevance to developmental neurobiology. The Discussion is excellent, it brings this study into the context of previous work and proposes very appealing suggestions on the evolutionary relevance and underlying mechanisms of the asymmetric inheritance of centrosomes. The main weakness of the study is that it tackles asymmetric inheritance only using fixed organoid samples. Although the authors developed a reasonable mode to assign the clonal relationships in their images, this study would be much stronger if the authors could apply time-lapse microscopy to show the asymmetric inheritance of centrosomes.
Showers and shared kitchen facilities are in a warm, permanent building, rather than the canvas tents used sixty miles away in Seattle. Every tiny house has a porch and a bathroom. As an equal proportion of the development’s total price tag, each house costs $88,000; on an individual basis they are $19,000 per unit.
Homeless people are actually getting their basic necessities covered and it helps with better hygiene that other people living in those cabins and camps desperately needed.
Author Response
The following is the authors’ response to the original reviews.
We thank the reviewers for their thoughtful assessment of our work and their valuable critiques which we will address in the “Recommendations for the authors” section below. In particular, we appreciate Reviewer #3 noting the value of the C. elegans model system and our efforts to bridge models with our study. We agree with the reviewer that there is a need to clarify the rationale, presentation and interpretation of our results. We have substantially revised the text in our manuscript and Figure legend to address this issue, and provided extensive new commentary and citations to lay out the logic behind our experiments. Indeed, it was our oversight not being more thorough about this initially. We have further adjusted our conclusions to be less unequivocal. Finally, we added an RPM-1 signaling diagram (Fig. 8A) to more clearly annotate the players in the RPM-1/MYCBP2 signaling network that were evaluated genetically in Fig. 8. Importantly, we provide clearer commentary on how genetic enhancer effects with known RPM-1 binding proteins and the absence of genetic suppression in vab-1/Eph receptor double mutants with components of the RPM-1/FSN-1 ubiquitin ligase complex are consistent with the biochemical finding that MYCBP2 stabilizes but does not degrade EphB2. Text edits reflecting these points are in the abstract, the C. elegans results section starting on line 411, and the discussion on lines 499, 502-504 and 541.
Following extensive discussions between the three reviewers, all three agree that the C. elegans data, as presented, does not add to, and in fact might harm, your bottom line. Our combined suggestion is to take this data out unless you plan to improve it substantially. All reviewers are perplexed by Figure 2F and the presumed interactions of cytosolic proteins with the extracellular domain of EPHB2. At the very least, please provide some suggestions/model/interpretation.
We have adjusted our manuscript substantially to address this. Please see detailed comments in the individual Reviewer sections below.
We would like to thank the reviewers for their thorough examination of our manuscript, constructive criticisms, and helpful suggestions.
Reviewer #1 (Recommendations For The Authors):
The work is extensive in my view, and mostly of high quality. See minor comments on some of the figures below.
Thank you very much.
Two more major comments :
- I don't think the C. elegans work adds to - in fact I think it hurts - the statement that this regulatory mechanism is specific to EphB2. I would advise the authors to take it out.
We agree that C. elegans has a sole Eph receptor called VAB-1 and is therefore not a specific model for EPH2B. However, testing MYCBP2 specificity for EPHB2 was not the goal or our perceived value for the C. elegans experiments. We now clarify this in the text of the Results section.
Rather, we are providing evidence that the C. elegans ephrin receptor interacts genetically with known MYCBP2/RPM-1 binding proteins. Moreover, we now provide an extensive array of citations to note that genetic enhancer interactions between different RPM-1/MYCBP2 binding proteins is well established. The reviewer has nicely highlighted for us that we handled the C. elegans genetics in too cursory a fashion in our original manuscript. We appreciate this being noted and have now aimed to make this substantially clearer. We hope the reviewer agrees that our revised C. elegans section accomplishes this goal.
Furthermore, we extensively revised the text of the Results to emphasize a key point: our observation that axon termination defects are not suppressed in vab-1; fsn-1 and vab-1; rpm-1 double mutants excludes the possibility that the VAB-1 Eph receptor is a substrate that is inhibited or degraded by the RPM-1/FSN-1 ubiquitin ligase complex. If the VAB-1 Eph receptor were ubiquitinated and degraded by the RPM-1/FSN-1 complex, we would have observed a suppression of phenotype in vab-1; rpm-1 double mutants. The precedent for this genetic relationship between the RPM-1 ubiquitin ligase and its substrates that are degraded has been established by several prior studies (PMID: 15707898; PMID: 31676756; PMID: 35421092). We now more clearly note that the absence of genetic suppression in vab-1; rpm-1 double mutants and vab-1; fsn-1 double mutants is consistent with the non-canonical stabilizing role of MYCBP2 on EPHB2 that was observed in our biochemical experiments with mammalian cells.
We also adjusted the text of the manuscript to stress that we are testing genetic interactions between the VAB-1 Eph receptor and known RPM-1 binding proteins. This is a key point, as genetic enhancer interactions are consistent with the Eph receptor functioning in the RPM-1 signaling network. This concept has been well established for RPM-1 binding proteins as now noted in our revised text with an extensive number of additional citations to published work.
Based on the above arguments, we respectfully disagree with the reviewer that our C. elegans data should be removed from the paper. To re-iterate, we are not trying to evaluate specificity for MYCBP2 and EPHB2 in C. elegans. Rather, our goals are twofold: 1) To ask whether there is an evolutionarily conserved functional genetic link between Eph receptors and known RPM-1 binding proteins. 2) To provide further in vivo genetic evidence invalidating the hypothesis that Ephrin receptors could be ubiquitination substrates that are inhibited/degraded by MYCBP2.
Text edits reflecting these points are in the abstract, the C. elegans results section starting on line 411, and the discussion on lines 499, 502-504 and 541.
- The cellular responses are not robust and the effects of MYCBP2 KO - although significant - are minor in most cases. But I don't think more experiments will help here.
We interpret the comment about the robustness to mean that the extent to which a given cellular response is affected by the loss of MYCBP2 is minor. First, the cellular responses themselves are typical of previous studies and depend on the cellular biology underlying them. For example, a growth collapse of ~50-60% over a background of 10% (Fig. 7) is typical for these sorts of assays (PMID: 37369692; PMID: 33972524; PMID: 17785182). A decrease of cell area by ~25% (Fig. 3) is quite substantial if one considers how much of a cell’s volume is taken up by the nucleus and organelles. Second, the phenotypes elicited by the loss of MYCBP2 are likely brought on by a decrease in EphB2 protein levels, but not its complete absence, as suggested by our biochemical experiment. Given that EphB2 complete loss only affects the cellular responses to a limited extent, the minor effects are not a surprise (e.g. for GC collapse: PMID: 23143520). Nevertheless, the subtle changes in cellular phenotypes, elicited by EPHB2 signaling are often sufficient to achieve proper cell positioning and cell response to guidance cues. For instance, regulation of the growth cone collapse of the outgrowing axons requires delicate changes that are dynamic and temporal.
Minor:
Fig 1C - EPHA3 and EPHB2 seem to run in different sizes, is this the case? In 2A they run at the same size.
We believe this size discrepancy is due to different percentages of SDS-PAGE gels used to resolve proteins. In Fig. 1C, we used a 6% gel for a Western blot analysis of both EPHA3/-B2-FLAG (~130 kDa) and MYCBP2 (~510 kDa). In Fig. 2A however, we performed Western blot analysis using 10% resolving gel to separate and detect EPHA3/-B2-FLAG along with MYC-FBXO45 (~30 kDa). We have reviewed the results obtained from additional biological replicates of this experiment, and observed a similar pattern in gel migration of EPHA3/-B2-FLAG across all replicates.
Fig1F - I can't trust the MYCBP2 blot.
Indeed, the MYCBP2-EPHB2 co-IP with endogenous proteins was not convincing. We now repeated this experiment using rat cortical neurons, and the results replace the previous Fig. 1F panel as mentioned on line 158.
In Fig2b the authors claim that there is enhancement in the binding of MYCBP2 and EPHB2 upon FBXO45 expression. For this type of statement quantification is required.
The quantification is now included in Fig. 2C and its significance is mentioned on line 180. Our conclusion about the enhancement stands.
Fig2G - it remained unclear to me where the binding site to MYCBP2 is, how long is the cytoplasmic tail in the DeltaICD protein?
Based on our experimental observations from Fig. 2E-H, we concluded that the fragment encompassing the extracellular domain(s) and/or transmembrane (TM) domain of EPHB2 is necessary for the protein complex formation with MYCBP2. We would like to accentuate that the EPHB2-MYCBP2 interaction might not be direct, and might involve other transmembrane protein(s) acting as a scaffold for EPHB2 and MYCBP2 binding. We did not pursue experiments to determine the exact region of the extracellular-TM portion of EPHB2 that is required for the interaction with MYCBP2.
The cytoplasmic tail in ΔICD protein consists of 25 aa of the N-terminal fragment of EPHB2 juxtamembrane (JM) region, which is adjacent to the TM helix, and followed by the 8 aa FLAG tag (EPHB2 ΔICD domain composition: extracellular domains – TM domain – 25 aa fragment of JM region – FLAG). We have determined the TM and JM sequences based on Hedger et al. (PMID: 25779975) and included the N-terminal portion of the JM region to facilitate proper ΔICD protein localization within the plasma membrane (PMID: 35793621). We modified the schematic in Fig. 2G to better visualise the EPHB2 truncations and now provide information on their size in the figure legend.
Always good to have a model of how all these proteins work together.
While we acknowledge that this would be helpful, we do not have a clear answer on how the EPHB2-MYCBP2 complex formation occurs. This requires further elucidation of the putative proteins involved in this ternary complex or testing the possibility that a MYCBP2 fragment is extruded extracellularly. Without these experiments there are too many possibilities to summarise into a clear model figure. We thus did not make any edits regarding these possibilities in the section starting on line 195.
Reviewer #2 (Recommendations For The Authors):
Overall, the experiments are classical experiments of co-immunoprecipitations, swapping experiments, collapse assays, and stripe assays which all are well carried out and are convincing.
Thank you for your encouraging comments.
Controls for the stripe assay may include Fc / Fc stripe assays.
We have performed these control experiments and now include their quantifications in the results sectioning concerning Fig. 3, starting on line 249, and those concerning Fig. 6 on line 381.
It is not clear to me why SD and not SEM has been used here for presentations.
Standard deviation (SD) measures the dispersion of a dataset relative to its mean. The standard error of the mean (SEM) measures how much discrepancy is likely in a sample’s mean compared with the population mean. Thus, SEM includes a statistical inference about the sampling distribution while SD is a less “processed” measurement that by definition is larger than SEM. SEM might make the data look less dispersed and many journals encourage the use of SD in bar graphs (PMID: 16223828).
Fig 7A: it is rather difficult to see 'branches' in Fig. 7A, better pictures and close-ups should be provided. How are branches defined? This piece of work needs more attention.
To remedy this shortcoming, we now provide inverted images with GFP signal in dark pixels overlaid on Fc (white) / eB2 (pink) stripes next to the original images.
Reviewer #3 (Recommendations For The Authors):
1) My most important suggestion to the authors would be to more carefully describe the results and their interpretation of the results. Sometimes, the distinction is not clear.
We modified the text throughout the manuscript to address this.
2) There are several cases, when the authors report on trends that are not statistically significant (1D, for example), or report no change, when it is clear that the addition of one more sample could have dramatically made a difference (4M - see point 12).
We agree that some of the nonsignificant differences could become significant if we added more Ns. But we prefer not to move our experimental design towards N-chasing and p-hacking (PMID: 25768323). The number of biological replicates is normally pre-determined before the onset of the experiment. Of course, some replicates can be discarded if there is a valid reason, such as a technical issue with the experiment or a positive control not working but this is not relevant for the dataset we have provided.
3) Data in 1F is very difficult to interpret.
As in response to Reviewer #1: Indeed, the MYCBP2-EPHB2 co-IP with endogenous proteins was not convincing. We now repeated this experiment using rat cortical neurons, and the improved results are in revised Fig. 1F.
4) Figure 2 puts Figure 1 in a strange perspective. If I understand correctly, fig 2 claims that EPHB2 interaction with MYCBP2 depends on FBXO45 - if that is the case then how does the binding in Figure 1 occur?
Indeed, we propose that the EPHB2-MYCBP2 interaction depends on FBXO45. In Fig. 2, we reveal that FBXO45 enhances the formation of the EPHB2-MYCBP2 complex. Thus, we suspect that the endogenous FBXO45 present in HeLa cells and neurons would mediate the interaction between EPHB2 and MYCBP2 in Fig. 1 experiments. We were unable to show this by Western blotting due to lack of reliable commercial antibodies against FBXO45, the complex containing endogenous FBXO45 and EPHB2 is also implied by our AP-MS data (Fig. 1B) and published databases.
5) I am still trying to wrap my mind around the results in 2G-H. So do MYCBP2 and FBXO45 bind the extracellular domain of EPHBP2? What does that mean?
(see also our response to Reviewer #1, end of their section) Based on our experimental observations from Fig. 2G-H, we conclude that the fragment encompassing the extracellular domain(s) and/or transmembrane domain of EPHB2 is necessary for the protein complex formation with MYCBP2 and FBXO45. Although there is a possibility that MYCBP2 directly binds the extracellular portion of EPHB2, we have not formally tested this hypothesis. MYCBP2 has been previously shown to interact with the extracellular portion of transmembrane N-cadherin (CDH2) via BioID proximity labeling and AP-MS proteomics approaches (PMID: 32341084).
Considering the results in Fig. 2A-B, we suspect that EPHB2-MYCBP2 interaction is indirect, as FBXO45 enhances this association. Secretion of FBXO45 and direct binding of FBXO45 to the extracellular cadherin (EC1-2) domains of N-cadherin has been documented (PMID: 25143387; PMID: 32341084). Although, not tested, this is also a possibility for EPHB2-FBXO45 mode of interaction. Nevertheless, we also cannot rule out the possibility that an unknown transmembrane protein binds EPHB2 extracellularly and the same unknown protein binds MYCBP2/FBXO45 intracellularly. Resolving this model is beyond the scope of this study and will require us to pursue extensive new lines of investigation.
6) I don't understand the stable Hela cell line CRISPR - is this a stable MYCBP2 deletion? In which case why is there only a reduction, not complete elimination of the protein? Or, is this a stable integration of a plasmid generating gRNA against MYCBP2? In which case, I would expect a homozygous null to emerge at some point. In any case, this is not well explained.
These lines are not derived from single cells infected with the CRISPR sgRNA-carrying viruses, therefore they are not clonal and probably contain some cells that express normal levels of MYCBP2, hence its detection on a Western. This is now clarified starting on line 221 and on line 608.
7) In 3C - is this the right statistical analysis?? I would say you want to claim the different effect of the control +/- eB2 compared to the effect in the mutant +/- eB2. Still should be significant but I think a more correct analysis.
We now include this comparison in Fig. 3C as well in the results section starting on line 234.
8) The robustness of the assay in Figure 3D is underwhelming – how was the area measured?
This is a live imaging experiment. Fig. 3D plots cell area at 60 minutes after ephrin-B2 addition as a fraction of the same cell’s area at 0 minutes (ephrin-B2 addition). For control cells that is a decrease of ~25%. If one considers that a cell’s nucleus and organelles like the Golgi Apparatus take up most of its volume, the magnitude is not that surprising.
9) Figure 3F – did you try to plot the relative area of overlap divided by the total cellular area? You might get a more striking phenotype. Also – claiming that this confirms that MYCBP2 is REQUIRED for EPHB2 function is a bit overstated, especially given that we don’t know (do you?) the EPHB2 mutant phenotype in this assay.
We preferred to stay with the original method of image quantification which we use for other assays. With respect to the requirement of MYCBP2 for EPHB2 function in the stripe assay, our logic is rooted in the observation that native HeLa cells do not respond to ephrin-B2 stripes (45.46 ± 7.62% of cells on eB2 stripes v. Fc; data not shown). When they are transfected with EPHB2 expression plasmids they do, therefore we assume that EPHB2 expression endows them with a sensitivity to eB2 stripes. A loss of MYCBP2 attenuates this sensitivity. We clarified this starting on line 246 and on line 251.
10) I didn't quite get the difference between 4A and 4B.
We apologize for the confusion. In Fig 4A, we used a stable HeLa cell line that has tetracycline-inducible expression of EPHB2-FLAG. Using these cells, we subsequently generated CTRLCRISPR or MYCBP2CRISPR cells. In these cells we then induced EPHB2 expression with tetracycline and observed that deletion of MYCBP2 resulted in the reduction of EPHB2 protein levels. To confirm this observation and to rule out the possibility that EPHB2 protein reduction is an effect of the CRISPR lines generation, we tested whereas MYCBP2 deletion reduces EPHB2, which has been transiently overexpressed (Fig. 4B). We hence conclude that loss of MYCBP2 decreases EPHB2 that was either expressed from a stable locus (Fig. 4A) or from transient transfection (Fig. 4B). We modified the Results section starting on line 262 to make this point clear.
11) The entire link to lysosomal degradation should be strengthened. Perhaps I am confused, but if the reduced EPHB2 levels in MYCBP2 mutant cells result from impaired lysosomal degradation then inhibiting the lys-deg should bring the protein levels back to normal (i.e. CRISPR control) - no? As currently presented, I do not understand nor do I think the claim is strongly supported by the data.
Before treatment with inhibitors, EPHB2 levels in MYCBP2CRISPR cells are already 40% lower than they are in CTRLCRISPR cells and in all our attempts, inhibitors can only rescue/restore EPHB2 in MYCBP2CRISPR cells to a level that is lower than in CTRLCRISPR cells. But this restoration is greater in MYCBP2CRISPR than in MYCBP2CTRL cells (BafA1: 19% increase in CTRL cells and 40% in MYCBP2CRISPR cells; CoQ: 10% comparing to 35%). This indicates that EPHB2 degradation through the lysosomal pathway in MYCBP2CRISPR cells is stronger, explaining why EPHB2 degradation is promoted in MYCBP2CRISPR cells, compatible with reduced EPHB2 levels and enhanced EPHB2 ubiquitination.
12) 4M, O - reporting ns based on these data seems a bit strange to me... Add one point and it will be strongly significant.
See our response to point (2), above. We prefer not to invoke potential p-hacking.
13) 7d - so what are you claiming? That the cellular response to eB1 but not eB2 is affected by the addition of FBD1? this is almost the opposite of what you wrote in the text...
We treated the cells with two different ephrin-B ligands to make a stronger conclusion. When using ephrin-B1, growth cone collapse in FBD1 WT is not significant comparing to Fc treatment. When using ephrin-B2, growth cone collapse in FBD1 WT is not as significant as it is in FBD1 mut group (* versus ). We interpret this as meaning that the EPHB2-mediated growth cone collapse to both ligands is dampened, when we disrupt the EPHB2-MYCBP2 association. The difference between these two ligands might be due to their different affinities for the receptor or signalling kinetics.
14) By far the weakest link in this paper is the worm part. I think it's a pity because strengthening this would affect the significance of the finding. First, the authors mention new genes without introducing their relationship to the signaling pathway tested. Second, the textual logics should be strengthened. Finally and most importantly, when the difference between the phenotypic severity is so strong (vab-1 and rpm-1) then I think it's impossible to say anything from the double mutant.
We appreciate the reviewer noting that they appreciate the value and importance of the C. elegans model. The goals of our C. elegans experiments were twofold:
1) To evaluate genetic interactions between the VAB-1 Eph receptor and known RPM-1 binding proteins. This was not clearly explained in the original manuscript nor was the published precedent for these types of genetic enhancer experiments provided. We have now rectified this by substantially revising the text of the Results C. elegans section starting on line 431 and by adding several citations.
2) Our C. elegans genetics confirmed that the VAB-1 Eph receptor is not inhibited/degraded by the RPM-1/MYCBP2 ubiquitin ligase complex. We have now revised the text to draw this point out more clearly.
To further address the reviewer’s concerns, we have added a new schematic (Fig. 8A) to show the relationship between the RPM-1 and the RPM-1 binding proteins (FSN-1/FBXO45 and GLO-4/SERGEF) we are testing. We chose FSN-1 because it is part of the RPM-1 ubiquitin ligase complex and we chose GLO-4 because it functions outside the context of RPM-1 ubiquitin ligase signaling via the GLO-1 Rab GTPase to influence late endosomal/lysosomal biogenesis.
Regarding the reviewer’s concern that different penetrance/frequency of defects between rpm-1 mutants and vab-1 mutants means outcomes with vab-1; rpm-1 double mutants cannot be interpreted. We respectfully disagree. An extensive number of published studies have demonstrated that RPM-1 binding proteins have milder phenotypes than rpm-1 mutants and display genetic enhancer effects as double mutants with one another (PMID:17698012, PMID: 22357847, PMID: 25010424, PMID: 24810406). We now make this point much more clearly. While the frequency of axon termination defects in rpm-1 mutants is high it is not completely saturated as the defect is not 100%. Moreover, a major point of the vab-1; rpm-1 double mutants is that they do not have a significant reduction in phenotypic penetrance/frequency. Thus, our system is fully capable of resolving genetic suppression, which did not occur. We now make this point much more carefully and clearly.
To further address the reviewer’s concern, we have softened language about the VAB-1/Eph receptor functioning in the same pathway as RPM-1 throughout the manuscript. While we think this is still the case, because the frequency of axon termination defects is not fully saturated in rpm-1 mutants and defects could potentially become more severe (i.e. the hook might occur closer to the head of the animal rather than in the midbody). Nonetheless, this is not a critical point and we think it is more important to be clear about the two major goals and objectives of our C. elegans experiments. We hope the reviewer agrees that our rationale, logic and conclusions are more clearly and accurately drawn in the revised paper.
Beyond just audio recordings so for that reason two of our senior 00:15:02 researchers Benjamin Hoffman and Maddie cusumano have also developed a biologer benchmark data set and so a biologer is an animal born tag like the one in the image on the right here 00:15:14 and these produce very valuable data because they can inform us about animal ecophysiology and allow us to improve conservation by monitoring animal movements and behaviors with very high 00:15:27 resolution
GET/active/Return the content of the active file open in Obsidian. Returns the content of the currently active file in Obsidian.返回Obsidian中当前活动文件的内容。 If you specify the header Accept: application/vnd.olrapi.note+json, will return a JSON representation of your note including parsed tag and frontmatter data as well as filesystem metadata. See "responses" below for details.如果您指定头部 Accept: application/vnd.olrapi.note+json ,将返回包括解析标签、前置数据以及文件系统元数据在内的笔记的JSON表示。有关详细信息,请参见下面的"responses"。
该接口返回当前活动文件内容
How do you think about the relationship between social media and “real life”?
We have to admit that people nowadays cannot live without social media. Living in 2023, I believe that social media is no longer a tool to communicate, it is becoming more and more diverse and combines all kinds of functions such as a platform to show people "who they are". I know too many people who are social phobias but are a "stars" on social media. Back then, people like them would be judged as "people who don't live on the Internet", however, no one would tag them as "strange" anymore. Therefore, yes, social media has been included in real life and is making real life better and more diverse.
Starting a blog .t3_16v8tfq._2FCtq-QzlfuN-SwVMUZMM3 { --postTitle-VisitedLinkColor: #9b9b9b; --postTitleLink-VisitedLinkColor: #9b9b9b; --postBodyLink-VisitedLinkColor: #989898; } Hey everyone- I’m still trying to wrap my head on how to organize this.I have my antinet growing and I want to start a blog with the use of one of my notes as a springboard.Do I9 votesWork on the blog and store the index cards after the note that I’m drawing inspiration fromCreate a new blog section in my antinet and place them thereStore them in wherever and create an hub note that points to them
reply to u/RobThomasBouchard at https://www.reddit.com/r/antinet/comments/16v8tfq/starting_a_blog/
The answer is:<br /> D: Start a "blog" where you post your notes as status updates and interlink them a bit. When you've got enough, you organize them into a mini thesis and write a longer article/blog post about it.
Examples: - https://hypothes.is/users/chrisaldrich?q=tag%3A%22thought%20spaces%22 and - https://indieweb.org/commonplace_book#The_IndieWeb_site_as_a_Commonplace_book
tl;dr: Use your website like a public, online zettelkasten. 🕸️🗃️
We’d like to set additional cookies to understand how you use GOV.UK, remember your settings and improve government services.
test
And the rise of deepfakes—wholly AI-generated content, as opposed to the Pelosi video’s edited clips from real (fact-checkable) speeches—will only make this problem more thorny and more urgent.
With the introduction of deepfakes into society it is very important that online users educate themselves on how to properly spot these videos. It would be nice if platforms could tag these videos with warnings but since most don't, it is crucial to fact check each piece of content that may seem suspicious.
o a URL or multiple URLs. Including a document’s DOI in the metadata of a web page will ensure that annotations appear on that document regardless of where it’s hosted.For example, an article published by Cell includes the tag<meta name="citation_doi" content="10.1016/j.
this I don't quite understand
Jerry Michalski says that The Brain provides him with a "neighborhood perspective" of ideas when he reduces the external link number for his graph down to 1.
This is similar to Nicholas Luhmann's zettelkasten which provided neighborhoods of related notes based on distance from any particular note.
Also similar to oral cultures who relied on movement through their environment for encoding memories and later remembering them. [I'll use the tag "environmental memory" to track this until a better name comes along.]
Hi, I just started to use Zettlr for my thoughts, in stead of just individual txt-files. I find it easy to add tags to notes. But if you read manuals how to use ZettelKasten, most seem to advice to link your notes in a meaningful way (and describe the link). Maybe it's because I just really started, but I don't find immediate links when I have a sudden thought. Sometimes I have 2 ideas in the same line, but they're more like siblings, so tagging with the same keyword is more evident. How do most people do this?
reply to u/JonasanOniem at https://www.reddit.com/r/Zettelkasten/comments/16ss0yu/linking_new_notes/
This sort of practice is harder when you start out in most digital apps because there is usually no sense of "closeness" of ideas in digital the way that is implied by physical proximity (or "neighborhood") found in physical cards sitting right next to or around each other. As a result, you have to create more explicit links or rely on using tags (or indexing) when you start. I've not gotten deep into the UI of Zettlr, but some applications allow the numbering (and the way numbered ideas are sorted in the user interface) to allow this affordance by creating a visual sense of proximity for you. As you accumulate more notes, it becomes easier and you can rely less on tags and more on direct links. Eventually you may come to dislike broad categories/tags and prefer direct links from one idea to another as the most explicit tag you could give a note . If you're following a more strict Luhmann-artig practice, you'll find yourself indexing a lot at the beginning, but as you link new ideas to old, you don't need to index (tag) things as heavily because the index points to a card which is directly linked to something in the neighborhood of where you're looking. Over time and through use, you'll come to recognize your neighborhoods and the individual "houses" where the ideas you're working with all live. As an example, Luhmann spent his life working in sociology, but you'll only find a few links from his keyword register/subject index to "sociology" (and this is a good thing, otherwise he'd have had 90,000+ listings there and the index entry for sociology would have been utterly useless.)
Still, given all this, perhaps as taurusnoises suggests, concrete examples may help more, particularly if you're having any issues with the terminology/concepts or how the specific application affordances are being presented.
Note: This rebuttal was posted by the corresponding author to Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Reviewer #1:
- If doable, image dynein and dynactin simultaneously in the Halo-DYNC1H1/DCTN4-SNAP iNeurons. Co-movement of dynein and dynactin towards the somatodendritic compartment and their separate movement in the anterograde direction along the axon would provide the most convincing evidence for the key claims of the manuscript.
Please see the planned revision section for our response
Reviewer #2:
Major comment (requires additional experimentation)
- While the data presented do certainly suggest that dynein and Lis1 are transported anterogradely on separate vesicular cargoes from dynactin and Ndel1, the study would be much stronger if supported by dual imaging of dynein and dynactin to prove that these proteins do indeed move in association with separate vesicular populations. I would like to see dual-color kymograph traces showing that the proteins move independently. The authors should be able to accomplish this using their dual Halo-DYNC1H1/DCTN4-SNAP hESC line. To acquire and analyze this data might take several months, but it would greatly strengthen this paper. If the authors do this experiment, they may also be able to address the mechanism of reversal of anterograde cargoes which they speculate about in the Discussion, which would add even more interest and insight.
Please see the planned revision section for our response
Minor comments (addressable without additional experimentation)
- The authors deduce that 1-4 Halo fluorochromes corresponds to 1-2 dynein molecules. This implies that the cells are homozygous for the Halo tag, but I do not see this addressed explicitly. The authors should state explicitly whether the lines generated for their study are heterozygous or homozygous for the tag. If the cells are heterozygous, which would seem most likely, then they may be underestimating the number of dyneins per spot and should take this into account.
We have added whether lines are homozygous or heterozygous to the manuscript. We also include a new Supplementary Figure panel (Fig S6) showing the genotyping data. In summary, all lines are homozygous except for PAFAH1B1-Halo (hESCs) which is heterozygous.
- Why are the moving spots lower in intensity than the NEM-treated static spots. It appears to suggest that they may be associated with different structures. This should be clarified and discussed.
Our data suggest that the fast-moving spots have fewer dyneins than NEM treated static spots. We suggest this is because the fast-moving cargos are smaller than the average cargo and therefore have fewer dyneins on them. This is also supported by the smaller number of dyneins reported previously on endosomes as compared to the large lysosomes. We have clarified this in the discussion (page 7-8).
- The authors state in the Results that most of the dynein spots were diffusing, often along microtubules, but they do not visualize microtubules so how do they know this? They may need to remove the phrase "often along microtubules".
This has been removed.
- At the end of the Introduction the authors state that their data "allow us to understand how the dynein machinery drives long-range transport in the axon". This is an overstatement. The "how" in this sentence is not addressed in this study.
We have softened the sentence by adding the phrase “better understand”.
- The conclusion that dynein binds to cargos stably throughout their transport along the axon is based on measurements of the fastest moving cargoes but the authors do not provide data on the distribution of velocities for the entire population of retrograde cargoes. It is not valid to extrapolate the behavior of a small number of cargoes to the entire population. The average may be much slower than the fastest cargoes. Moreover, even for the fastest organelles the authors cannot say that the dynein is stably bound because they did not track single cargoes and thus do not know that the cargoes moved continuously in one single bout of movement for 500 µm; it is possible that the cargoes moved in multiple consecutive bouts interrupted by brief pauses and dynein motors may have exchanged between bouts.
We have added a section to the discussion to highlight that other cargos may behave differently from the fastest ones (page 7). We have also clarified the assumptions that lead us to expect a slower arrival time of the first signal (page 5).
- The authors say that "it is clear that at least some dyneins remain on cargoes throughout their transport along the axon". As explained above, the data do not prove this so this statement should be removed.
We have softened this sentence from “it is clear” to “our results suggest” and explained in more detail why we make this conclusion
- The authors note that most of the dynein spots were not moving processively and state that this is consistent with prior studies showing that only a subset of dynein is actively involved in transport. However, as they note elsewhere, dynein is both motor and cargo and most axonal dynein is transported at slow average velocities so maybe they should be more explicit about what they mean by "involved in transport".
We have clarified we mean fast axonal transport and thank the reviewer for highlighting this point.
- When the authors note that most of the dynein in axons is transported in the slow component of axonal transport, they should also cite the work of Pfister and colleagues who were the first to show this (PMID 8824315 and 8552592).
This was an omission on our part. The references have now been added.
- The authors propose that dynein and Lis1 are transported together but there were significantly fewer anterogradely transported Lis1 particles than dynein particles. This should be discussed.
We have added more information to the discussion. Although we cannot rule out this effect being due to the heterozygous tagging of our LIS1 cell line, we do not witness the same decrease in events in the retrograde direction. Therefore, we believe there is a subset of anterogradely moving dynein lacking LIS1. As discussed in the manuscript, this subset may already be bound to dynactin and therefore not require LIS1.
- For the statistical analysis, the authors should provide p values in the legends for the comparisons that are judged to be "not significant". The authors should also be consistent in how they label differences that are not significant - they mark them as "ns" in Fig. 1, but in the other figures they do not, leaving some ambiguity about whether particular comparisons were not tested or were found to be not significant. For example, in Fig. 4C the average speed of the dynactin is about 0.5 µm/s greater than for the other proteins and the spread in the data suggest that this could be significant, but no significance is indicated on the plot, implying p>0.05. It is not clear how confident we can be that there is no difference.
We have now included all p values in the figure legends and have removed the “ns” in Fig 1D. In our revised manuscript, only significant differences are highlighted in the figures.
Reviewer #3:
- if I look at the kymographs, trajectories appear rather complex, pausing, standing still, moving and everything mixed. The explanation of how actual trajectories are extracted and on what basis is very short, too short for me. I think the authors should expand this. Furthermore, I think it would be good if the authors would present, in their kymographs examples of the tracked (and also the not included) tracks. Maybe in supplementary info.
The analysis of this data used the Trackmate Fiji plugin. This tracks spots frame to frame in a movie and then outputs the data of the tracks. No data was extracted from kymographs but they were used as a graphical illustration of the moving spots. To better explain our analysis pipeline, we have expanded our methods section and have added an example of a tracked movie (Video 15) as well as highlighted the tracked spots in one kymograph example (Figure 7S).
- I found 'velocity' ill defined. I get the impression, judging from the number of points (compared to the other parameters) that the authors determine the average velocity of each individual trajectory. That is an important parameter (but should indeed be called 'trajectory averaged' velocity), but might not be the only one useful to learn from the data, where trajectories do not always appear to have constant speeds (pausing, etc.). Why do the authors not determine point-to-point velocities and plot histograms of those for all the trajectories (simply plot histograms of all the displacements between subsequent data points in trajectories)? This might provide great insight into the actual maximum velocity and the fraction of pausing or moving in opposite direction etc., providing much more molecular detail than currently extracted from the data.
The reviewer is correct. We have measured the average velocity of the spots from the beginning of the track to the end. We have clarified this in the text. Furthermore, as stated above in the revision plan, we are currently doing the additional analysis and will include it in the final revision
- I was a bit surprised to read that the authors have gone to the effort to create a dual-color labeled cell line, but did not do actual correlative two-color measurements (or at least show them). It would be so insightful to see dynein and dynactin move separately in the anterograde direction.
Please see the planned revision section for our response.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Summary - The authors use a CRISPR knock-in gene editing strategy to label endogenous dynein, dynactin (p62 or Arp11) and dynein regulators (Ndel1 and Lis1) with Halo or SNAP tags. They do this in human iPSC and ESC cell lines engineered to express doxycycline-inducible NGN2 cloned into a "safe harbor" site of the genome. They induce the cells to differentiate into iNeurons using doxycycline and image the tagged proteins in axons with single molecule sensitivity using HILO illumination. The paper is clearly written, the description of the methods is thorough, and the data and figures (including the videos) are of good quality. The use of gene editing to knock the tags into the endogenous gene loci is a superior strategy to classic overexpression strategies. The authors also make effective use of microfluidic chambers to ensure the axons are uniformly orientated and coaligned over a distance of 500µm.
Major comment (requires additional experimentation)
Minor comments (addressable without additional experimentation)
Referee Cross-Commenting
There seems to be agreement among all three reviewers that the authors should perform dual imaging of dynein and dynactin to prove that these proteins do indeed move together in the retrograde direction but separately in the anterograde direction. This would strengthen the study greatly.
General assessment - There are now multiple papers that have analyzed axonal transport of cargoes in iPSC-derived neurons, but this one appears to be the first to do it by tagging dynein motors and with single-molecule sensitivity. The principal conclusions are (1) that dynein is capable of long-range movement in axons and (2) that dynein moves dynein/Lis1 complexes are transported anterogradely in association with distinct cargoes from dynactin/Ndel1 complexes. The former is a modest conclusion and is entirely expected so not very impactful, but the latter is interesting and novel. The difference between the average velocities for the four proteins in the anterograde and retrograde directions is striking. All four move at similar velocities in the retrograde direction but in the anterograde direction, dynein and Lis1 move significantly faster than dynactin and Ndel1. Given these data, it is reasonable to infer that these proteins are being transported in two separate sets of cargoes. As the authors note in their Discussion, this is important because it could provide a mechanism for transporting dynein components anterogradely in a less active form that could then be activated when the components come together in the distal axon. However, I feel that one critical experiment is missing, which is to perform dual labeling of anterogradely transported dynein and dynactin in the same cells (see major comment). Without this experiment, the evidence is indirect.
Audience - If confirmed by the dual labeling experiment, the authors' conclusions would represent a conceptual and mechanistic insight into the mechanism of bidirectional transport in axons that would be of broad interest to neuronal cell biologists studying neuronal trafficking.
Expertise - This reviewer has expertise in the neuronal cytoskeleton, live imaging and axonal transport and has some experience working with iPSC-derived neurons.
Note: This preprint has been reviewed by subject experts for Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
To image dynein in the axon at a single-molecule level, Fellows et al. used neuron-inducible human stem cell lines to Halo/SNAP tag endogenous dynein components by gene editing, and visualized fluorescently labeled protein molecules in differentiated neurons in microfluidic chambers by HILO microscopy-based live imaging. Using those cutting edge technologies, the authors demonstrate that in the axon, not only dynein and dynactin but also the dynein regulators LIS1 and NDEL1 can move long distance retrogradely towards the somatodendritic compartment. They also show that dynein /LIS1 move faster than dynactin/NDEL1 in the anterograde direction, suggesting that they are delivered separately to the distal end of the axon. The approach to study subcellular motility of endogenous dynein/dynactin is creative, the data are solid. I would like to suggest one experiment to support more strongly the authors' conclusions:<br /> If doable, image dynein and dynactin simultaneously in the Halo-DYNC1H1/DCTN4-SNAP iNeurons. Comovement of dynein and dynactin towards the somatodendritic compartment and their separate movement in the anterograde direction along the axon would provide the most convincing evidence for the key claims of the manuscript.
Referee Cross-Commenting
I agree with Reviewer 2 that the authors should clarify whether the knockin lines for dynein are homozygous. I also agree with both Reviewers 2 and 3 that the authors should do more analysis of the kymographs to obtain more information.
This is an elegant study on dynein motility and transport in vivo. The experimental approaches and findings presented in this manuscript are very valuable contributions to the field of dynein/dynactin and axonal transport. The results showing that dynein/dynactin can move long-range retrogradely in the axon are in good agreement with previous findings that dynein-driven cargo transport is highly processive, and the data suggesting that dynein and dynactin/NDEL1 are trafficked separately to the distal tip of the axon provide new insights into the regulatory mechanisms for the subcellular distribution and activity of molecular motors. Together these findings provide conceptual advances for understanding axonal transport. They will be of great interest to not only scientists in the field of intracellular transport but also those in cellular neurobiology.
# Set the priority when loading # e.g., zsh-syntax-highlighting must be loaded # after executing compinit command and sourcing other plugins # (If the defer tag is given 2 or above, run after compinit command) zplug "zsh-users/zsh-syntax-highlighting", defer:2
[!NOTE]
zplug load中,要设置加载优先级,可以使用?flashcard
defertag - If the defer tag is given 2 or above, run after compinit command
blood pressure chart
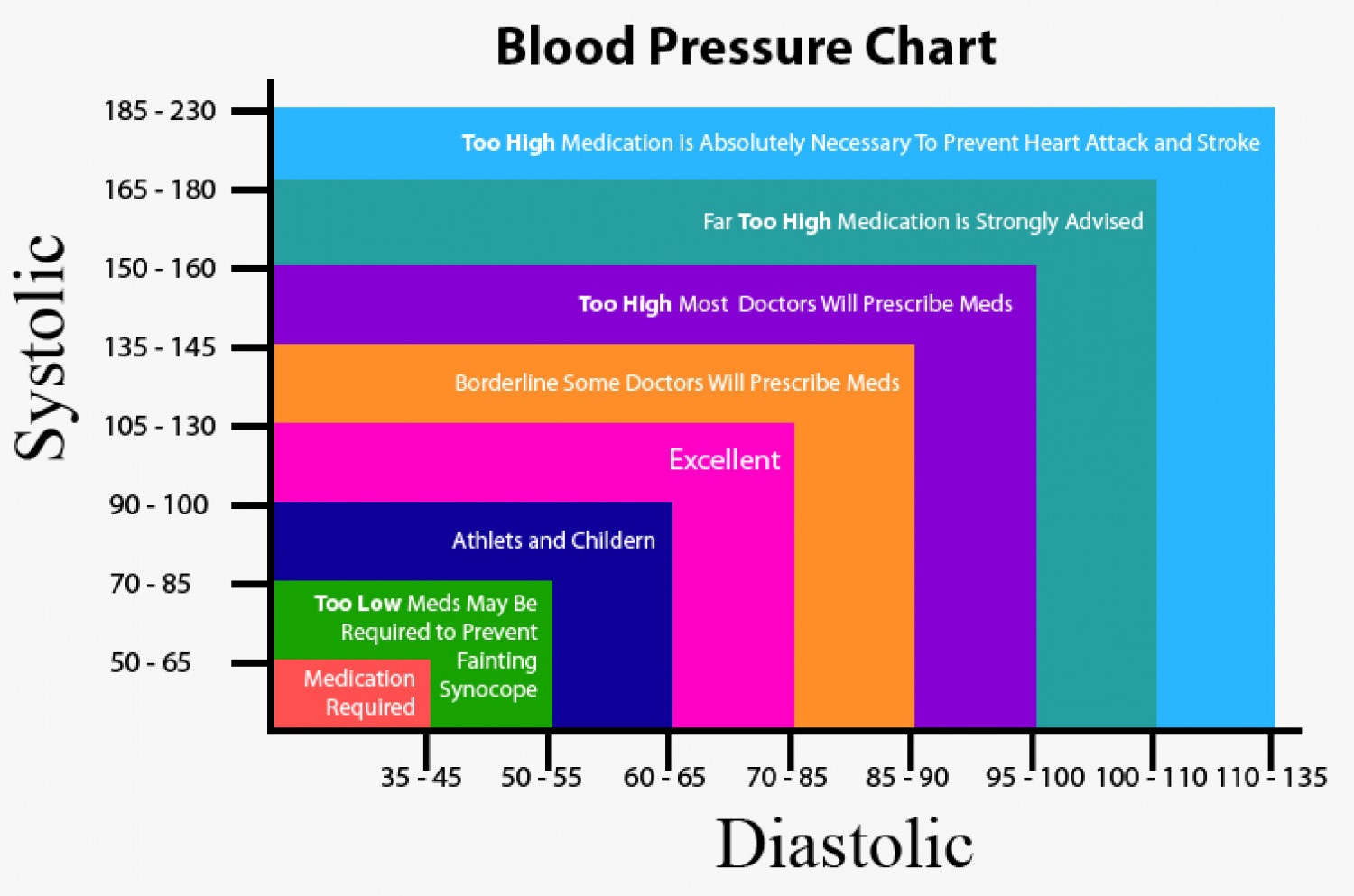
Reviewer #2 (Public Review):
Summary:<br /> The paper sought to determine the number of myosin 10 molecules per cell and localized to filopodia, where they are known to be involved in formation, transport within, and dynamics of these important actin-based protrusions. The authors used a novel method to determine the number of molecules per cell. First, they expressed HALO tagged Myo10 in U20S cells and generated cell lysates of a certain number of cells and detected Myo10 after SDS-PAGE, with fluorescence and a stained free method. They used a purified HALO tagged standard protein to generate a standard curve which allowed for determining Myo10 concentration in cell lysates and thus an estimate of the number of Myo10 molecules per cell. They also examined the fluorescence intensity in fixed cell images to determine the average fluorescence intensity per Myo10 molecule, which allowed the number of Myo10 molecules per region of the cell to be determined. They found a relatively small fraction of Myo10 (6%) localizes to filopodia. There are hundreds of Myo10 in each filopodia, which suggests some filopodia have more Myo10 than actin binding sites. Thus, there may be crowding of Myo10 at the tips, which could impact transport, the morphology at the tips, and dynamics of the protrusions themselves. Overall, the study forms the basis for a novel technique to estimate the number of molecules per cell and their localization to actin-based structures. The implications are broad also for being able to understand the role of myosins in actin protrusions, which is important for cancer metastasis and wound healing.
Strengths:<br /> The paper addresses an important fundamental biological question about how many molecular motors are localized to a specific cellular compartment and how that may relate to other aspects of the compartment such as the actin cytoskeleton and the membrane. The paper demonstrates a method of estimating the number of myosin molecules per cell using the fluorescently labeled HALO tag and SDS-PAGE analysis. There are several important conclusions from this work in that it estimates the number of Myo10 molecules localized to different regions of the filopodia and the minimum number required for filopodia formation. The authors also establish a correlation between number of Myo10 molecules filopodia localized and the number of filopodia in the cell. There is only a small % of Myo10 that tip localized relative to the total amount in the cell, suggesting Myo10 have to be activated to enter the filopodia compartment. The localization of Myo10 is log-normal, which suggest a clustering of Myo10 is a feature of this motor.
Weaknesses:<br /> One main critique of this work is that the Myo10 was overexpressed. Thus, the amount in the cell body compared to the filopodia is difficult to compare to physiological conditions. The amount in the filopodia was relatively small - 100s of molecules per filopodia so this result is still interesting regardless of the overexpression. However, the overexpression should be addressed in the limitations.<br /> The authors have not addressed the potential for variability in transfection efficiency. The authors could examine the average fluorescence intensity per cell and if similar this may address this concern.<br /> The SDS PAGE method of estimating the number of molecules is quite interesting. I really like this idea. However, I feel there are a few more things to consider. The fraction of HALO tag standard and Myo10 labeled with the HALO tagged ligand is not determined directly. It is suggested that since excess HALO tagged ligand was added we can assume nearly 100% labeling. If the HALO tag standard protein is purified it should be feasible to determine the fraction of HALO tagged standard that is labeled by examining the absorbance of the protein at 280 and fluorophore at its appropriate wavelength. The fraction of HALO tagged Myo10 labeled may be more challenging to determine, since it is in a cell lysate, but there may be some potential approaches (e.g. mass spec, HPLC).<br /> In Figure 1B, the stain free gel bands look relatively clean. The Myo10 is from cell lysates so it is surprising that there are not more bands. I am not surprised that the bands in the TMR fluorescence gel are clean, and I agree the fluorescence is the best way to quantitate.<br /> In Figure 3C, the number of Myo10 molecules needed to initiate a filopodium was estimated. I wonder if the authors could have looked at live cell movies to determine that these events started with a puncta of Myo10 at the edge of the cell, and then went on to form a filopodia that elongated from the cell. How was the number of Myo10 molecules that were involved in the initiation determined? Please clarify the assumptions in making this conclusion.<br /> It is stated in the discussion that the amount of Myo10 in the filopodia exceeds the number of actin binding sites. However, since Myo10 contains membrane binding motifs and has been shown to interact with the membrane it should be pointed that the excess Myo10 at the tips may be interacting with the membrane and not actin, which may prevent traffic jams.
Note: This rebuttal was posted by the corresponding author to Review Commons. Content has not been altered except for formatting.
Learn more at Review Commons
Reviewer 1 major comments:
The authors show one configuration of the E1-E2 heterodimer in Figure 4d. As shown, the E1 protein is exterior to the E2 protein and would suggest E1 is on the surface on the spike complex and virus surface. However, another configuration of the glycoproteins has E2 on the exterior of E1 and also on the exterior of the virus. The latter conformation is what has been observed in cryoEM studies of alphaviruses. The first configuration represents the E1-E2 between the three heterodimers which are important for spike assembly. The reason the orientation of the E2-E1 dimer is important is the authors speculate on the importance of the 6 CHIK residues not found in ONNV based on the structure, but the structural interpretation is, in my opinion, not correct.
We thank reviewer 1 for pointing out the correct E2-E1 heterodimer configuration. To address this, we corrected the position of E2 and E1 in Figure 4 based on previous cryoEM study1, keeping E2 always on the exterior in the E2-E1 heterodimer. We also replaced the Indian Ocean Lineage (IOL) E2-E1 structure1 in the original Figure 4 with the CHIKV 181/clone 25 structure which was recently analyzed by Katherine Basore et al.2. In a single E2-E1 heterodimer, all six unique CHIKV positive selection sites are located on the outside of the structure after correcting the configuration. In addition, we investigated two of the unique CHIKV positively selected sites that are important for virion production, E2-V135 (V460 in the original manuscript version) and E1-V220 (V1029 in the original manuscript version), in trimerized structure of E2-E1 heterodimers. We found that the E2-V135 and E1-V220 residues in one heterodimer are facing E2 of the neighboring heterodimer on either side. Interestingly, while V135 is embedded between the E2 proteins of two different heterodimers, E1-V220 is partially embedded by E1 and the neighboring E2 and partially exposed to the outside. This suggests that even though both E2-V135 and E1-V220 might be crucial for CHIKV E2-E1 trimerization, E1-V220 provides an additional docking site for host factor interactions. We thank review 1 again for this important comment leading to these new findings. We have updated Figure 4F-4G and the corresponding result section (lines 201-209) in this partially revised manuscript.
- Validation of E1 interaction with SPSC3 and eIF3k needs to be stronger. Some concerns/questions are listed below. A myc tag was inserted between E3 and E2. How efficiently does furin cleave E3 from E2 in this virus and how are viral titers of the myc-tagged virus compared to the non-tagged virus? I ask because is the IP looking at what is being pulled down by E2 or E3-myc-E2 that could be part of the spike polyprotein? The authors found E2 interacts with E3, E1 and a list of other host proteins. These results suggest several interactions including E2-host factor, E2-E1, E2-E3, E2-E1-host factor, E2-E3-E1, E2-E3-host factor. In figure 6d, and the subsequent conclusions, the authors suggest E1 is interacting with the host factor and do not see E2 alone and very low amounts of E3-E2-6K-E1. based on how the IP was performed I am not sure how an interaction between E1 and SPCS3 alone, without E2, would be detected. I would also like to see a reciprocal pull down using E1 and also E2 to see if these host factors are pulled down.
We thank the reviewer for these concerns. Given the low viral protein expression in macrophages (Figure 1A), we need an efficient system to enrich for large amounts of CHIKV glycoproteins for identifying host interactors through mass spectrometry. Adding tag/reporter proteins, such as mCherry, between E3 and E2 have been used to label alphavirus glycoproteins in previous study2, which is why we chose to use this myc tag labeling strategy coupled with myc Ab-conjugated agarose beads for AP-MS. However, like reviewer 1 speculated, inserting myc tag between E3 and E2 does attenuate CHIKV infectivity according to the reduced supernatant viral titers of 293T cells transfected with CHIKV/myc-E2 genomic RNA in comparison to those of cells transfected with unmodified CHIKV vaccine strain 181/clone 25 genomic RNA (shown in revision plan). Despite the attenuation, CHIKV/myc-E2 harvested from transfected 293T cells still reaches a titer over 108 pfu/ml, which allowed us to identify interactors by AP-MS.
We further analyzed the cleavage efficiency of glycoproteins by comparing the expression levels of E3-E2-6K -E1, E3-E2 (p62), E2, and E3 in 293T cells transfected with unmodified CHIKV or CHIKV/myc-E2 genomic RNA (result shown in revision plan). We didn’t detect any uncleaved forms of glycoproteins in cells transfected with either unmodified CHIKV or CHIKV/myc-E2 RNA when we probed with E2 antibody. However, probing with E3 antibody prior to longer exposure of the immunoblot showed higher E3-E2-6k-E1 and E3-E2 (p62) levels in cells transfected with CHIKV/myc-E2 RNA, suggesting that both mature E2 and E2-containing precursor polyproteins are available to be pulled down. Overall, the expression levels of mature E2 detected by E2 antibody are similar.
We thank reviewer 1 for providing a thorough dissection of all the possible interactions between the identified host factors and cleaved/uncleaved glycoproteins. This is a very interesting question. As reviewer 1 mentioned that E1 usually appears with E2 or E3-E2 in heterodimer forms, we were also surprised to find that E2 does not interact with either of the two host factors. To address this, we plan to conjugate E2 and E1 to protein A/G beads, respectively, for a reciprocal pulldown to validate CHIKV glycoprotein interactions with SPCS3 and eIF3k. Results from this experiment will be included in the fully revised manuscript.
- If CHIK E1 is interacting with the host factors and that is antagonizing the antiviral response of SPSC3 (as one example), then what do pull downs using ONNV structural proteins look like? One would expect reduced interactions because the different amino acid causes a different E2-E1 dimer or attenuates the E1-host factor binding site.
We thank Reviewer 1 for this insightful suggestion. We agree that it would be informative to examine the interactions between ONNV glycoproteins and identified host factors (SPCS3 and eIF3k). Unfortunately, there is no commercial ONNV glycoprotein antibody available making this experiment unfeasible. Interestingly, we did observe reduced interactions between the host factors SPCS3 and eIF3k and the CHIKV E1-V220I mutant (V1029I in original manuscript version) where the positively selected site in E1 was mutated to the homologous ONNV residue (please refer to our response to Reviewer 3’s major comment #1). This result suggests that the ONNV glycoproteins likely have an attenuated E1-host factor binding site as the reviewer speculated.We have included this as Figure 7A in partially revised manuscript.
- E1 and E2 are thought to interact during polyprotein translation and the initial dimer forms in the ER. If E1 is interacting with SPSC3 in the ER, is E2 also present? Or is a population of E1 not interacting with E2 in order to inhibit SPSC3? I would love a model of how the authors see all these factors coming together for this new role of E1.
We thank Reviewer 1 for proposing this interesting hypothesis. Given the unexpected absence of E2 in our validation of host factor-E1 pulldown, we speculate that a group of free E1 proteins with distinct function is interfering with host factors in the ER, which is a model worth further investigation and discussion. A great example of this is the alphavirus nonstructural protein 3 (nsP3) that plays essential roles in RNA replication, although depending on the alphavirus not all of the nsP3 in the cell colocalizes with dsRNA, suggesting there is a separate distinct pool of nsP3 outside of active viral replication complex that interacts with host factors in these observed larger cytoplasmic aggregates3. To address this, we plan to use laser confocal microscopy to observe the interactions between host factors (SPCS3, eIF3k), and CHIKV E2 and E1. We will include this result as well as our proposed model in the fully revised manuscript.
Reviewer 1 minor comments:
- In Figure 1c, (-) RNA is shown but in the rest of the figures (+) RNA is shown. Show both or select one. I do find it interesting the (-) RNA levels are similar over time, even at 4 hours post transfection (early time). Related to this, ONNV has higher levels of (-) RNA but what is known about structural protein levels in ONNV and CHIK in macrophages? Are there comparable levels of CP and GP being produced?
We thank Reviewer 1 for this comment. The (-) RNA is synthesized before the synthesis of subgenomic mRNA and therefore can reflect more accurately early viral replication and nonstructural protein functions. This is the reason why we consider the (-) RNA levels evaluated by specific nsP1 TaqMan probes to be more appropriate for determining early stage differences between ONNV and CHIKV replication in Figure 1 as the goal of that figure is to define the steps in CHIKV life cycle that are more efficient than those of ONNV in THP-1 derived macrophages. On the other hand, the (+) RNA evaluated by E1 primers that we used in the later figures monitors viral RNA synthesis over time in the reflection of genomic (+) RNA and subgenomic mRNA transcribed from (-) RNA templates. Similar levels of (+) RNA and contrasting virion titers really point the difference to the later stages of subgenomic mRNA translation, viral glycoprotein secretion, and assembly.
We have generated ONNV/myc-E2 reporter virus and assessed viral glycoprotein expression through flow cytometry using a FITC -conjugated anti-myc antibody in the THP-1 derived macrophages transfected with CHIKV/myc-E2 and ONNV/myc-E2 (shown in revision plan). The results show that the expression of ONNV glycoproteins is more inhibited than that of CHIKV glycoproteins, though both of their expression levels in macrophages seem to be suppressed. Since there is no commercial ONNV antibody available, we were unable to compare capsid expression levels between the two viruses. Overall, differences in the myc-tagged glycoprotein expression levels of the two viruses reveals ONNV defect in either structural protein translation or glycoprotein maturation .
- Figure 2e and figure 3 have ONNV has the first bar followed by CHIK. In figure 1 and 2b, CHIK is first and then ONNV. helps the reader to have the controls in the same order.
We thank Reviewer 1 for this suggestion. We have changed the order of ONNV and CHIKV bars in figure 2E and figure3 so the CHIKV bar consistently comes first in all the figures.
- Line 143-145 the authors discuss that when ONNV is the backbone and CHIK proteins are inserted the infection is more attenuated because of the E2 and E1 are from CHIK and ONNV, not the same virus (could also be E2-CP interactions are disrupted). However the chimeras made with the CHIK backbone (in Figure 2) have a mismatch between E2 and E1 as well.
We thank Reviewer 1 for this informative comment. We agree that the incompatible E2-E1 heterodimer formation may not be the only reason that causes attenuation of ONNV/CHIKV E1 and ONNV/CHIKV E2. There may be multiple factors contributing to the fitness of the chimeras, which requires more in-depth mechanistic investigations and is out of the scope of this study. We have now removed the explanation “potentially due to incompatible heterodimer formation between ONNV E2 and CHIKV E1” in line 144.
- When discussing the residues that were found in the FEL and MEME analysis, the authors start the amino acid numbering from CP and continue along the polyprotein. Usually when discussing amino acids in the structural proteins, each protein starts at amino acid 1. So V460 would be E2-V135. It would also be useful to know what the residues in ONNV were at these positions to see if amino acids changed in charge, size, bond forming potential, etc. Showing these residues in the E2-E1 conformation found in the virion would also allow one to find adjacent residues that could explain differences in spike assembly and potentially where/how E1 is binding to a host protein.
We thank Reviewer 1 for this comment. We revised the amino acid numbers in the manuscript to start from the beginning of each structural protein. To look more into these residues in ONNV, we aligned CHIKV and ONNV from different lineages and compared the 6 positively selected sites (refer to our response to Reviewer 1’s minor comment #5). We found that E2-135 and E1-220 which are essential for CHIKV production are valines in all the aligned CHIKV strains. For the aligned ONNV strains, E2-135 are all leucines and E1-220 are all isoleucines. While valine, leucine and isoleucine are all amino acids with hydrophobic side chains, valine has the shortest side chain. The length of the side chains may lead to different hydrophobic properties that affect protein folding, which warrants further structural analysis.
- How effective is a non-attenuated CHIK strain in infecting macrophages? Could you make a SINV-La Reunion chimeric virus (which is BSL2) to see if a higher percentage of macrophages are infected and is this potentially contributing to the increased pathogenesis of La Reunion? Also how different is 181/25 with a pathogenic strain in the E2 and E1 residues? and compared to ONNV?
We thank Reviewer 1 for this question, which is also raised by Reviewer 2. In order to address this question, we plan to use the virulent CHIKV La Reunion strain to study the infection of THP-1 derived macrophages with non-attenuated CHIKV in BSL-3. We are getting trained in the BSL-3 facility and will soon be certified.
We thank Reviewer 1 for this insightful suggestion on investigating the conservation of these positively selected sites in different strains. We have aligned the sequences of ONNV and CHIKV strains from different lineages, including CHIKV vaccine strain 181/clone 25 and Thai strain AF15561 (the parental strain of CHIKV 181/clone 25) (alignment shown in revision plan). We found that the two positively selected sites with negative effects on virion production, E2-135 and E1-220 (sites 460 and 1029 in original manuscript version), are very conserved in either CHIKV or ONNV strains. CHIKV E2-135 is always valine (V) regardless of the lineages, while ONNV E2-135 is always leucine (L). CHIKV E1-220 is always V, while ONNV E1-220 is always isoleucine (I).
We also analyzed the amino acid heterogeneity of E2-135 and E1-220 in 397 CHIKV patient sequences from NCBI Virus database. Most of the amino acids at these 2 sites are V. The counts of each amino acid at E2-135 and E1-220 is summarized in table below. This result suggests that valine residues at E2-135 and E1-220 are crucial for CHIKV fitness and strongly selected during viral evolution. The sequence alignment and table will be included and discussed in the fully revised manuscript .
E2-135
E1-220
Valine (V)
394
392
Alanine (A)
1
3
Methionine (M)
1
0
Glutamic acid (E)
0
1
Glycine (G)
1
0
Isoleucine (I)
0
1
- When describing the last results section, "CHIKV E1 binding proteins exhibit potent anit-CHIV activities" the authors use macrophages. In the rest of the text they consistently use THP-1 macrophages or human primary monocyte derived macrophages. The details of the cell type are extremely useful to the reader and having those in the last results section would be great.
We thank Reviewer 1 for pointing out the importance of cell type clarification in the last results section. We now consistently use “THP-1 derived macrophages” instead of “macrophages” in this section.
- The paper is well-written. There is a slight disconnect as the authors go from discussing results in Figure 4 to Figure 5.
We thank Reviewer 1 for the comment regarding the disconnection of the last two figures in this paper which is also shared by the other reviewers. We have taken 3 approaches to address this comment: 1) We performed a pulldown of the host factors (SPCS3, eIF3k) identified in Figure 5 with CHIKV positively selected mutants examined in Figure 4 with deficient virion production. The result is presented in our response to Reviewer 3’ s major comment #1, suggesting that the positively selected site in E1 is essential for CHIKV glycoprotein interaction with host factors. 2) To complement our first experiment, we will also determine structural protein expression and processing of parental and E1 mutant CHIKV in eIF3k CRISPR knockout 293T cells. 3) Finally, we plan to perform CORUM analysis to identify high confidence functional protein complexes using our 14 hits found in both mass spec experiments, which will provide mechanistic insights into how these identified cellular complexes and processes might modulate CHIKV infection.
Reviewer 2’s major comments
The authors elegantly demonstrate that CHIKV structural proteins confer an advantage over ONNV structural proteins in a step in the replication cycle downstream of virus RNA synthesis, possibly virion assembly. This point would be strengthened determining the particle-to-PFU ratio of the parental viruses and the chimeras . Presumably, the ratio would increase in the chimeras containing CHIKV structural proteins.
We thank Reviewer 2 for this comment. We agree that determining particle-to-PFU ratios of parental and chimeric viruses will strengthen this study. To obtain the particle-to-PFU ratio, we infected THP-1 derived macrophages with CHIKV, ONNV and chimeras containing CHIKV glycoproteins (Chimera I, and ONNV/CHIKV E2+E1) for 24 h. To quantify the secreted viral particles, we extracted viral RNA in the supernatant and detected (+) viral RNA through TaqMan assay with specific nsp1 probes. The released infectious virions were evaluated through plaque assay. The particle-to-PFU ratios are summarized in the table below. The results show that ONNV has the highest particle-to-PFU ratio (41398), suggesting defective ONNV genome encapsidated in particles leading to defective virion production. On the other hand, the particle-to-PFU ratio of CHIKV (747) is 55-fold lower than that of ONNV. Replacing E3-E2-6K-E1 of ONNV with CHIKV homologous proteins reduces the particle-to-PFU ratio by 8 fold to 4875. Replacing E2 and E1 of ONNV with the ones from CHIKV (ONNV/CHIKV E2+E1) reduces the particle-to-pfu ratio by 20 fold to 2017, suggesting that CHIKV glycoproteins enhance the infectivity of viral progenies produced by THP-1 derived macrophages. We have included the results in Figure 3D-3E in our partially revised manuscript and described in lines 149-158.
- Additionally, the authors should consider performing virion assembly blocking assays with a small molecule inhibitor to determine if this abrogates the virus production advantage of CHIKV structural proteins within the ONNV backbone.
We thank Reviewer 2 for this insightful comment. As the secretory pathway is commonly important for alphavirus glycoprotein maturation and assembly, it will be informative to interrogate CHIKV glycoprotein trafficking and assembly through this pathway using specific inhibitors, such as dihydropyridine FLI-06 and golgicide A . Golgicide A is a reversible inhibitor of the cis-Golgi GBF1, which leads to rapid disassembly of the Golgi and trans-Golgi network (TGN)4. FLI-06 is a new inhibitor that interferes with cargo recruitment to ER-exit sites and disrupts Golgi without depolymerizing microtubules or interfering GBF15. We pretreated THP-1 derived macrophages with 10 uM FLI-06 or golgicide A for 30 mins prior to infection with CHIKV, ONNV, Chimera I, or ONNV/ CHIKV E2+E1. After 1 hour of virus adsorption in PBS with 1% FBS in the absence of the inhibitors, the cells were treated with the inhibitors at the same concentration (10uM) in complete medium for 24 h. The plaque assay result shows that all the viruses are sensitive to secretory pathway inhibition, however, the production of viruses containing CHIKV glycoproteins is significantly more attenuated by FLI-06 and golgicide A. This suggests that CHIKV glycoproteins-mediated trafficking and assembly is more heavily dependent on the host secretory pathway . We will include this result in the fully revised manuscript.
- Finally, the authors should perform competition experiments with the chimeric viruses and ONNV in macrophages to determine if the chimeras can outcompete the parental ONNV strain. Based on their data, the chimeric viruses should outcompete.
We thank Reviewer 2 for this inspiring suggestion. The competition experiment is an innovative and informative way to evaluate whether CHIKV glycoproteins confer a selective advantage on virion production in THP-1 derived macrophages. We plan to infect THP-1 derived macrophages with ONNV and ONNV/CHIKV E2+E1 and detect the viral glycoproteins secreted in the supernatant by western blot, although there is a possibility that this experiment might not work due to superinfection exclusion. Given that there is no commercial antibody of ONNV available, we need to use tagged viruses for this competition experiment. We constructed ONNV/CHIKV myc-E2+E1 that has a myc tag at the N-terminus of CHIKV E2, and ONNV/HA-E2 that has a HA tag at the N-terminus of ONNV E2. Our first attempt at concentrating the viral progenies released by THP-1 derived macrophages infected with the two tagged viruses has not been successful. We performed sucrose gradient ultracentrifugation of the supernatant viral particles but the myc and HA tags were not detected in the expected sucrose layer. Next, we plan to use myc-Ab and HA-Ab conjugated beads to pull down the supernatant viral particles to detect the ratio of ONNV/CHIKV myc-E2+E1 and ONNV/HA-E2 secreted by THP-1 derived macrophages. This will determine whether ONNV containing CHIKV glycoproteins can outcompete ONNV in co-infected cells due to increased viral fitness.
- The authors use both primary macrophages and macrophage cell lines as their in vitro model system and make one of their major points (listed in the title) that the determinants they identified in the CHIKV structural proteins convert macrophages into dissemination vessels; however, they do not show: 1) an in vivo model that the CHIKV-ONNV chimeras disseminate more efficiently than the parental ONNV; and 2) that these chimeras generate virus more efficiently specifically in macrophages. It would be useful to show that ONNV and CHIKV have equivalent virion production in other cell lines and that the advantage conferred by CHIKV structural proteins in the ONNV backbone is specific to macrophages. The authors should also change their title to reflect that dissemination is not directly being addressed in their study; the implications of their in vitro experimentation in a mammalian host would be more appropriate for the discussion.
We acknowledge the limitations of the study, which include a lack of direct demonstration of in vivo dissemination. To address these concerns, we will include further discussion of our in vitro findings in the context of viral dissemination in mammalian hosts in the fully revised manuscript. We are also testing ONNV, CHIKV, Chimera I and ONNV/CHIKV E2+E1 infections in 293T cells to investigate whether the advantage conferred by CHIKV glycoproteins are macrophage specific.
We have also updated the title to accurately reflect the significance of this research: “Chikungunya virus glycoprotein targeting of host factors increases viral fitness in human macrophage”.
Reviewer 2’s optional comments
- The authors use CHIKV-ONNV chimeras but it would be interesting to test other chimeras to determine if CHIKV structural proteins confer the same advantage in the backbone of other arthritogenic alphaviruses. The study would also be strengthened by using a pathogenic strain of CHIKV instead of the vaccine strain, as this is significantly attenuated in vivo.
We thank Reviewer 2 for this suggestion which is also suggested by Reviewer 1 in their minor comment #5. We plan to use virulent CHIKV La reunion strain and carry out infection experiments in BSL-3 to strengthen this study. We are getting trained in the BSL-3 facility and will be certified soon.
- In Figure 4, the authors identify residues in the CHIKV structural proteins that appear to be under positive selection in human subjects and generate point mutants in these residues with the corresponding ONNV residues. They find that one mutation, V1029I located in E1, completely abolishes virion production in THP-1 macrophage cell lines. However, in their previous chimeric experiments, they find that neither CHIKV E1 or E2 was sufficient to increase virus production in the ONNV backbone. The authors should address this discrepancy, otherwise they should consider moving the data in their point mutation experiments to a supplementary figure. While worthy of reporting, especially given the patient data, these experiments do not buttress the points made in the previous figures.
We thank Reviewer 2 for this insightful comment. According to previous studies, E2 and E1 always interact with each other from the step of the formation of single heterodimer in the ER to heterodimer trimerization before viral particle assembly. Although the E1-V220 site (previously called V1029) on the exterior of a single E2-E1 heterodimer appears to not be engaged in the E2-E1 interaction E1-V220 is partially exposed and protruding into the groove formed by E1 and the E2 of neighboring heterodimer, accessible to host factors. As such, mutating CHIKV E1-V220 to the ONNV residue (E1-V220I) may not only disrupt E2-E1 trimerization but also interfere viral glycoprotein interaction with host factors(presented in our response to Reviewer 1’s major comment #1). Similarly, solely swapping E2 or E1 with CHIKV substitute in the ONNV backbone would also affect the interaction between neighboring E2 and E1 in trimerized spike, which may explain why neither ONNV/CHIKV E2 or ONNV/CHIKV E1 rescues virion production in THP-1 derived macrophages . We have included this in the partially revised discussion section lines __ __296-313.
- The authors conclude their manuscript with an assessment of several host proteins, namely SPCS3 and eIF3k, that were identified by mass spectrometry and whose knockdown results in increased virion production. The authors speculate about the role of these proteins but do not provide any mechanistic detail on how they might be playing a role. It is unclear that the putative antiviral role of these proteins involves steps downstream of virus replication, especially given that the authors speculate translation might be affected by eIF3k which, if the case, RNA synthesis should also be expected to be affected.
We thank Reviewer 2 for this comment. We acknowledge that we have yet a full mechanistic understanding of how SPCS3 and eIF3k impact virion production. We plan to investigate their antiviral roles in our follow-up studies. For our partial revision, we have constructed several single eIF3k knockout (KO) clones of 293T cells. The eIF3k sgRNA we designed targets exon 3 which would eliminate expression of all 3 splice isoforms of eIF3k (KO schematic and sequence verification of CRISPR KO shown in revision plan). Unfortunately, we failed to obtain single clones of 293T cells with SPCS3 complete KO, consistent with a previous study by Rong Zhang et al6 that were unable to recover SPCS3 KO clones likely due to the importance of SPCS3 in cell survival. We infected an eIF3k KO clone (clone 9) with CHIKV vaccine strain 181/clone 25, ONNV SG650, and SINV Toto1101. Interestingly, we found that the antiviral activity of eIF3k is specific to CHIKV as CRISPR KO of eIF3k increases CHIKV production by 2.5 fold but not ONNV or SINV production (shown in revision plan). We have included this in the partially revised manuscript in__ line 272-282 (Figure 7B-7D).__
We presume that Reviewer 2’s inference of eIF3k’s potential effects on viral RNA synthesis is based on our speculation of its antiviral role in viral translation, which may affect viral nonstructural gene expression. We would like to clarify that eIF3k is not an initiation factor traditionally needed for cap-dependent translation. It is also not clear what translation process (nonstructural polyprotein translation from viral genomic RNA or structural polyprotein translation from viral subgenomic mRNA) involves eIF3k if it indeed affects viral protein expression. Notably, previous SINV studies imply that alphavirus structural polyprotein translation may employ unique mechanisms without the requirement of several crucial initiation factors4,5. It will be interesting to see whether eIF3k participates in viral subgenomic mRNA translation as that would affect viral glycoprotein expression leading to reduced virion production. We have now included additional discussion on eIF3k antiviral mechanisms in the partially revised manuscript in lines 345-353.
- Overall, while the initial chimeric virus and domain swap approach is strong, the manuscript would benefit with a more thorough examination of virion assembly steps and a mechanistic link to virion production. Otherwise, the authors should revise the structure of their manuscript by de-emphasizing points about virion assembly and leave room for other mechanistic explanations of their chimeric data that more clearly link the host antiviral factor/E1 binding studies.
We thank the reviewer for these positive comments and suggestions. We have addressed this by further interrogating the production kinetics of CHIKV, ONNV, and the chimeras containing CHIKV glycoproteins through determining their particle-to-PFU ratios as well as treating infected cells with secretory pathway inhibitors (refer to our responses to Reviewer 2 major comments #1 and #2). We have also included additional discussion on eIF3k antiviral mechanisms specifically on how it may affect other steps of the viral life cycle in the partially revised manuscript in lines 345-353 (refer to our response to Reviewer 2 optional comment #3).
Reviewer 3’s critique comments
- Overall, the manuscript is well written but in its current state it is more like two different stories because the effects of envelope proteins and list of interactors are not brought together in one story. A possible fix to this problem would be inclusion of ONNV and CHIKV containing env mutations that do and do not restore viral release from macrophages into the pulldown/association experiments shown in Figure 6.
We thank Reviewer 3 for the insightful suggestions to better connect the first (CHIKV determinants) and second (CHIKV glycoprotein interactors) parts of the manuscript. In response to the Reviewer’s comment, we tested the binding of SPCS3 and eIF3k to CHIKV E1 with E1-V220I (V1029I in original manuscript version) mutation (shown in revision plan) which was shown to abrogate virion production in THP-1 derived macrophages in Figure 4E. We transfected plasmids expressing 3XFLAG-tagged SPCS3/eIF3k or empty vector for 24 h followed by transfection with plasmids expressing either the parental CHIKV vaccine strain 181/clone 25 poly-glycoproteins (E3-myc-E2-6K-E1) or poly-glycoproteins with the E1-V220I mutation. Interestingly, we found that mutating CHIKV E1-V220 to the homologous ONNV residue reduces the binding to either SPCS3 or eIF3k. This result strongly suggests that the positively selected E1-V220 is located in the interaction interface between E1 and SPCS3/eIF3k, confirming the genetic conflict between E1 and these host factors to be one of the major drivers of CHIKV evolution observed at site E1-V220. We have included this result in partially revised manuscript in Figure 7A and in lines 265-271.
- The other major issue is the lack of protein data for the viral mutants relative to WT ONNV and CHIKV and assessment of viral RNA in the supernatants to determine whether the block is release or an earlier event since viral RNA levels in the cell seems to be the same or at least normalized.
We thank Reviewer 3 for pointing out the insufficient clarification of the block leading to defective CHIKV mutant virion production. We previously detected E2 expression from 293T cells transfected with poly-glycoproteins (E3-myc-E2-6K-E1) containing E2-V135L (V460L in original manuscript version), E2-A164T (A489T in original manuscript version), E2-A246S (A571S in original manuscript version) and E1-V220I (V1029I in original manuscript version). We found that only E2-V135L mutation can lead to unexpected E2 cleavage (shown in revision plan) as we mentioned but not shown in the original manuscript. This explains why E2-V135L mutation attenuates infectious CHIKV production.
The E2 expression of E1-V220I appears to be not affected in 293T cells transfected with poly-glycoproteins with E1-V220I (shown in revision plan ). In addition, the E1-host factor binding result in our response to Reviewer 3’s major comment #1 showed that E1 with the positively selected site mutation V220I can also be successfully expressed in 293T cells after transfection with poly-glycoprotein. Based on these current data, E1-V220I mutation likely abrogates virion production without affecting glycoprotein expression.
Our previous result of the ONNV particle-to-PFU ratio reveals that ONNV RNA is released but encapsidated in defective particles causing its attenuation in infected macrophages. Thus, even though the glycoproteins of E1-V220I can be expressed, the diminished virion production of CHIKV E1-V220I can still be ascribed to 1) blocked viral particle release and 2) production of defective particles like ONNV. Given that it is not feasible to obtain particle-to-PFU ratio of E1-V220I mutant which fails to form plaques, Reviewer 3’s suggestion to assess the supernatant viral RNA will be a nice approach to address this question. To further address this concern, we plan to transfect THP-1 derived macrophages with CHIKV E1-V220I mutant RNA to detect the intracellular viral glycoprotein expression and supernatant viral RNA levels through western blot and TaqMan assay, respectively.
- Lastly, knockdown experiments indicate an effect of things like OAS3 or other innate immune modulators. There are no controls to demonstrate that these are specific to CHIKV infection or if knockdown would assist growth of ONNV as well.
We also thank Reviewer 3 for the suggestion to check whether the identified host factors specifically target CHIKV or inhibit the infection of ONNV as well. We previously tried but were facing some issues. Since only a small fraction of macrophages can be infected with CHIKV and even a smaller fraction can be infected with ONNV (Figure 1A), it is hard to elucidate the roles of these identified host factors in ONNV infection by siRNA knockdown. We decided to take a more rigorous approach to investigate the antiviral specificity of identified host factors, especially understudied SPCS3 and eIF3k, to different alphaviruses by generating complete knockout 293T single cell clones. Despite the fact that we did not successfully generate SPCS3 complete KO, we obtained an eIF3k KO single cell clone and infected it with CHIKV, ONNV and SINV (refer to our response to Reviewer 2 optional comment #3). We found that eIF3k only has antiviral activity against CHIKV with almost no effects on ONNV or SINV infection. We have included this in our partially revised manuscript in line 272-282 (Figure 7B-7D).
Reviewer 3's minor comments:
Other points to consider:
- The title does not fit the manuscript findings and should be modified.
We thank Reviewer 3 for this important comment, which was also brought up by Reviewer 2. We have now changed our title to “Chikungunya virus glycoprotein targeting of host factors increases viral fitness in human macrophage”, which more accurately reflects the significance of our research.
- It is unclear why the authors show results for SINV and RRV in Figure 1. Either these should be removed or the viruses should be carried throughout the experiments described in the Figure. Better yet would be to add additional alphaviruses to this analysis to determine if there are additional viruses that act similarly to CHIKV.
We apologize for the confusion caused by including SINV and RRV results in Figure 1. We intended to show the superiority of CHIKV in infecting primary monocyte derived macrophages among arthritogenic alphaviruses, which we speculate may provide the molecular basis for macrophage-mediated CHIKV dissemination and disease. We would like to keep the SINV and RRV infection results in Figure 1 to highlight the relative susceptibility of macrophages to CHIKV. To echo the additional alphaviruses tested in Figure 1 and bring the story full circle, we included the result of SINV infection of eIF3k CRISPR KO 293T cells in Figure 7B-7D. These results uncover inhibitory activities of eIF3k that are specific to CHIKV.
- Is the data presented in Figure 1A significant?
We thank Reviewer 3 for this question. We infected both THP-1 derived macrophages and primary monocyte derived macrophages with EGFP-expressing alphaviruses each in duplicates for two independent times. The general low expression of EGFP in all virus-infected groups refrains us from drawing conclusions based on statistically significant differences observed with MFI, hence we chose to show representative scatter plots in the original manuscript. To address Reviewers 3's question, we plotted the infected cell (EGFP+) based on the percentages of the experimental duplicates (shown in revision plan), and found CHIKV infection to be the most significantly different from that of the other alphaviruses in primary monocyte derived macrophage . The numbers above the bar charts are the mean percentages of EGFP+ cells.
- The justification for inclusion of Figure 4A is lacking. It is unclear what this panel is supposed to be demonstrating.
This is an excellent suggestion as the host factors identified by AP-MS not only contain interactors of CHIKV mature E2 but also those of uncleaved E2-containing precursor polyproteins. We modified Figure 4A to reflect all E2/E2-containing poly-glycoproteins present in CHIKV-infected cells (shown in revision plan).
- There is little justification for the candidates assessed in
We understand Reviewer 3’s concern. Due to the nature of mass spectrometry studies which predict protein-protein interactions rather than direct functional validation, we acknowledge that we may miss some host candidates that have anti- or pro-CHIKV activities. Although justification of hit selection from mass spectrometry datasets is more difficult than that from CRISPR KO screen datasets, we set up specific criteria to identify host protein candidates with the greatest potential to functionally interact with CHIKV glycoproteins. Most of the proteins we chose to validate (Figure 6a) were identified in both of our independent AP-MS experiments, which both pass through a P-value threshold of 0.05 and log2 fold change of 0.
- Extended data Figure 3 is very difficult to read due to the small font size.
We apologize for the small font in Extended data Figure 3. We plan to replace Figure EV3 ( Extended data 3 in unrevised version) with a CORUM protein-protein interaction network that centers on the significant hits identified by both AP-MS experiments, but includes hits from either one of the two experiments in these functional protein complexes. The figure will be more concise and centralized, and the font will be bigger.
- Just to be clear, the blots shown in Figure 6D are different from those depicted in Extended data Figure 4b, because some of them look very similar.
We thank Reviewer 3 for this question. In Figure 6D, we expressed CHIKV glycoproteins through transfecting CHIKV genomic RNA into 293T cells, while, in Figure 4B, we expressed CHIKV glycoproteins through transfecting poly-glycoprotein plasmid (pcDNA3.1-E3-myc-E2-6K-E1) into 293T cells, which are complementary approaches to express CHIKV glycoproteins to validate their interactions with identified host factors. We have now added schematics to illustrate the different experimental strategies above the figures in this partially revised manuscript (shown in revision plan).
References:
Voss, J. E. et al. Glycoprotein organization of Chikungunya virus particles revealed by X-ray crystallography. Nature 468, 709–712 (2010). Jose, J., Tang, J., Taylor, A. B., Baker, T. S. & Kuhn, R. J. Fluorescent Protein-Tagged Sindbis Virus E2 Glycoprotein Allows Single Particle Analysis of Virus Budding from Live Cells. Viruses 7, 6182–6199 (2015). Götte, B., Liu, L. & McInerney, G. M. The Enigmatic Alphavirus Non-Structural Protein 3 (nsP3) Revealing Its Secrets at Last. Viruses 10, 105 (2018). Saenz, J. B. et al. Golgicide A reveals essential roles for GBF1 in Golgi assembly and function. Nat. Chem. Biol. 5, 157–165 (2009). Krämer, A. et al. Small molecules intercept Notch signaling and the early secretory pathway. Nat. Chem. Biol.9, 731–738 (2013). Zhang, R. et al. A CRISPR screen defines a signal peptide processing pathway required by flaviviruses. Nature 535, 164–168 (2016).